
Trends
Sci.
2026;
23(5):
12554
Mechanotransduction in Cellular Physiology: From Cytoskeleton Dynamics to Gene Expression
Aswin
Rafif Khairullah1,
![]() ,
Abdul
Hadi Furqoni2,
,
Abdul
Hadi Furqoni2,
![]() ,
Analinasari Analinasari3,
,
Analinasari Analinasari3,
![]() ,
,
Arif Nur Muhammad Ansori4,5,6,
![]() ,
Bantari Wisynu Kusuma Wardhani7,
,
Bantari Wisynu Kusuma Wardhani7,
![]() ,
,
Bima
Putra Pratama8,
![]() ,
Dea Anita Ariani Kurniasih9,
,
Dea Anita Ariani Kurniasih9,
![]() ,
Eny Martindah1,
,
Eny Martindah1,
![]() ,
,
Fadhila
Utari7,
![]() ,
Ilma Fauziah Maruf7,
,
Ilma Fauziah Maruf7,
![]() ,
Imam Mustofa10,
,
Imam Mustofa10,
![]() ,
,
Mohammad
Sukmanadi11,*,
![]() ,
Riza Zainuddin Ahmad1,
,
Riza Zainuddin Ahmad1,
![]() ,
Saifur Rehman12,
,
Saifur Rehman12,
![]() ,
,
Siti
Hamidatul Aliyah2,
![]() and Wita
Yulianti13,
and Wita
Yulianti13,
![]()
1Research Center for Veterinary Science, National Research and Innovation Agency (BRIN), Bogor,
West Java 16911, Indonesia
2Center for Biomedical Research, National Research and Innovation Agency (BRIN), Bogor,
West Java 16911, Indonesia
3Postgraduate Program in Applied Master of Food Security, Lampung State Polytechnic (Polinela), Bandar Lampung, Lampung 35144, Indonesia
4Postgraduate School, Universitas Airlangga, Kampus B Dharmawangsa, Surabaya, East Java 60286, Indonesia
5Uttaranchal Institute of Pharmaceutical Sciences, Uttaranchal University, Uttarakhand 248007, India
6Medical Biotechnology Research Group, Virtual Research Center for Bioinformatics and Biotechnology, Surabaya, East Java 60493, Indonesia
7Research Center for Pharmaceutical Ingredients and Traditional Medicine, National Research and Innovation Agency (BRIN), Bogor, West Java 16911, Indonesia
8Research Center for Process Technology, National Research and Innovation Agency (BRIN), South Tangerang,
Banten 15310, Indonesia
9Research Center for Public Health and Nutrition, National Research and Innovation Agency (BRIN), Bogor,
West Java 16911, Indonesia
10Division of Veterinary Reproduction, Faculty of Veterinary Medicine, Universitas Airlangga, Kampus C Mulyorejo, Surabaya, East Java 60115, Indonesia
11Division of Basic Veterinary Medicine, Faculty of Veterinary Medicine, Universitas Airlangga, Kampus C Mulyorejo, Surabaya, East Java 60115, Indonesia
12Department of Pathobiology, Faculty of Veterinary and Animal Sciences, Gomal University,
Dera Ismail Khan 27000, Pakistan
13Research Center for Biota Systems, National Research and Innovation Agency (BRIN), Bogor,
West Java 16911, Indonesia
(*Corresponding author’s e-mail: [email protected])
Received: 6 November 2025, Revised: 3 January 2026, Accepted: 9 January 2026, Published: 15 January 2026
Abstract
Mechanotransduction is the process through which cells convert mechanical cues from their microenvironment into biochemical signals that regulate gene expression and cellular behavior. This review aims to critically integrate current evidence on how mechanical forces are transmitted from the extracellular matrix through adhesion complexes and cytoskeletal structures to the nucleus, thereby shaping transcriptional regulation. The discussion focuses on the coordinated roles of cytoskeletal dynamics and mechanosensitive signaling pathways, including YAP/TAZ, MRTF-A, and RhoA/ROCK, in controlling cellular proliferation, differentiation, and tissue adaptation. The physiological relevance of these mechanisms is examined in key biological contexts such as stem cell lineage specification, skeletal muscle adaptation to mechanical loading, and endothelial regulation under hemodynamic forces. In parallel, the review addresses how disruptions in mechanotransduction contribute to disease pathogenesis, particularly in cancer, fibrosis, and atherosclerosis, where altered mechanical environments lead to aberrant gene regulatory responses. Finally, this article highlights emerging methodological advances, including three-dimensional culture systems, organ-on-chip platforms, and integrated multi-omics approaches, as essential tools for improving mechanobiological modeling and identifying novel therapeutic targets related to mechanically driven cellular dysfunction.
Keywords: Mechanotransduction, Cytoskeleton, Disease, Integrins, Gene expression, Mechanical signaling
Introduction
Mechanical forces are key factors that significantly determine cellular function and behavior [1]. Every living cell operates in an environment that constantly exerts mechanical influences through tension, pressure, strain, and shear forces originating from the extracellular matrix (ECM), intercellular interactions, and tissue fluid movement [2]. Variations in these forces not only influence cell morphology and architecture but also regulate important biological processes such as migration, proliferation, differentiation, and apoptosis [3]. In the context of tissue physiology, mechanical forces serve as primary signals that enable cells to adapt to microenvironmental dynamics, maintain functional balance, and preserve structural integrity and organ homeostasis [4].
Mechanotransduction is a fundamental mechanism by which cells convert mechanical stimuli into biochemical responses that play a role in regulating cellular function [5]. This process involves coordinated interactions between cytoskeletal elements, adhesion complexes, mechanosensory proteins, and various intracellular signaling components [6]. Through mechanotransduction, physical changes in the cell's environment are translated into changes in protein conformation, activation of signaling cascades, and regulation of gene expression [7]. The cytoskeleton—composed of actin filaments, microtubules, and intermediate filaments—acts as a dynamic structural network that transmits mechanical forces from the plasma membrane to the nucleus and organelles, thus linking mechanical responses to genetic regulatory mechanisms within the cell [8].
The physiological significance of mechanotransduction is evident in a wide range of biological processes. In stem cell differentiation, the mechanical properties of the extracellular matrix play a role in determining the direction of cell specialization: A matrix with high stiffness promotes osteogenic differentiation, while a softer matrix favors differentiation toward neurogenic or adipogenic lineages [9]. In the cardiovascular system, shear forces generated by blood flow influence the activation of mechanosensitive transcription factors such as krüppel-like factor 2 (KLF2) and endothelial nitric oxide synthase (eNOS), which play a role in maintaining vasodilation and blood pressure stability [10]. In skeletal muscle tissue, repetitive strain during contraction stimulates the phosphoinositide 3-kinase–protein kinase B–mechanistic target of rapamycin (PI3K–Akt–mTOR) and mitogen-activated protein kinase/extracellular signal-regulated kinase (MAPK/ERK) pathways, thereby increasing protein synthesis, muscle hypertrophy, and mitochondrial biogenesis in response to increased mechanical load [11]. Furthermore, mechanotransduction plays a crucial role in wound healing, tissue remodeling, and embryonic development, where the integration of mechanical and chemical signals ensures the spatial and temporal coordination of cell growth, migration, and differentiation [12].
Dysregulation of mechanotransduction has important implications for various pathological processes. Increased tissue stiffness in fibrosis can trigger overactivation of mechanosensitive pathways such as Yes-associated protein/Transcriptional coactivator with PDZ-binding motif (YAP/TAZ), which promotes fibroblast differentiation into myofibroblasts and accelerates extracellular matrix deposition [13]. In the context of cancer, changes in the mechanical characteristics of the tumor microenvironment increase cytoskeletal tension, contributing to uncontrolled cell proliferation, epithelial–mesenchymal transition (EMT), and increased metastatic potential [14]. Meanwhile, in atherosclerosis, nonlaminar blood flow inhibits endothelial anti-inflammatory gene expression and favors vascular plaque formation [15]. Overall, these phenomena emphasize that the balance of mechanotransduction plays a crucial role in determining whether cellular responses will be adaptive or pathological [5].
The aim of this review is to provide a comprehensive understanding of mechanotransduction in cellular physiology, with an emphasis on the interplay between cytoskeleton dynamics, mechanosensors, and gene expression regulation. This article will discuss the key signaling pathways that translate mechanical forces into transcriptional activity, highlighting the physiological role of mechanotransduction in cell differentiation, tissue adaptation, and maintenance of homeostasis. Furthermore, this review will evaluate the impact of mechanotransduction dysregulation on various pathological conditions and highlight advances in experimental technology that enable quantitative analysis of mechanotransduction, providing a basis for further research and developing mechanobiology-based therapeutic strategies.
Data collection method
This scoping review was conducted in accordance with the PRISMA-ScR guidelines. A comprehensive literature search was performed in PubMed/MEDLINE, Scopus, and Web of Science to identify English-language studies published between January 1, 2008, and November 3, 2025. Controlled vocabulary terms and relevant keywords were used to capture studies related to mechanotransduction and mechanosensing; integrins, cadherins, and PIEZO channels; focal adhesion and cytoskeletal signaling pathways, including FAK/Src, RhoA/ROCK, MAPK/ERK, and PI3K/AKT; nucleo-cytoskeletal coupling mechanisms such as the LINC complex and lamins; and nuclear and chromatin-associated regulators, including YAP/TAZ and MRTF-A, in relation to gene expression.
Eligible studies included in vitro and in vivo primary research as well as mechanistic reviews that explicitly linked mechanical stimuli to biochemical signaling and gene regulatory outcomes. Non-peer-reviewed publications, non-English articles, and studies lacking defined mechanical perturbation or measurable mechanobiological readouts were excluded. Data extraction was independently performed by two reviewers using a standardized framework encompassing experimental system, mechanical stimuli, signaling routes, nuclear or chromatin effects, functional readouts, and biological context, with discrepancies resolved through consensus. Study design features and potential sources of bias were assessed qualitatively, and the findings were synthesized narratively along a force-to-gene continuum without conducting a meta-analysis.
Basic mechanisms of mechanotransduction
Mechanotransduction plays an essential role in maintaining the physiological balance of cells by transforming mechanical stimuli from the environment into biochemical signals that regulate function and gene expression. This process involves a network of structural and molecular components that interact synergistically to detect, transmit, and integrate mechanical forces into appropriate biological responses at the cellular level.
Main components of mechanotransduction
Mechanotransduction is increasingly understood not as a linear signaling cascade, but as an integrated, multiscale system in which mechanical cues are sensed, transmitted, and interpreted through coordinated interactions between the plasma membrane, the extracellular matrix (ECM), and the cytoskeleton [16,17]. While early models emphasized single mechanosensors, more recent frameworks highlight the cooperative and context-dependent nature of these components, where mechanical information is distributed across multiple structural elements rather than processed by a single dominant pathway [18].
At the cell surface, the plasma membrane functions as the primary interface for mechanical input; however, its mechanosensory capacity largely depends on associated adhesion complexes rather than the lipid bilayer itself [19]. Integrins and cadherins represent two mechanistically distinct but complementary adhesion systems. Integrin-based adhesions primarily mediate cell–ECM interactions and are optimized for sensing substrate stiffness and tensile forces, whereas cadherin-based junctions specialize in transducing intercellular tension within tissues [20]. This functional divergence explains why integrin signaling dominates in studies of matrix stiffness, while cadherin-mediated pathways are more prominent in morphogenesis and collective cell behavior.
Integrin-mediated mechanotransduction has been extensively characterized through focal adhesion complexes involving talin, vinculin, paxillin, and focal adhesion kinase (FAK) [21]. In stiffness-driven models, force-induced unfolding of talin exposes vinculin-binding sites, reinforcing integrin–actin coupling and activating downstream signaling cascades such as RhoA/ROCK, PI3K/AKT, and MAPK [22]. However, the relative contribution of these pathways varies across experimental systems. For example, RhoA/ROCK signaling dominates force transmission in mesenchymal cells, whereas PI3K/AKT signaling appears more relevant in epithelial or endothelial contexts. These differences underscore the limitation of extrapolating mechanotransduction mechanisms from single cell types or two-dimensional substrates.
In contrast, cadherin-based adhesions transmit mechanical forces between neighboring cells and regulate tissue-scale mechanics [23]. Tension across cadherins modulates β-catenin availability and transcriptional activity, linking mechanical coupling to cell fate decisions during development and tissue remodeling [24]. Notably, some studies suggest that cadherin-mediated mechanotransduction can attenuate YAP/TAZ activity under high cell density, highlighting a mechanistic counterbalance to integrin-driven signaling. This apparent contradiction illustrates how mechanical inputs can produce divergent transcriptional outcomes depending on adhesion context and tissue architecture.
The ECM acts not merely as a passive scaffold but as an active regulator of mechanotransduction by defining the magnitude, direction, and persistence of mechanical forces [25]. Variations in ECM stiffness, elasticity, and topology modulate cellular contractility and focal adhesion maturation, thereby influencing intracellular tension [26]. While increased ECM stiffness is frequently associated with YAP/TAZ activation and pro-proliferative gene expression, this relationship is not universal. In highly confined three-dimensional environments, for example, nuclear deformation rather than matrix stiffness may dominate mechanosensitive signaling, suggesting that stiffness-centric models have limited applicability outside simplified culture systems [27].
The cytoskeleton integrates signals from adhesion complexes and distributes mechanical forces throughout the cell, forming a continuous force-transmission network from the plasma membrane to the nucleus [31]. Actin filaments, microtubules, and intermediate filaments contribute distinct mechanical functions: Actin generates contractile tension, microtubules provide compressive resistance and spatial organization, and intermediate filaments enhance mechanical resilience under large deformations [32-34]. Importantly, these components do not act independently. Crosstalk between actin and microtubules fine-tunes intracellular tension, while intermediate filaments modulate force dissipation, particularly under pathological mechanical stress.
Changes in cytoskeletal tension can directly influence nuclear shape, chromatin organization, and transcriptional accessibility [35]. However, the extent to which cytoskeletal forces alone regulate gene expression remains debated, as biochemical signaling pathways often operate in parallel. Thus, while the cytoskeleton is indispensable for mechanical signal propagation, its regulatory role must be interpreted within a broader mechano-biochemical framework rather than as a standalone determinant of gene regulation [36].
Conversion of mechanical forces into biochemical signals
The defining feature of mechanotransduction lies in the conversion of physical forces—such as tension, compression, and shear stress—into biochemical signals that regulate cellular behavior [37]. Rather than relying on a single mechanosensor, cells employ multiple force-sensitive proteins that respond through conformational changes, altered binding affinities, and activation of downstream signaling pathways [38]. This redundancy enhances robustness but also introduces complexity and variability across experimental models.
Focal adhesion proteins such as talin, vinculin, and paxillin serve as prototypical mechanosensitive elements [39,40]. Talin unfolding under tensile load exposes cryptic vinculin-binding sites, reinforcing integrin–actin coupling and stabilizing focal adhesions [41]. While this mechanism is well supported in vitro, its quantitative contribution in vivo remains difficult to resolve, as force magnitudes and loading rates differ substantially across tissues. Moreover, not all integrin-dependent signaling requires talin unfolding, suggesting alternative force-sensing modes within adhesion complexes [42].
Mechanosensitive ion channels, particularly Piezo1, represent a distinct class of mechanotransducers that directly couple membrane deformation to intracellular signaling [43]. Piezo1-mediated Ca²⁺ influx activates downstream pathways such as CaMKII and calcineurin–NFAT, influencing transcriptional programs related to calcium homeostasis and inflammation [44]. Compared to adhesion-based mechanotransduction, ion channel–mediated signaling operates on faster timescales but may lack the spatial specificity provided by focal adhesions. This functional distinction highlights how different mechanotransduction pathways are preferentially engaged depending on the nature and duration of mechanical stimuli.
At the nuclear level, YAP and TAZ function as central integrators of mechanical signals derived from both cytoskeletal tension and adhesion-mediated signaling [45]. In rigid or highly contractile environments, YAP/TAZ translocate to the nucleus and activate transcriptional programs that promote proliferation and differentiation [46]. However, emerging evidence indicates that YAP/TAZ activity can also be modulated independently of substrate stiffness, for example through nuclear deformation or changes in cell geometry. These findings challenge stiffness-centric models and suggest that multiple mechanical parameters converge on YAP/TAZ regulation [47].
A unifying principle across mechanosensitive proteins is their capacity for force-induced conformational change, which exposes hidden binding sites and alters signaling networks [48,49]. Talin stretching and lamin A/C deformation exemplify how mechanical forces directly modify protein structure and function [50,51]. Nonetheless, detecting these conformational changes in living tissues remains technically challenging, limiting direct validation of many proposed mechanisms [52].
Downstream of mechanosensors, signaling pathways such as RhoA/ROCK, MAPK/ERK, and PI3K/AKT coordinate cytoskeletal remodeling, survival, and transcriptional responses [21,53-56]. While these pathways are often presented as parallel outputs of mechanotransduction, evidence suggests substantial cross-regulation and context dependence. For instance, RhoA/ROCK-driven contractility can enhance YAP/TAZ activity, whereas PI3K/AKT signaling may modulate mechanosensitivity indirectly through metabolic regulation. This interdependence complicates efforts to isolate linear cause–effect relationships.
Overall, mechanotransduction emerges as a distributed signaling network in which mechanical inputs are decoded through multiple, partially overlapping biochemical pathways [57]. The conversion of mechanical forces into biochemical signals is therefore not a singular event but a dynamic and context-dependent process shaped by cell type, dimensionality, and mechanical history. Recognizing these limitations is essential for interpreting experimental findings and translating mechanotransduction principles into physiological and pathological contexts [38]. Bottom of Form
As shown in Figure 1, ECM stiffness is sensed by integrins/cadherins and PIEZO1, transmitted through focal adhesions and the cytoskeleton, and culminates in YAP/TAZ and MRTF-A/SRF–driven gene expression.
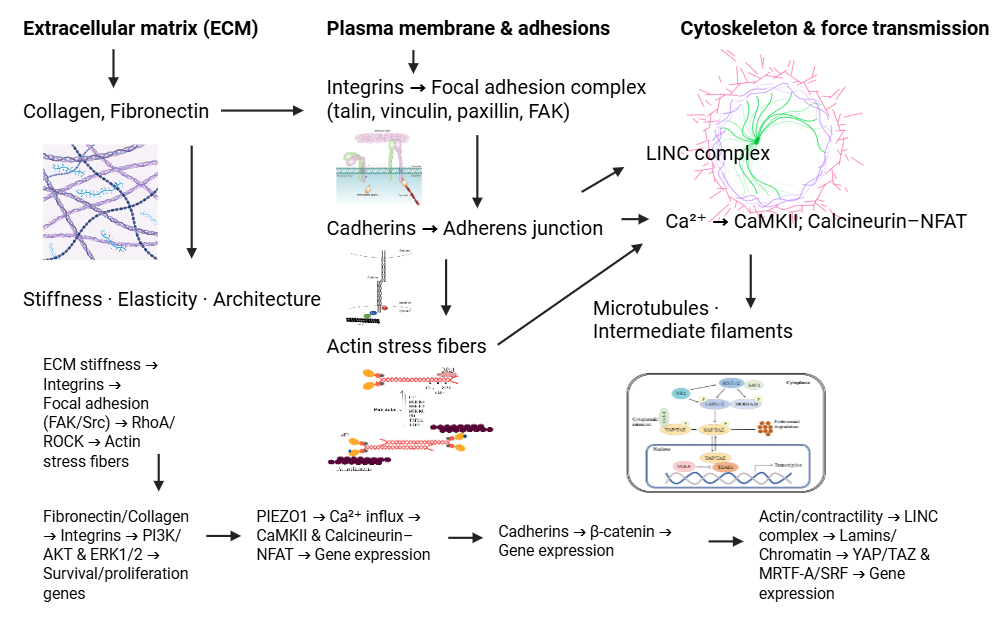
Figure 1 Mechanotransduction across the ECM–adhesion–cytoskeleton–nucleus axis.
Cytoskeleton dynamics in mechanotransduction
The cytoskeleton functions as a dynamic structural network, transmitting mechanical forces from the external environment into the cell while coordinating the resulting biochemical responses. Changes in cytoskeletal organization and tension are key elements of mechanotransduction, allowing cells to adjust their shape, adhesion, and genetic activity according to mechanical stimuli. Table 1 summarizes the major roles of cytoskeleton components, including actin and microtubules, their interactions, and focal adhesion complexes, in cellular mechanotransduction.
Table 1 The role of cytoskeleton dynamics and adhesion complexes in cellular mechanotransduction.
Component |
Role in mechanotransduction |
Signaling pathway/Mediator |
Functional impact/Cellular response |
Actin filaments |
Generates and transmits contractile force through interaction with myosin II and forms stress fibers |
RhoA/ROCK, MLC phosphorylation, YAP/TAZ, and MRTF-A |
Cytoskeleton reorganization, translocation of mechanosensitive transcription factors to the nucleus, regulation of gene expression, cell differentiation, and structural adaptation |
Microtubules |
Balancing actin contractile forces, distributing intracellular tension, and transporting vesicles and organelles |
Dynein, Kinesin, and RhoA regulation |
Maintaining mechanical homeostasis, cellular plasticity, spatiotemporal tension regulation, and coordination of cell migration and shape |
Actin–Microtubule interactions |
Coordination of contractile and compressive forces and regulation of membrane protrusions (lamellipodia/filopodia) |
Formins, CLASP, IQGAP1, and the LINC complex |
Transmission of force to the nucleus, directed cell migration, and physiological adaptation to mechanical stress |
Focal adhesion complex |
The interface of internal and external forces and converting mechanical stimulation into molecular responses |
Integrin, FAK, Src, talin, and paxillin |
Adhesion stability, cytoskeleton reorganization, force orientation, activation of MAPK/ERK, PI3K/AKT, and Rho GTPases pathways |
Integrin |
Force sensors and transducers and bonding with the ECM |
Talin and vinculin |
Force reinforcement, stable cell adhesion, and adaptive responses to external mechanical stress |
Actomyosin contraction |
Major source of internal contractile force; transmits tension to adhesion complexes and ECM |
Actin–myosin II, MLC phosphorylation, and RhoA/ROCK |
Regulates cytoskeletal tension, intracellular force orientation, and mediates force transmission to the nucleus via LINC |
The role of actin and microtubules in force transmission
Actin and microtubules are the main components of the cytoskeleton, playing a central role in the transmission of mechanical forces within cells [58]. These two structures not only provide support for cellular architecture but also function as dynamic mediators that transmit forces from the plasma membrane to the nucleus and intracellular organelles [31]. Through coordinated interactions, the actin-microtubule network enables cells to detect, transmit, and adapt in response to changes in forces from both the external and internal environments [59].
Actin filaments form a dynamic network that functions in generating and transmitting contractile force through interactions with the motor protein myosin II [60]. The formation of stress fibers around focal adhesion points allows the tensile force generated by actomyosin contraction to be transmitted to the adhesion complex and the ECM [61].
This activity is mediated by the RhoA/ROCK pathway, which regulates the phosphorylation of myosin light chains (MLCs) to enhance cytoskeleton contractility [62]. Furthermore, actin filament reorganization directly influences the cell's mechanosensitive response [63]. Increased actin tension can stimulate the translocation of mechanosensitive transcription factors such as YAP/TAZ and Myocardin-Related Transcription Factor A (MRTF-A) into the nucleus, thereby modulating the expression of genes that regulate cell growth and differentiation [64]. Thus, actin functions not only to maintain the mechanical integrity of the cell but also as a functional bridge connecting physical forces to the regulation of genetic expression [65].
Microtubules act as a counterbalance to the contractions generated by actin filaments [66]. Composed of α- and β-tubulin heterodimers, microtubules form a semi-rigid tubular network that can dynamically polymerize and depolymerize [67]. This structure serves to distribute forces throughout the cell and maintain mechanical balance between focal adhesion sites and the nucleus [68].
Furthermore, microtubules play a crucial role in the transport of vesicles and organelles through the motor proteins dynein and kinesin, which also help distribute tension within the cell [69]. Microtubule instability, for example, due to depolymerizing agents like nocodazole, can increase actin contractility and modify cell shape and stiffness [70].
The reciprocal interaction between microtubules and actin allows for spatiotemporal force regulation, where microtubule depolymerization can trigger RhoA activation, while microtubule stabilization tends to suppress contractile signals [71]. This cross-linking makes microtubules key regulators in maintaining mechanical balance and cell plasticity [72].
Actin and microtubules function in an integrated manner through linker proteins such as formins, Cytoplasmic Linker-Associated Protein (CLASP), and IQ Motif Containing GTPase Activating Protein 1 (IQGAP1), which help coordinate the dynamics of both components [66]. This collaboration is crucial for regulating cell migration, the formation of membrane protrusions such as lamellipodia and filopodia, and the transmission of force to the nucleus via the Linker of Nucleoskeleton and Cytoskeleton (LINC) complex [73].
By combining the contractile forces generated by actin filaments and the compressive forces of microtubules, cells are able to precisely adjust their mechanical response to environmental changes [74]. This synergy maintains a balance between structural strength, elasticity, and the physiological adaptation capacity of cells to various mechanical stresses [33].
Adhesion complex and contractile forces
The cell adhesion complex acts as a central hub for transmitting mechanical forces while also regulating biochemical signals that influence cell behavior [20]. This structure enables integration between external forces from the ECM and internal forces generated by the actomyosin system [75]. Through precise coordination between adhesion proteins, cytoskeletal elements, and signaling enzymes, the adhesion complex serves as a dynamic interface that translates mechanical stimuli into molecular responses that regulate cell morphology, motility, and gene expression [76].
Actomyosin contraction is the primary source of contractile force in cells and plays a key role in regulating cytoskeletal tension [77]. This process relies on the interaction between actin filaments (F-actin) and myosin II, which is controlled through phosphorylation [78]. Activation of RhoA by Rho-associated kinase (ROCK) increases myosin light chain (MLC) phosphorylation, thereby activating myosin II ATPase activity and enhancing actin filament contractility [79].
The force generated by actomyosin contraction is transmitted to the focal adhesion complex and the extracellular matrix via integrins, forming a mechanical feedback loop that regulates adhesion stability and force direction within the cell [80]. The dynamics of this contraction are spatiotemporal, with the magnitude and direction of the force determining the cell's response to microenvironmental conditions [81]. For example, increased internal tensile forces can trigger focal adhesion reorganization, increasing cell stiffness, and enhancing force transmission to the nucleus through the LINC complex [82].
FAK is a non-receptor tyrosine kinase that plays a key role in mechanotransduction in focal adhesion complexes [83]. FAK activation begins when integrins bind to ECM ligands, such as fibronectin or collagen, triggering integrin oligomerization and FAK recruitment to the adhesion site [84]. Next, FAK undergoes autophosphorylation at tyrosine residue 397, forming a binding site for Src kinase and other adaptor proteins, including paxillin and talin [85].
Activation of the focal adhesion kinase–steroid receptor coactivator (FAK–Src) complex triggers a series of downstream signaling cascades, including the MAPK/ERK, PI3K/AKT, and rho guanosine triphosphate hydrolases (Rho GTPases) pathways, which play a role in regulating cytoskeleton reorganization, proliferation, and cell survival [86]. Furthermore, integrin- and FAK-mediated signals can also modulate gene expression through the activation of mechanosensitive transcription factors such as YAP/TAZ and Nuclear Factor kappa-light-chain-enhancer of activated B cells (NF-κB) [87].
Integrins function as both sensors and conductors of mechanical force [88]. Upon tensile stress, integrins undergo conformational changes that increase their affinity for ECM ligands and strengthen their association with actin filaments via talin and vinculin [89]. This mechanism forms a force reinforcement mechanism, allowing cells to maintain stable adhesion while adaptively responding to changes in external mechanical stress [90]. As shown in Figure 2, external mechanical cues are relayed from the ECM through integrins and focal adhesions to actin and microtubules, then to the nucleus via the gene expression complex.
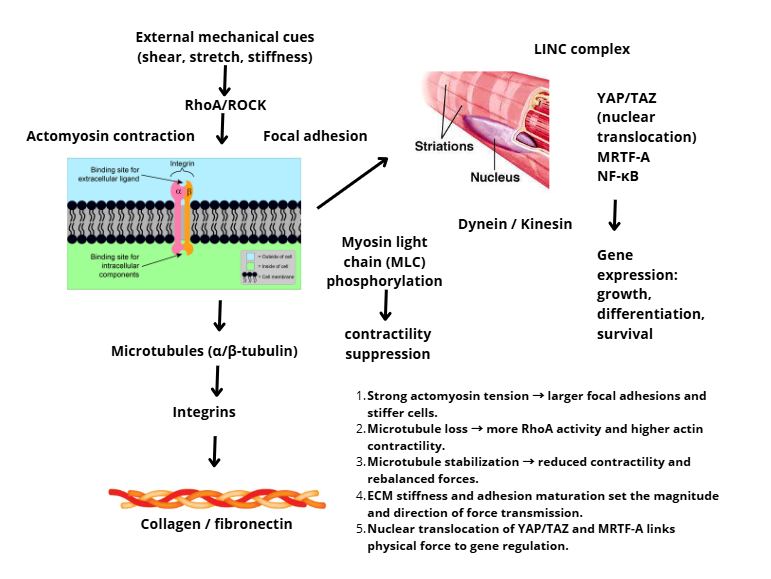
Figure 2 Cytoskeletal force transmission in mechanotransduction.
Signaling pathways and regulation of gene expression
Mechanical signals received by cells are not confined to the cytoskeleton but are transmitted to the nucleus to regulate gene transcription and expression. This process involves complex interactions between cellular structures, adaptor proteins, and transcription factors that collectively determine the cell's response to mechanical forces. Table 2 presents a summary of the major signaling pathways and transcription factors that link mechanical forces to the regulation of gene expression in mechanotransduction.
Table 2 Mechanotransduction signaling pathways and regulation of cellular gene expression.
Components/Factors |
Activation mechanism |
Target/Downstream path |
Functional impact/Cellular response |
LINC Complex (Nesprin-SUN) |
Connects the cytoskeleton to the nuclear lamina and transmits mechanical stress to the nucleus |
Lamin A/C and chromatin remodeling |
Regulates nuclear rigidity, chromatin accessibility, and expression of genes related to proliferation and differentiation |
YAP / TAZ |
Dephosphorylation due to stiff substrate or high tension and translocation to the nucleus |
TEAD → CTGF, CYR61, and ANKRD1 |
Activation of proliferative genes and tissue remodeling and supports cell differentiation and growth |
MRTF-A |
Detach from G-actin as actin polymerizes and enters the nucleus |
SRF → ACTA2, VCL, and MYH9 |
Regulates cytoskeletal gene expression, contractility, and cell morphology |
NF-κB |
Membrane deformation or cytoskeletal tension |
ICAM1, VCAM1, and IL6 |
Triggers pro-inflammatory gene expression and vascular and immune responses |
β-catenin |
Activation by mechanical stretching |
Wnt target genes |
Supports cell proliferation and tissue repair |
KLF2 |
Response to shear stress (shear flow) |
Anti-inflammatory and antithrombogenic genes |
Suppresses pro-inflammatory gene expression and maintains vascular homeostasis |
Epigenetics |
Nuclear deformation and cytoskeletal stress |
Histone acetyltransferase (p300/CBP) and DNMT1 |
Chromatin modification, regulation of mechanosensitive gene expression, and long-term regulation of cell responses |
Mechanical force responsive gene |
Activation by YAP/TAZ, MRTF-A, NF-κB, and β-catenin |
CTGF, CYR61, EGR1, FOS, JUN, and TGFβ1 |
Structural adaptation of cells, autocrine/paracrine signals, tissue remodeling, and proliferation or pathological responses |
Signal transmission from the cytoskeleton to the nucleus
Signal transmission from the cytoskeleton to the nucleus represents a critical physical phase of mechanotransduction, in which externally applied or internally generated mechanical forces are propagated across cellular structures to directly deform the nucleus and its chromatin architecture [16,91]. Rather than acting solely through biochemical signaling cascades, this process relies on a continuous mechanical linkage that enables force transfer from the plasma membrane to the nuclear interior, thereby influencing nuclear mechanics and chromatin accessibility in response to changes in the mechanical microenvironment [92].
Central to this force transmission is the Linker of Nucleoskeleton and Cytoskeleton (LINC) complex, a multiprotein assembly that mechanically couples cytoskeletal filaments to the nuclear lamina [93]. The LINC complex is composed of nesprin proteins located at the outer nuclear membrane, which bind actin filaments, microtubules, or intermediate filaments, and SUN domain proteins embedded in the inner nuclear membrane that connect to lamin A/C [94]. Together, these components form a load-bearing molecular bridge that allows mechanical forces generated at adhesion sites or within the cytoskeleton to be transmitted directly to the nucleus and chromatin [95].
When cells experience mechanical cues such as substrate stiffness, tensile strain, or shear stress, cytoskeletal remodeling alters the magnitude and direction of forces conveyed through the LINC complex [96]. These forces induce nuclear deformation, manifested as changes in nuclear shape, volume, and stiffness, which are largely determined by the composition and organization of the nuclear lamina [97]. Lamin A/C, in particular, functions as a key mechanical determinant of nuclear rigidity, with increased mechanical loading promoting lamin A/C expression and nuclear stiffening, thereby enhancing resistance to deformation and preserving genomic stability [98-100].
Conversely, disruption of LINC complex components or alterations in lamin organization weaken nuclear mechanical integrity, resulting in increased nuclear deformability and aberrant force dissipation [101]. Such mechanical imbalance affects the spatial organization of chromatin and can lead to altered accessibility of genomic regions, especially those located near the nuclear periphery [99]. These observations indicate that nuclear mechanics act as a mechanical filter, modulating how external forces are translated into structural changes within the genome.
Mechanical deformation of the nucleus also influences chromatin accessibility through mechano-epigenetic mechanisms [102]. Physical strain on the nucleus can reorganize chromatin compaction states, affecting histone modifications such as H3K9 acetylation and H3K27 trimethylation, which are associated with permissive or repressive chromatin configurations, respectively [103]. Importantly, these changes can occur independently of classical signaling intermediates, highlighting that direct mechanical coupling between the cytoskeleton and chromatin constitutes a distinct regulatory layer in mechanotransduction.
Collectively, the cytoskeleton–LINC–nucleus axis establishes a bidirectional mechanical continuum, in which forces transmitted from the extracellular matrix reshape nuclear architecture, while nuclear stiffness feeds back to influence cytoskeletal tension distribution [94,110]. This mechanically integrated system enables cells to adapt structurally to physical stress and maintain tissue integrity. Failure of this force transmission pathway, due to defects in LINC components or nuclear lamins, is associated with a range of mechanopathologies, including cancer, laminopathies, and connective tissue disorders characterized by impaired nuclear mechanoadaptation [111].
Transcription factors and genes responsive to stress
Transcription factors constitute the regulatory layer of mechanotransduction, translating mechanically induced structural cues into specific gene expression programs that govern cell fate, function, and tissue homeostasis [112,113]. Once mechanical forces have been transmitted to the nucleus, cells respond by selectively activating or suppressing mechanosensitive transcription factors, enabling adaptive transcriptional outputs in response to variations in cytoskeletal tension, substrate stiffness, or fluid shear stress [114].
Among these factors, Yes-associated protein (YAP) and transcriptional coactivator with PDZ-binding motif (TAZ) act as central transcriptional effectors of mechanical signaling [115]. Under conditions of high mechanical load or rigid substrates, inhibition of the Hippo kinase cascade (LATS1/2) permits YAP/TAZ dephosphorylation and nuclear accumulation, where they associate with TEAD transcription factors to drive the expression of genes such as CTGF, CYR61, and ANKRD1 [116,117]. These target genes promote cell proliferation, migration, and extracellular matrix remodeling, supporting tissue growth and mechanical adaptation. In contrast, low mechanical stress or compliant matrices maintain YAP/TAZ in an inactive cytoplasmic state, thereby limiting mechanosensitive gene expression and restraining excessive growth responses [118].
Myocardin-related transcription factor A (MRTF-A) represents another key mechanosensitive regulator whose activity is tightly coupled to actin dynamics [119]. Mechanical stimuli that promote actin polymerization reduce the pool of cytoplasmic G-actin, releasing MRTF-A and enabling its translocation to the nucleus [109]. There, MRTF-A cooperates with Serum Response Factor (SRF) to induce the transcription of cytoskeletal and contractile genes, including ACTA2, VCL, and MYH9, reinforcing actomyosin tension and cellular structural integrity [120,121]. This feed-forward mechanism links mechanical force sensing directly to transcriptional programs that stabilize cellular mechanics.
In addition to YAP/TAZ and MRTF-A, other transcription factors contribute to context-dependent mechanotranscriptional responses. Nuclear factor kappa B (NF-κB) is activated by mechanical deformation and shear stress, particularly in vascular cells, driving the expression of pro-inflammatory genes such as ICAM1, VCAM1, and IL6 [122,123]. β-catenin, traditionally associated with Wnt signaling, can also be mechanically activated to promote genes involved in proliferation and tissue repair [124]. Conversely, Kruppel-like factor 2 (KLF2) functions as a protective mechanosensitive transcription factor in endothelial cells, where shear stress–induced activation suppresses inflammatory and thrombogenic gene expression, thereby preserving vascular homeostasis [125].
The transcriptional outcomes of mechanotransduction reflect the integrated activity of these mechanosensitive regulators. Mechanical stimulation commonly induces immediate-early and remodeling-associated genes, including CTGF, CYR61, EGR1, FOS, JUN, and TGFβ1, which coordinate cytoskeletal adaptation, extracellular matrix remodeling, and paracrine signaling within tissues [126,127]. In parallel, mechanical cues can modulate the expression of epigenetic regulators such as histone acetyltransferases (p300/CBP) and DNA methyltransferases (DNMT1), suggesting that sustained mechanical environments can imprint longer-term transcriptional memory through chromatin regulation [128].
Importantly, the balance among mechanosensitive transcription factors determines physiological versus pathological outcomes. Coordinated activation of YAP/TAZ, MRTF-A, and NF-κB favors proliferative, fibrotic, or inflammatory gene programs, as observed in cancer progression and tissue fibrosis [130]. In contrast, dominance of protective pathways, such as KLF2 signaling accompanied by restrained YAP/TAZ activity, supports anti-inflammatory responses and tissue homeostasis [131,132]. Thus, transcriptional mechanotransduction functions as a decision-making layer that integrates mechanical inputs to drive either adaptive remodeling or disease-associated gene expression programs. As shown in Figure 3, mechanical cues from the ECM travel through the cytoskeleton and the LINC complex to remodel chromatin and reprogram gene expression.
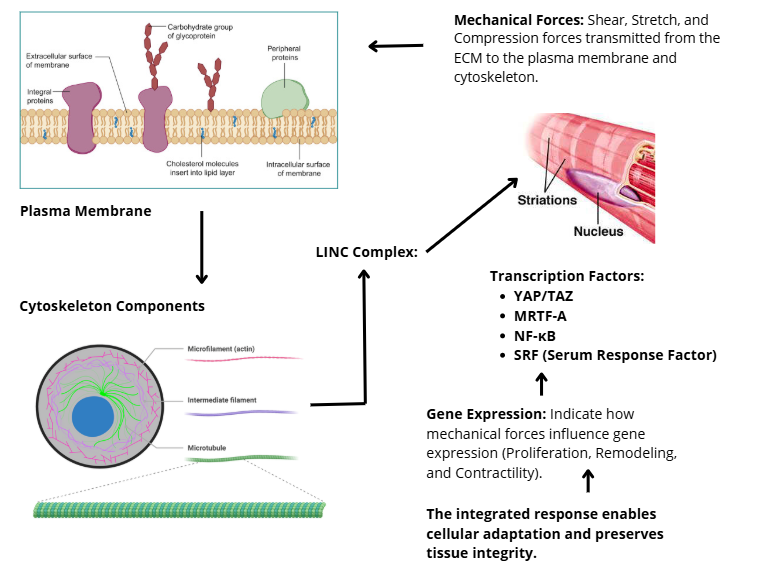
Figure 3 Mechanotransduction from the cytoskeleton to the nucleus via the LINC complex.
Mechanotransduction in physiology and pathophysiology
Mechanotransduction plays a crucial role in maintaining the balance between cellular structure and function under physiological conditions, while also contributing to disease when mechanosensitive pathways are disrupted. Understanding mechanotransduction in the context of physiology and pathophysiology provides insight into how mechanical forces influence cellular adaptation or dysfunction. Table 3 summarizes the role of mechanotransduction under normal and pathological conditions in various tissues.
Table 3 The role of mechanotransduction in cellular physiology and pathophysiology.
Condition |
System/Network |
Mechanotransduction mechanism |
Signaling pathways/ Transcription factors |
Mobile impact/response |
Normal physiology |
Stem cells |
Mechanotransduction mediates differentiation based on matrix stiffness |
YAP/TAZ, RhoA/ROCK, and integrin–FAK |
Osteogenic, neurogenic, or adipogenic differentiation and specialization of cells according to the mechanical environment |
Skeletal muscle |
Contraction and stretching strains trigger activation of mechanoreceptors |
PI3K–Akt–mTOR and MAPK/ERK |
Muscle hypertrophy, mitochondrial biogenesis, and increased protein synthesis and metabolic capacity |
|
Vascular system (endothelium) |
Hemodynamic force sensor (shear stress) |
KLF2 and eNOS |
NO production, vasodilation, blood pressure homeostasis, and vascular integrity |
|
Pathological physiology |
Tumor/cancer microenvironment |
Increased matrix stiffness and interstitial pressure |
YAP/TAZ, β-catenin, RhoA/ROCK, and integrin–FAK |
Abnormal proliferation, cell migration and invasion, epithelial–mesenchymal transition (EMT), and metastasis |
Fibrosis |
Increased tissue stiffness and fibroblast activation |
TGF-β/SMAD, YAP/TAZ, and α-SMA |
Transformation of fibroblasts into myofibroblasts, excessive collagen production, fibrosis progression, and inhibition of tissue resolution |
|
Cardiovascular system (endothelium + smooth muscle) |
Turbulent blood flow and stiff arterial walls |
KLF2, eNOS, and NF-κB |
Decreased NO, increased oxidative stress, proinflammatory gene expression (VCAM1 and ICAM1), smooth muscle proliferation, vascular remodeling, hypertension, and atherosclerosis |
Normal physiology
Mechanotransduction plays an essential role in a variety of normal physiological processes by enabling cells to detect and respond to changes in mechanical forces in their environment [5]. By integrating mechanical and biochemical signals, mechanotransduction dynamically regulates cell differentiation, tissue function, and organ homeostasis [133]. This process underpins physiological adaptations in the musculoskeletal, cardiovascular, and nervous systems, while maintaining a balance between cellular structure and function [134].
Stem cell differentiation is strongly influenced by the mechanical characteristics of the extracellular environment, including matrix stiffness and the stress experienced by the cells [135]. Stiff matrices tend to direct cells toward osteogenic differentiation, while softer matrices promote neurogenic or adipogenic differentiation [136]. Mechanosensitive pathways such as YAP/TAZ, RhoA/ROCK, and integrin–FAK play key roles in determining cell fate by regulating tissue-specific gene expression [137]. Thus, mechanotransduction provides a link between physical stimuli and transcriptional regulation that determines cell specialization [138].
In skeletal muscle tissue, mechanical forces generated during repeated contraction and stretching activate mechanoreceptors and signaling pathways such as PI3K–Akt–mTOR and MAPK/ERK, which control hypertrophy and mitochondrial biogenesis [139]. Activation of these pathways increases protein synthesis and metabolic capacity, allowing muscles to adapt to increased workload [140]. Furthermore, shear and tension forces on the plasma membrane influence the expression of contractile genes and cytoskeletal proteins, strengthening muscle structure to maintain optimal function [141]. Thus, mechanotransduction is the molecular basis of muscle functional adaptation to physical activity [19].
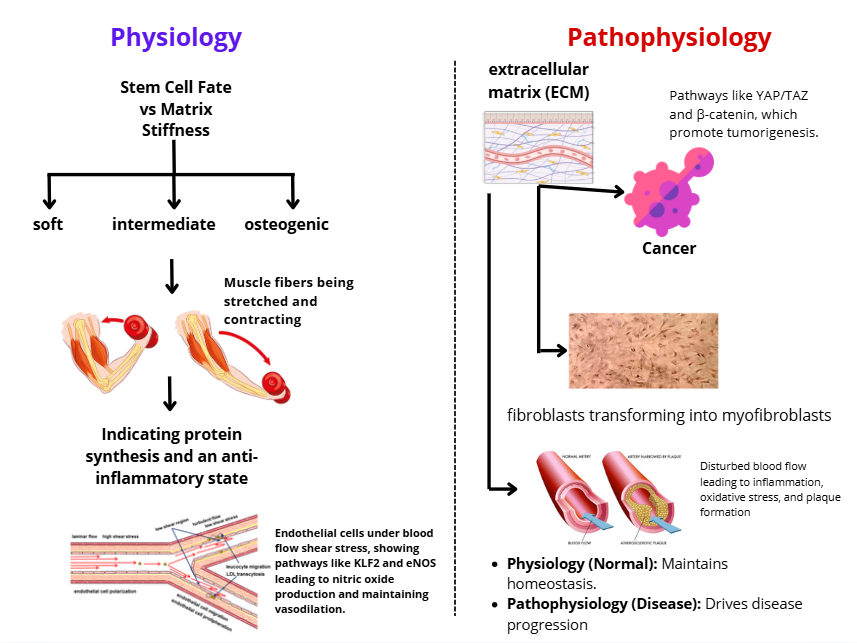
Figure 4 Mechanotransduction across physiology and pathophysiology.
In the vascular system, endothelial cells serve as primary sensors for hemodynamic forces, such as shear stress from blood flow [142]. Endothelial mechanotransduction regulates the activation of the krüppel-like factor 2–endothelial nitric oxide synthase (KLF2–eNOS) pathway, which triggers nitric oxide (NO) production to maintain vasodilation and lower blood pressure [143]. Conversely, endothelial mechanotransduction dysfunction can lead to increased expression of pro-inflammatory and procoagulant genes, thereby increasing the risk of vascular disorders [144]. Thus, mechanotransduction plays a crucial role in maintaining blood pressure homeostasis and vascular integrity through adaptive responses to mechanical forces [145].
Pathological physiology
Dysregulation of mechanotransduction plays a crucial role in the development of various diseases, including cancer, fibrosis, and cardiovascular disorders [5]. An imbalance between mechanical forces, tissue stiffness, and cellular responses induces profound alterations in cell behavior, gene expression, and cell–matrix interactions, ultimately driving chronic pathological remodeling [146]. Beyond its pathogenic significance, growing evidence indicates that mechanotransduction represents a promising therapeutic entry point, enabling the development of mechanobiology-driven interventions that target force-sensitive signaling pathways and cellular mechanosensors [147].
In the tumor microenvironment, increased extracellular matrix stiffness and elevated interstitial pressure actively promote malignant transformation rather than merely reflecting tumor progression [148]. Aberrant activation of mechanosensitive pathways such as YAP/TAZ, β-catenin, and RhoA/ROCK enhances cancer cell proliferation, invasion, and metastatic dissemination [149]. Importantly, this mechanistic insight has led to the emergence of YAP/TAZ-targeted therapeutic strategies, including small-molecule inhibitors, transcriptional disruptors, and upstream Hippo pathway modulators, which have shown potential in suppressing tumor growth and overcoming therapy resistance in preclinical cancer models [150,151]. Furthermore, targeting integrin–FAK signaling and cytoskeletal tension has been proposed as a complementary strategy to disrupt mechano-driven epithelial–mesenchymal transition (EMT) and tumor invasiveness.
Fibrosis is another pathological condition in which aberrant mechanotransduction plays a central role. Excessive extracellular matrix deposition increases tissue stiffness, which in turn activates fibroblasts and sustains their differentiation into α-smooth muscle actin (α-SMA)-positive myofibroblasts [152,153]. Persistent activation of mechanosensitive pathways, particularly TGF-β/SMAD and YAP/TAZ signaling, reinforces a feed-forward loop between mechanical stiffening and fibroblast activation, accelerating fibrotic progression [154]. These findings have stimulated the development of anti-fibrotic mechanotherapies, including YAP/TAZ inhibitors, integrin antagonists, and mechanoresponsive biomaterials designed to soften the cellular microenvironment and promote fibrosis resolution [155]. Such approaches highlight the therapeutic potential of modulating mechanical cues rather than targeting biochemical pathways alone.
In the cardiovascular system, endothelial mechanotransduction is critically influenced by hemodynamic forces, particularly shear stress generated by blood flow [156]. Disturbed or turbulent flow suppresses protective mechanosensitive pathways such as KLF2–eNOS signaling, leading to reduced nitric oxide (NO) bioavailability, increased oxidative stress, and activation of pro-inflammatory transcription factors including NF-κB [157,158]. These molecular events promote leukocyte adhesion, vascular inflammation, and atherosclerotic plaque formation. Translationally, organ-on-chip and vessel-on-chip platforms have emerged as powerful tools to model pathological flow patterns and screen drugs that restore endothelial mechanosensitivity, enabling more physiologically relevant evaluation of cardiovascular therapeutics [159].
Collectively, these insights underscore the growing relevance of mechanopharmacology, an emerging field that integrates mechanical cues into drug discovery and therapeutic design. Mechanosensitive scaffolds in regenerative medicine, force-responsive drug delivery systems, and microengineered organ-on-chip platforms are increasingly being used to modulate cell fate, enhance tissue regeneration, and improve predictive accuracy in drug screening. Future research integrating mechanotransduction with multi-omics profiling, advanced biomaterials, and microphysiological systems will be essential for translating mechanobiological principles into effective clinical interventions across cancer, fibrosis, and cardiovascular disease.
Experimental technologies and approaches
Advances in experimental technology now allow for a more detailed understanding of mechanotransduction mechanisms at the molecular, cellular, and tissue levels [160]. Multidisciplinary approaches combining cell biology, biophysics, and microengineering enable quantitative analysis of mechanical forces and cellular responses to them [161]. A variety of experimental techniques are used to visualize, manipulate, and quantify the forces that influence cell behavior and regulate gene expression [162].
Atomic force microscopy (AFM) is one of the most sensitive methods for assessing plasma membrane stiffness, adhesion forces, and the elastic response of cells to external forces [163]. Using force spectroscopy, AFM can map the topography and force distribution at the nanometer scale, providing valuable information about the local stiffness and dynamics of the cytoskeleton [164].
Additionally, micropatterning techniques are used to regulate adhesion geometry and cell orientation on specific substrates, allowing for a more focused analysis of how cell shape and internal tension influence the activation of mechanosensitive pathways such as YAP/TAZ and RhoA/ROCK [165].
On the other hand, traction force microscopy (TFM) allows the measurement of traction forces generated by cells on fluorescent elastic substrates [166]. This approach provides in-depth insights into the internal force distribution and stress transmission through focal adhesion complexes, making it an important method for studying cell–matrix interactions in real time [167].
Three-dimensional (3D) culture models have transformed the approach to mechanotransduction studies by providing an environment that more closely mimics physiological conditions than traditional two-dimensional culture [168]. In these systems, cells interact spatially with the extracellular matrix and receive forces from multiple directions, allowing for more accurate analysis of mechanical responses and cell differentiation [169].
Additionally, organ-on-chip technology based on microfluidic platforms enables the simulation of physiological forces, including shear stress, cyclic strain, and hydrostatic pressure [170]. These systems support mechanotransduction studies in complex tissues such as vascular endothelium, alveolar epithelium, or myocardium, while also enabling pharmacological evaluation to assess the response to therapeutic interventions that utilize mechanics [171].
RNA sequencing (RNA-seq) approaches have become a key technique for identifying genes and molecular pathways involved in mechanosensitive responses [172]. Through comprehensive transcriptomic analysis, RNA-seq can detect changes in gene expression resulting from variations in mechanical force, substrate stiffness, or cell deformation [173]. Integrating RNA-seq data with epigenomics and proteomics allows mapping complex regulatory networks between mechanosensitive transcription factors, such as YAP/TAZ, MRTF-A, and KLF2, and target genes that regulate cell adaptation to the mechanical environment [174]. This approach provides systemic insights into gene expression dynamics and helps identify mechanogenomic signatures characteristic of both physiological and pathological conditions [175].
Challenges and future research directions
Although understanding of the fundamental mechanisms of mechanotransduction has advanced significantly, several unresolved scientific questions remain that limit the translation of this knowledge into broader biological and clinical applications [176]. The inherent complexity of mechanotransduction—arising from the intertwined nature of mechanical forces, biochemical signaling, and gene regulation—necessitates a systemic, quantitative, and interdisciplinary research framework [177].
One major open question concerns how mechanical signals can be reliably disentangled from biochemical cues within complex cellular systems [147]. While multi-omics approaches integrating transcriptomics, epigenomics, and proteomics have enabled large-scale mapping of force-responsive gene networks, it remains unclear how mechanical inputs independently shape gene regulatory programs across different cell types and tissue contexts [178]. Moreover, the lack of standardized experimental pipelines for mechanogenomic data acquisition and integration hampers cross-study comparability and reproducibility, particularly in pathological models [179].
Another unresolved challenge lies in predicting cellular fate decisions under dynamic and heterogeneous mechanical environments. Computational modeling and biomechanical simulations have shown promise in linking matrix stiffness, cytoskeletal tension, and molecular signaling to outcomes such as differentiation or apoptosis [180]. However, current models often oversimplify biological variability, raising the question of how accurately in silico predictions can reflect in vivo mechanotransduction dynamics [134]. The integration of machine learning and Artificial Intelligence (AI) into mechanobiology offers new opportunities to address this limitation, yet the interpretability and biological generalizability of AI-driven models remain significant concerns [181].
From a translational perspective, a critical open question is how mechanotransduction principles can be effectively harnessed for therapeutic intervention [147]. Although mechanosensitive biomaterials and stiffness-tunable scaffolds have demonstrated potential in tissue engineering and regenerative medicine, their long-term biological stability and clinical scalability require further validation [182]. Similarly, while anti-fibrotic strategies targeting YAP/TAZ or integrin–FAK signaling pathways show therapeutic promise, the risk of disrupting physiological mechanoadaptive functions remains insufficiently understood [183].
Finally, the emergence of organ-on-chip platforms raises important questions regarding the extent to which these systems can faithfully recapitulate multiscale mechanical cues present in native tissues. While organ-on-chip and mechanopharmacology approaches provide powerful tools for drug screening and disease modeling, their standardization, validation, and integration with omics-based readouts remain ongoing challenges [184]. Addressing these open questions will be essential for advancing mechanotransduction research from descriptive frameworks toward predictive, translational, and clinically actionable paradigms.
Conclusions
Mechanotransduction constitutes a fundamental regulatory axis through which mechanical cues are integrated with biochemical signaling to control gene expression and cellular fate. Rather than functioning solely as a passive force-transmission system, the cytoskeleton–adhesion–nucleus continuum actively shapes transcriptional programs via mechanosensitive regulators such as YAP/TAZ and MRTF-A, thereby determining whether mechanical stimuli elicit adaptive or pathological responses. Emerging evidence highlights that context-dependent factors—including matrix dimensionality, cell type, and force magnitude—critically influence mechanotransduction outcomes, underscoring the need for integrative and comparative mechanobiological models. Future advances will depend on combining three-dimensional culture systems, organ-on-chip platforms, and multi-omics approaches to disentangle mechanical and biochemical signaling with higher precision. Such efforts are expected to accelerate the translation of mechanotransduction principles into mechanotherapy, mechanoresponsive biomaterials, and targeted interventions for fibrosis, cancer, and cardiovascular diseases.
Acknowledgements
The authors would like to express their sincere gratitude to the National Research and Innovation Agency (BRIN) and the Indonesia Endowment Fund for Education (LPDP) for financial support through the Research and Innovation for Advanced Indonesia (RIIM) Program – Competitive Wave 7, as stipulated in the Decree of the Deputy for Research and Innovation Facilitation of BRIN No. 61/II.7/HK/2024. The authors also acknowledge material and immaterial support from the Research Organization for Agriculture and Food (ORPP BRIN) and the valuable assistance of the Research Center for Mineral Technology, BRIN, in the preparation and completion of this manuscript. Collaborative contributions from Institut Teknologi Sumatera (ITERA) are gratefully appreciated.
Declaration of Generative AI in Scientific Writing
The authors declare that no generative AI tools were used in the writing or preparation of this manuscript.
CRediT Author Statement
Aswin Rafif Khairullah: Conceptualization, Literature review, and Writing – original draft. Abdul Hadi Furqoni: Literature review, Data curation, and Writing – original draft. Analinasari Analinasari: Literature review, Data curation, and Writing – original draft. Arif Nur Muhammad Ansori: Reference management, Data curation, and Validation. Bantari Wisynu Kusuma Wardhani: Conceptualization, Supervision, Funding acquisition, and Writing – review and editing. Bima Putra Pratama: Writing – review and editing, Validation, and Supervision. Dea Anita Ariani Kurniasih: Manuscript preparation, Critical review, and Validation. Eny Martindah: Manuscript preparation, Critical review, and Validation. Fadhila Utari: Manuscript preparation, Critical review, and Validation. Ilma Fauziah Ma’ruf: Methodological review, Critical revision, and Validation. Imam Mustofa: Writing – review and editing, Supervision, and Validation. Mohammad Sukmanadi: Reference management, Resources, and Validation. Riza Zainuddin Ahmad: Manuscript preparation, Critical review, and Validation. Saifur Rehman: Reference management, Resources, and Validation. Siti Hamidatul Aliyah: Project administration, Supervision, Writing – review and editing, and Validation. Wita Yulianti: Writing – review and editing, Visualization, and Validation.
References
P Elblová, M Lunova, A Dejneka, M Jirsa and O Lunov. Impact of mechanical cues on key cell functions and cell-nanoparticle interactions. Discover Nano 2024; 19(1), 106.
Y Zeng, X Cui, H Li, Y Wang, M Cheng and X Zhang. Extracellular vesicles originating from the mechanical microenvironment in the pathogenesis and applications for cardiovascular diseases. Regenerative Therapy 2024; 26(1), 1069-1977.
P Nitti, A Narayanan, R Pellegrino, S Villani, M Madaghiele and C Demitri. Cell-Tissue Interaction: The biomimetic approach to design tissue engineered biomaterials. Bioengineering (Basel) 2023; 10(10), 1122.
J Su, Y Song, Z Zhu, X Huang, J Fan, J Qiao and F Mao. Cell-cell communication: New insights and clinical implications. Signal Transduction and Targeted Therapy 2024; 9(1), 196.
X Di, X Gao, L Peng, J Ai, X Jin, S Qi, H Li, K Wang and D Luo. Cellular mechanotransduction in health and diseases: From molecular mechanism to therapeutic targets. Signal Transduction and Targeted Therapy 2023; 8(1), 282.
I Muhamed, F Chowdhury and V Maruthamuthu. Biophysical tools to study cellular mechanotransduction. Bioengineering 2017; 4(1), 12.
C Cavallini, E Olivi, R Tassinari and C Ventura. Mechanotransduction, cellular biophotonic activity, and signaling patterns for tissue regeneration. Journal of Biological Chemistry 2024; 300(11), 107847.
A Rojas, S Dahlen, F Zhang and S Liu. The cytoskeletal structure in cardiomyocyte maturation and proliferation. Cells 2025; 14(19), 1494.
J Na, Z Yang, Q Shi, C Li, Y Liu, Y Song, X Li, L Zheng and Y Fan. Extracellular matrix stiffness as an energy metabolism regulator drives osteogenic differentiation in mesenchymal stem cells. Bioactive Materials 2024; 35(1), 549-563.
Q Zheng, Y Zou, P Teng, Z Chen, Y Wu, X Dai, X Li, Z Hu, S Wu, Y Xu, W Zou, H Song and L Ma. Mechanosensitive channel PIEZO1 senses shear force to induce KLF2/4 expression via CaMKII/MEKK3/ERK5 axis in endothelial cells. Cells 2022; 11(14), 2191.
Y Wang, J Meng, J Zhang, L Tian, W Wei, X Tang, Q Zhang, D Ding, X Wang, Z Guo and Y He. Cell biomechanics on muscle atrophy: From intricate mechanisms to therapeutic frontiers. Annals of Medicine 2025; 57(1), 2540598.
R Cao, H Tian, Y Tian and X Fu. A Hierarchical Mechanotransduction System: From Macro to Micro. Advanced Science (Weinh) 2024; 11(11), e2302327.
R Li and W Huang. Yes-associated protein and transcriptional coactivator with PDZ-binding motif in cardiovascular diseases. International Journal of Molecular Sciences 2023; 24(2), 1666.
M Liaghat, S Ferdousmakan, SH Mortazavi, S Yahyazadeh, A Irani, S Banihashemi, FSS Asl, A Akbari, F Farzam, F Aziziyan, M Bakhtiyari, MJ Arghavani, H Zalpoor and M Nabi-Afjadi. The impact of epithelial-mesenchymal transition (EMT) induced by metabolic processes and intracellular signaling pathways on chemo-resistance, metastasis, and recurrence in solid tumors. Cell Communication and Signaling 2024; 22(1), 575.
S Jebari-Benslaiman, U Galicia-García, A Larrea-Sebal, JR Olaetxea, I Alloza, K Vandenbroeck, A Benito-Vicente and C Martín. Pathophysiology of atherosclerosis. International Journal of Molecular Sciences 2022; 23(6), 3346.
F Martino, AR Perestrelo, V Vinarský, S Pagliari and G Forte. Cellular Mechanotransduction: From Tension to Function. Frontiers in Physiology 2018; 9(1), 824.
GB Bais and J Giudice. The bidirectional interplay between RNA processing and mechanotransduction. Cell Reports 2025; 44(9), 116175.
IP Uray and K Uray. Mechanotransduction at the plasma Membrane-cytoskeleton Interface. International Journal of Molecular Sciences 2021; 22(21), 11566.
U Mukhopadhyay, T Mandal, M Chakraborty and B Sinha. The Plasma Membrane and Mechanoregulation in Cells. ACS Omega 2024; 9(20), 21780-21797.
P Wang, J Li and Q Wei. Understanding the interplay between cell force and cell adhesion processes. Engineered Regeneration 2023; 4(3), 277-288.
K Katoh. Integrin and Its associated proteins as a mediator for Mechano-signal transduction. Biomolecules 2025; 15(2), 166.
K Katoh. Signal transduction mechanisms of focal adhesions: Src and FAK-Mediated cell response. Frontiers in Bioscience 2024; 29(11), 392.
RM Mège and N Ishiyama. Integration of cadherin adhesion and cytoskeleton at adherens junctions. Cold Spring Harbor Perspectives in Biology 2017; 9(5), a028738.
WH Lin, LM Cooper and PZ Anastasiadis. Cadherins and catenins in cancer: Connecting cancer pathways and tumor microenvironment. Frontiers in Cell and Developmental Biology 2023; 11(1), 1137013.
Z Chen, C Du, S Liu, J Liu, Y Yang, L Dong, W Zhao, W Huang and Y Lei. Progress in biomaterials inspired by the extracellular matrix. Giant 2024; 19(1), 100323.
A Saraswathibhatla, D Indana and O Chaudhuri. Cell-extracellular matrix mechanotransduction in 3D. Nature Reviews Molecular Cell Biology 2023; 24(7), 495-516.
W Xie, X Wei, H Kang, H Jiang, Z Chu, Y Lin, Y Hou and Q Wei. Static and Dynamic: Evolving Biomaterial Mechanical Properties to Control Cellular Mechanotransduction. Advanced Science 2023; 10(9), e2204594.
X Cai, KC Wang and Z Meng. Mechanoregulation of YAP and TAZ in cellular homeostasis and disease progression. Frontiers in Cell and Developmental Biology 2021; 9(1), 673599.
M Zhang and B Zhang. Extracellular matrix stiffness: Mechanisms in tumor progression and therapeutic potential in cancer. Experimental Hematology & Oncology 2025; 14(1), 54.
T Zhao, Y Huang, J Zhu, Y Qin, H Wu, J Yu, Q Zhai, S Li, X Qin, D Wang, T Li and Y Liu. Extracellular matrix signaling cues: Biological functions, diseases, and therapeutic targets. MedComm 2025; 6(8), e70281.
T Hohmann and F Dehghani. The Cytoskeleton-A complex interacting meshwork. Cells 2019; 8(4), 362.
R Sanghvi-Shah and GF Weber. Intermediate filaments at the junction of mechanotransduction, migration, and development. Frontiers in Cell and Developmental Biology 2017; 5(1), 81.
T Svitkina. The actin cytoskeleton and actin-based motility. Cold Spring Harbor Perspectives in Biology 2018; 10(1), a018267.
E Infante and S Etienne-Manneville. Intermediate filaments: Integration of cell mechanical properties during migration. Frontiers in Cell and Developmental Biology 2022; 10(1), 951816.
J Salvador and ML Iruela-Arispe. Nuclear mechanosensation and mechanotransduction in vascular cells. Frontiers in Cell and Developmental Biology 2022; 10(1), 905927.
E Momotyuk, N Ebrahim, K Shakirova and E Dashinimaev. Role of the cytoskeleton in cellular reprogramming: Effects of biophysical and biochemical factors. Frontiers in Molecular Biosciences 2025; 12(1), 1538806.
EK Pillai and K Franze. Mechanics in the nervous system: From development to disease. Neuron 2024; 112(3), 342-361.
EK Paluch, CM Nelson, N Biais, B Fabry, J Moeller, BL Pruitt, C Wollnik, G Kudryasheva, F Rehfeldt and W Federle. Mechanotransduction: Use the force(s). BMC Biology 2015; 13(1), 47.
D Mohammed, I Hamid, B Vanhollebeke and M Martin. Microtubules and mechanosensing: Key players in endothelial responses to mechanical stimuli. Cellular and Molecular Life Sciences 2025; 82(1), 317.
K Legerstee and AB Houtsmuller. A layered view on focal adhesions. Biology 2021; 10(11), 1189.
H Wang, R Said, C Nguyen-Vigouroux, V Henriot, P Gebhardt, J Pernier, R Grosse and CL Clainche. Talin and vinculin combine their activities to trigger actin assembly. Nature Communications 2024; 15(1), 9497.
T Izard and DT Brown. Mechanisms and functions of vinculin interactions with phospholipids at cell adhesion sites. Journal of Biological Chemistry 2016; 291(6), 2548-2555.
P Ridone, M Vassalli and B Martinac. Piezo1 mechanosensitive channels: What are they and why are they important. Biophysical Reviews 2019; 11(5), 795-805.
G Yuan, Z Xiong, X Ke, G Wang, X Liu and Z Li. Exploring the multifactorial regulation of PIEZO1 in chondrocytes: Mechanisms and implications. International Journal of Medical Sciences 2025; 22(13), 3393-3411.
Q Zhang, X Han, J Chen, X Xie, J Xu, Y Zhao, J Shen, L Hu, P Xu, H Song, L Zhang, B Zhao, YJ Wang and Z Xia. Yes-associated protein (YAP) and transcriptional coactivator with PDZ-binding motif (TAZ) mediate cell density-dependent proinflammatory responses. Journal of Biological Chemistry 2018; 293(47), 18071-18085.
Y Wei, VLZ Hui, Y Chen, R Han, X Han and Y Guo. YAP/TAZ: Molecular pathway and disease therapy. MedComm 2023; 4(4), e340.
Y Li, J Wang and W Zhong. Regulation and mechanism of YAP/TAZ in the mechanical microenvironment of stem cells (review). Molecular Medicine Reports 2021; 24(1), 506.
M Ozu, L Galizia, JJ Alvear-Arias, M Fernández, A Caviglia, R Zimmermann, F Guastaferri, N Espinoza-Muñoz, M Sutka, L Sigaut, LI Pietrasanta, C González, G Amodeo and JA Garate. Mechanosensitive aquaporins. Biophysical Reviews 2023; 15(4), 497-513.
S Vajda, D Beglov, AE Wakefield, M Egbert and A Whitty. Cryptic binding sites on proteins: Definition, detection, and druggability. Current Opinion in Chemical Biology 2018; 44, 1-8.
Y Wang, M Yao, KB Baker, RE Gough, S Le, BT Goult and J Yan. Force-dependent interactions between talin and full-length vinculin. Journal of the American Chemical Society 2021; 143(36), 14726-14737.
A Buchwalter. Intermediate, but not average: The unusual lives of the nuclear lamin proteins. Current Opinion in Cell Biology 2023; 84(1), 102220.
J Jo, SA Nansa and DH Kim. Molecular regulators of cellular mechanoadaptation at Cell-material interfaces. Frontiers in Bioengineering and Biotechnology 2020; 8(1), 608569.
G Guan, RD Cannon, DE Coates and L Mei. Effect of the Rho-Kinase/ROCK Signaling pathway on cytoskeleton components. Genes 2023; 14(2), 272.
C Braicu, M Buse, C Busuioc, R Drula, D Gulei, L Raduly, A Rusu, A Irimie, AG Atanasov, O Slaby, C Ionescu and I Berindan-Neagoe. A comprehensive review on MAPK: A promising therapeutic target in cancer. Cancers 2019; 11(10), 1618.
ST Eblen. Extracellular-regulated kinases: Signaling from Ras to ERK substrates to control biological outcomes. Advances in Cancer Research 2018; 138(1), 99-142.
T Pungsrinont, J Kallenbach and A Baniahmad. Role of PI3K-AKT-mTOR pathway as a pro-survival signaling and resistance-mediating mechanism to therapy of prostate cancer. International Journal of Molecular Sciences 2021; 22(20), 11088.
I Tortorella, C Argentati, C Emiliani, F Morena and S Martino. Biochemical pathways of cellular mechanosensing/mechanotransduction and their role in neurodegenerative diseases pathogenesis. Cells 2022; 11(19), 3093.
DA Fletcher and RD Mullins. Cell mechanics and the cytoskeleton. Nature 2010; 463(7280), 485-492.
G Yuan, H Gao and T Yang. Exploring the role of the plant actin cytoskeleton: From signaling to cellular functions. International Journal of Molecular Sciences 2023; 24(20), 15480.
LS Mosby, N Hundt, G Young, A Fineberg, M Polin, S Mayor, P Kukura and DV Köster. Myosin II filament dynamics in actin networks revealed with interferometric scattering microscopy. Biophysical Journal 2020; 118(8), 1946-1957.
H Katsuta, M Sokabe and H Hirata. From stress fiber to focal adhesion: A role of actin crosslinkers in force transmission. Frontiers in Cell and Developmental Biology 2024; 12(1), 1444827.
K Bhadriraju, M Yang, SA Ruiz, D Pirone, J Tan and CS Chen. Activation of ROCK by RhoA is regulated by cell adhesion, shape, and cytoskeletal tension. Experimental Cell Research 2007; 313(16), 3616-3623.
K Okura, T Matsumoto, A Narita and H Tatsumi. Mechanical stress decreases the amplitude of twisting and bending fluctuations of actin filaments. Journal of Molecular Biology 2023; 435(22), 168295.
BV Fearing, L Jing, MN Barcellona, SE Witte, JM Buchowski, LP Zebala, MP Kelly, S Luhmann, MC Gupta, A Pathak and LA Setton. Mechanosensitive transcriptional coactivators MRTF-A and YAP/TAZ regulate nucleus pulposus cell phenotype through cell shape. The FASEB Journal 2019; 33(12), 14022-14035.
X Sun and GM Alushin. Cellular force-sensing through actin filaments. The FEBS Journal 2023; 290(10), 2576-2589.
A Pacheco and G Gallo. Actin filament-microtubule interactions in axon initiation and branching. Brain Research Bulletin 2016; 126, 300-310.
M Ezzo and S Etienne-Manneville. Microtubule-targeting agents: Advances in tubulin binding and small molecule therapy for gliomas and neurodegenerative diseases. International Journal of Molecular Sciences 2025; 26(15), 7652.
HV Goodson and EM Jonasson. Microtubules and microtubule-associated proteins. Cold Spring Harbor Perspectives in Biology 2018; 10(6), a022608.
J Singh, P Sanghavi and R Mallik. Microtubule motor driven interactions of lipid droplets: Specificities and opportunities. Frontiers in Cell and Developmental Biology 2022; 10(1), 893375.
A Rape, WH Guo and YL Wang. Microtubule depolymerization induces traction force increase through two distinct pathways. Journal of Cell Science 2011; 124, 4233-4240.
CS Tong, M Su, H Sun, XL Chua, D Xiong, S Guo, R Raj, NWP Ong, AG Lee, Y Miao and M Wu. Collective dynamics of actin and microtubule and its crosstalk mediated by FHDC1. Frontiers in Cell and Developmental Biology 2024; 11(1), 1261117.
B MacTaggart and A Kashina. Posttranslational modifications of the cytoskeleton. Cytoskeleton 2021; 78(4), 142-173.
H Yamaguchi and J Condeelis. Regulation of the actin cytoskeleton in cancer cell migration and invasion. Biochimica et Biophysica Acta 2007; 1773(5), 642-652.
JP Conboy, II Petitjean, AVD Net and GH Koenderink. How cytoskeletal crosstalk makes cells move: Bridging cell-free and cell studies. Biophysical Reviews 2024; 5(2), 021307.
CT Mierke. Extracellular matrix cues regulate mechanosensing and mechanotransduction of cancer cells. Cells 2024; 13(1), 96.
K Katoh. Effects of mechanical stress on endothelial cells in situ and In Vitro. International Journal of Molecular Sciences 2023; 24(22), 16518.
SC Carim, A Kechad and GRX Hickson. Animal cell cytokinesis: The Rho-dependent actomyosin-anilloseptin contractile ring as a membrane microdomain gathering, compressing, and sorting machine. Frontiers in Cell and Developmental Biology 2020; 8(1), 575226.
F Prieto-Ruiz, E Gómez-Gil, R Martín-García, AJ Pérez-Díaz, J Vicente-Soler, A Franco, T Soto, P Pérez, M Madrid and J Cansado. Myosin II regulatory light chain phosphorylation and formin availability modulate cytokinesis upon changes in carbohydrate metabolism. Elife 2023; 12(1), e83285.
M Hirano and K Hirano. Critical role of Rho proteins in myosin light chain di-phosphorylation during early phase of endothelial barrier disruption. The Journal of Physiological Sciences 2022; 72(1), 32.
OM Rossier, N Gauthier, N Biais, W Vonnegut, MA Fardin, P Avigan, ER Heller, A Mathur, S Ghassemi, MS Koeckert, JC Hone and MP Sheetz. Force generated by actomyosin contraction builds bridges between adhesive contacts. The EMBO Journal 2010; 29(6), 1055-1068.
KH Vining and DJ Mooney. Mechanical forces direct stem cell behaviour in development and regeneration. Nature Reviews Molecular Cell Biology 2017; 18(12), 728-742.
T Bouzid, E Kim, BD Riehl, AM Esfahani, J Rosenbohm, R Yang, B Duan and JY Lim. The LINC complex, mechanotransduction, and mesenchymal stem cell function and fate. Journal of Biological Engineering 2019; 13(1), 68.
X Tan, Y Yan, B Song, S Zhu, Q Mei and K Wu. Focal adhesion kinase: From biological functions to therapeutic strategies. Experimental Hematology & Oncology 2023; 12(1), 83.
EK Duperret, A Dahal and TW Ridky. Focal-adhesion-independent integrin-αv regulation of FAK and c-Myc is necessary for 3D skin formation and tumor invasion. Journal of Cell Science 2015; 128(21), 3997-4013.
K Katoh. FAK-Dependent Cell Motility and Cell Elongation. Cells 2020; 9(1), 192.
N Mosaddeghzadeh and MR Ahmadian. The RHO Family GTPases: Mechanisms of Regulation and Signaling. Cells 2021; 10(7), 1831.
J Hicks-Berthet and X Varelas. Integrin-FAK-CDC42-PP1A signaling gnaws at YAP/TAZ activity to control incisor stem cells. Bioessays 2017; 39(10), 1700116.
JZ Kechagia, J Ivaska and P Roca-Cusachs. Integrins as biomechanical sensors of the microenvironment. Nature Reviews Molecular Cell Biology 2019; 20(8), 457-473.
R Kolasangiani, TC Bidone and MA Schwartz. Integrin conformational dynamics and mechanotransduction. Cells 2022; 11(22), 3584.
P Atherton, B Stutchbury, D Jethwa and C Ballestrem. Mechanosensitive components of integrin adhesions: Role of vinculin. Experimental Cell Research 2016; 343(1), 21-27.
C Oses, MCD Rossi, L Bruno, P Verneri, MC Diaz, B Benítez, A Guberman and V Levi. From the membrane to the nucleus: Mechanical signals and transcription regulation. Biophysical Reviews 2023; 15(4), 671-683.
G Uzer, CT Rubin and J Rubin. Cell mechanosensitivity is enabled by the LINC nuclear complex. Current Molecular Biology Reports 2016; 2(1), 36-47.
RM McGillivary, DA Starr and GWG Luxton. Building and breaking mechanical bridges between the nucleus and cytoskeleton: Regulation of LINC complex assembly and disassembly. Current Opinion in Cell Biology 2023; 85(1), 102260.
M Hieda. Signal transduction across the nuclear envelope: Role of the LINC complex in bidirectional signaling. Cells 2019; 8(2), 124.
BRM Meqbel, M Gomes, A Omer, IE Gallouzi and HF Horn. LINCing senescence and nuclear envelope changes. Cells 2022; 11(11), 1787.
DH Kim and D Wirtz. Cytoskeletal tension induces the polarized architecture of the nucleus. Biomaterials 2015; 48(1), 161-172.
YA Miroshnikova and SA Wickström. Mechanical forces in nuclear organization. Cold Spring Harbor Perspectives in Biology 2022; 14(1), a039685.
TP Lele, RB Dickinson and GG Gundersen. Mechanical principles of nuclear shaping and positioning. Journal of Cell Biology 2018; 217(10), 3330-3342.
M Kloc and J Wosik. mechanical forces, nucleus, chromosomes, and chromatin. Biomolecules 2025; 15(3), 354.
N Dubik and S Mai. Lamin A/C: Function in normal and tumor cells. Cancers 2020; 12(12), 3688.
N Alcorta-Sevillano, I Macías, CI Rodríguez and A Infante. Crucial role of lamin A/C in the migration and differentiation of MSCs in bone. Cells 2020; 9(6), 1330.
Y Song, J Soto and S Li. Mechanical regulation of histone modifications and cell plasticity. Current Opinion in Solid State and Materials Science 2020; 24(6), 100872.
S Ibragić, S Dahija and E Karalija. The good, the bad, and the epigenetic: Stress-induced metabolite regulation and transgenerational effects. Epigenomes 2025; 9(2), 10.
R Melchionna, P Trono, A Tocci and P Nisticò. Actin cytoskeleton and regulation of TGFβ signaling: Exploring their links. Biomolecules 2021; 11(2), 336.
W Hong and KL Guan. The YAP and TAZ transcription co-activators: Key downstream effectors of the mammalian Hippo pathway. Seminars in Cell and Developmental Biology 2012; 23(7), 785-793.
NA Perez-Gonzalez, ND Rochman, K Yao, J Tao, MT Le, S Flanary, L Sablich, B Toler, E Crentsil, F Takaesu, B Lambrus, J Huang, V Fu, P Chengappa, TM Jones, AJ Holland, S An, D Wirtz, RJ Petrie, KL Guan and SX Sun. YAP and TAZ regulate cell volume. Journal of Cell Biology 2019; 218(10), 3472-3488.
S Paul, S Xie, X Yao and A Dey. Transcriptional regulation of the hippo pathway: Current understanding and insights from single-cell technologies. Cells 2022; 11(14), 2225.
Á Ortega, I Vera, MP Diaz, C Navarro, M Rojas, W Torres, H Parra, J Salazar, JB De Sanctis and V Bermúdez. The YAP/TAZ Signaling Pathway in the Tumor Microenvironment and Carcinogenesis: Current Knowledge and Therapeutic Promises. International Journal of Molecular Sciences 2021; 23(1), 430.
E Sidorenko and MK Vartiainen. Nucleoskeletal regulation of transcription: Actin on MRTF. Experimental Biology and Medicine 2019; 244(15), 1372-1381.
BR Frieden and RA Gatenby. Signal transmission through elements of the cytoskeleton form an optimized information network in eukaryotic cells. Scientific Reports 2019; 9(1), 6110.
AA Khilan, NA Al-Maslamani and HF Horn. Cell stretchers and the LINC complex in mechanotransduction. Archives of Biochemistry and Biophysics 2021; 702(1), 108829.
PA Janmey, RG Wells, RK Assoian and CA McCulloch. From tissue mechanics to transcription factors. Differentiation 2013; 86(3), 112-120.
K Wagh, M Ishikawa, DA Garcia, DA Stavreva, A Upadhyaya and GL Hager. Mechanical regulation of transcription: Recent advances. Trends in Cell Biology 2021; 31(6), 457-472.
KN Dahl, AJ Ribeiro and J Lammerding. Nuclear shape, mechanics, and mechanotransduction. Circulation Research 2008; 102(11), 1307-1318.
RB Mokhtari, N Ashayeri, L Baghaie, M Sambi, K Satari, N Baluch, DA Bosykh, MR Szewczuk and S Chakraborty. The hippo pathway effectors YAP/TAZ-TEAD oncoproteins as emerging therapeutic targets in the tumor microenvironment. Cancers 2023; 15(13), 3468.
Z Zhang, P He, L Yang, J Gong, R Qin and M Wang. Posttranslational modifications of YAP/TAZ: Molecular mechanisms and therapeutic opportunities. Cellular and Molecular Biology Letters 2025; 30(1), 83.
A Hooglugt, MMVD Stoel, RA Boon and S Huveneers. Endothelial YAP/TAZ Signaling in Angiogenesis and tumor vasculature. Frontiers in Oncology 2021; 10(1), 612802.
JX Shen, L Zhang, HH Liu, ZY Zhang, N Zhao, JB Zhou, LL Qian and RX Wang. The mechanical role of YAP/TAZ in the development of diabetic cardiomyopathy. Current Issues in Molecular Biology 2025; 47(5), 297.
A Lopez-Hernandez, S Sberna and S Campaner. Emerging principles in the transcriptional control by YAP and TAZ. Cancers 2021; 13(16), 4242.
F Reed, ST Larsuel, MY Mayday, V Scanlon and DS Krause. MRTFA: A critical protein in normal and malignant hematopoiesis and beyond. Journal of Biological Chemistry 2021; 296(1), 100543.
EN Olson and A Nordheim. Linking actin dynamics and gene transcription to drive cellular motile functions. Nature Reviews Molecular Cell Biology 2010; 11(5), 353-365.
S Mascharak, JL Guo, M Griffin, CE Berry, DC Wan and MT Longaker. Modelling and targeting mechanical forces in organ fibrosis. Nature Reviews Bioengineering 2024; 2(4), 305-323.
JM Cook-Mills, ME Marchese and H Abdala-Valencia. Vascular cell adhesion molecule-1 expression and signaling during disease: Regulation by reactive oxygen species and antioxidants. Antioxidants and Redox Signaling 2011; 15(6), 1607-1638.
L Hu, W Chen, A Qian and YP Li. Wnt/β-catenin signaling components and mechanisms in bone formation, homeostasis, and disease. Bone Research 2024; 12(1), 39.
SA Dabravolski, VN Sukhorukov, VA Kalmykov, AV Grechko, NK Shakhpazyan and AN Orekhov. The Role of KLF2 in the regulation of atherosclerosis development and potential Use of KLF2-Targeted Therapy. Biomedicines 2022; 10(2), 254.
K Tatebe, A Zeytun, RM Ribeiro, R Hoffmann, KS Harrod and CV Forst. Response network analysis of differential gene expression in human epithelial lung cells during avian influenza infections. BMC Bioinformatics 2010; 11(1), 170.
B Chaqour and M Goppelt-Struebe. Mechanical regulation of the Cyr61/CCN1 and CTGF/CCN2 proteins. The FEBS Journal 2006; 273(16), 3639-3649.
B Hu, D Zhou, H Wang, N Hu and W Zhao. Mechanical cues regulate histone modifications and cell behavior. Stem Cells International 2022; 2022(1), 9179111.
E Carley, MC King and S Guo. Integrating mechanical signals into cellular identity. Trends in Cell Biology 2022; 32(8), 669-680.
Z Lichner, M Ding, T Khare, Q Dan, R Benitez, M Praszner, X Song, R Saleeb, B Hinz, Y Pei, K Szászi and A Kapus. Myocardin-related transcription factor mediates epithelial fibrogenesis in polycystic kidney disease. Cells 2024; 13(11), 984.
MD Pankratova, AA Riabinin, EA Butova, AV Selivanovskiy, EI Morgun, SV Ulianov, EA Vorotelyak and EP Kalabusheva. YAP/TAZ signalling controls epidermal keratinocyte fate. International Journal of Molecular Sciences 2024; 25(23), 12903.
H Das, A Kumar, Z Lin, WD Patino, PM Hwang, MW Feinberg, PK Majumder and MK Jain. Kruppel-like factor 2 (KLF2) regulates proinflammatory activation of monocytes. Proceedings of the National Academy of Sciences of the United States of America 2006; 103(17), 6653-6658.
A Kindberg, JK Hu and JO Bush. Forced to communicate: Integration of mechanical and biochemical signaling in morphogenesis. Current Opinion in Cell Biology 2020; 66(1), 59-68.
M Lucariello, ML Valicenti, S Giannoni, L Donati, I Armentano, F Morena and S Martino. Mechanobiology in Action: Biomaterials, Devices, and the Cellular Machinery of Force Sensing. Biomolecules 2025; 15(6), 848.
LR Smith, S Cho and DE Discher. Stem cell differentiation is regulated by extracellular matrix mechanics. Physiology 2018; 33(1), 16-25.
S Nellinger and PJ Kluger. How mechanical and physicochemical material characteristics influence adipose-derived stem cell fate. International Journal of Molecular Sciences 2023; 24(4), 3551.
BC Heng, X Zhang, D Aubel, Y Bai, X Li, Y Wei, M Fussenegger and X Deng. Role of YAP/TAZ in cell lineage fate determination and related signaling pathways. Frontiers in Cell and Developmental Biology 2020; 8(1), 735.
JK Virdi and P Pethe. Biomaterials Regulate Mechanosensors YAP/TAZ in Stem Cell Growth and differentiation. Tissue Engineering and Regenerative Medicine 2021; 18(2), 199-215.
S Schiaffino, C Reggiani, T Akimoto and B Blaauw. Molecular mechanisms of skeletal muscle hypertrophy. Journal of Neuromuscular Diseases 2021; 8(2), 169-183.
MA Egerman and DJ Glass. Signaling pathways controlling skeletal muscle mass. Critical Reviews in Biochemistry and Molecular Biology 2014; 49(1), 59-68.
H Cheng, W Zhong, L Wang, Q Zhang, X Ma, Y Wang, S Wang, C He, Q Wei and C Fu. Effects of shear stress on vascular endothelial functions in atherosclerosis and potential therapeutic approaches. Biomedicine and Pharmacotherapy 2023; 158(1), 114198.
N Baeyens, C Bandyopadhyay, BG Coon, S Yun and MA Schwartz. Endothelial fluid shear stress sensing in vascular health and disease. Journal of Clinical Investigation 2016; 126(3), 821-828.
S Chatterjee. Endothelial mechanotransduction, redox signaling and the regulation of vascular inflammatory pathways. Frontiers in Physiology 2018; 9(1), 524.
P Theofilis, M Sagris, E Oikonomou, AS Antonopoulos, G Siasos, C Tsioufis and D Tousoulis. Inflammatory mechanisms contributing to endothelial dysfunction. Biomedicines 2021; 9(7), 781.
A Chalkias. Shear stress and endothelial mechanotransduction in trauma patients with hemorrhagic shock: Hidden coagulopathy pathways and novel therapeutic strategies. International Journal of Molecular Sciences 2023; 24(24), 17522.
HM Han, SY Kim and DH Kim. Mechanotransduction for therapeutic approaches: Cellular aging and rejuvenation. APL Bioengineering 2025; 9(2), 021502.
Y Yokoyama, N Domkam, H Kabir, A Mansour, S Tsukamoto, G Yerima, T Adachi and MRK Mofrad. Emerging mechanomedicines informed by mechanotransduction along the integrin-cytoskeleton-nucleus axis. APL Bioengineering 2025; 9(2), 021503.
E Henke, R Nandigama and S Ergün. Extracellular matrix in the tumor microenvironment and its impact on cancer therapy. Frontiers in Molecular Biosciences 2020; 6(1), 160.
M Cobbaut, S Karagil, L Bruno, MDCD de la Loza, FE Mackenzie, M Stolinski and A Elbediwy. Dysfunctional Mechanotransduction through the YAP/TAZ/Hippo pathway as a feature of chronic disease. Cells 2020; 9(1), 151.
JT Parsons, AR Horwitz and MA Schwartz. Cell adhesion: Integrating cytoskeletal dynamics and cellular tension. Nature Reviews Molecular Cell Biology 2010; 11(9), 633-643.
MR Zanotelli and CA Reinhart-King. Mechanical forces in tumor angiogenesis. Advances in Experimental Medicine and Biology 2018; 1092(1), 91-112.
BM Lozinski, S Ghorbani and VW Yong. Biology of neurofibrosis with focus on multiple sclerosis. Frontiers in Immunology 2024; 15(1), 1370107.
Y Tai, EL Woods, J Dally, D Kong, R Steadman, R Moseley and AC Midgley. Myofibroblasts: Function, formation, and scope of molecular therapies for skin fibrosis. Biomolecules 2021; 11(8), 1095.
H Liu, M Sun, N Wu, B Liu, Q Liu and X Fan. TGF-β/Smads signaling pathway, Hippo-YAP/TAZ signaling pathway, and VEGF: Their mechanisms and roles in vascular remodeling related diseases. Immunity, Inflammation and Disease 2023; 11(11), e1060.
B Hinz and D Lagares. Evasion of apoptosis by myofibroblasts: A hallmark of fibrotic diseases. Nature Reviews Rheumatology 2020; 16(1), 11-31.
X Wang, Y Shen, M Shang, X Liu and LL Munn. Endothelial mechanobiology in atherosclerosis. Cardiovascular Research 2023; 119(8), 1656-1675.
SA Dabravolski, VN Sukhorukov, VA Kalmykov, AV Grechko, NK Shakhpazyan and AN Orekhov. The role of KLF2 in the regulation of atherosclerosis development and potential Use of KLF2-Targeted therapy. Biomedicines 2022; 10(2), 254.
V Kopych, ADS Da Costa and K Park. Endothelial dysfunction in atherosclerosis: Experimental models and therapeutics. Biomaterials Research 2025; 29(1), 0252.
MJ Herzog, P Müller, K Lechner, M Stiebler, P Arndt, M Kunz, D Ahrens, A Schmeißer, S Schreiber and RC Braun-Dullaeus. Arterial stiffness and vascular aging: Mechanisms, prevention, and therapy. Signal Transduction and Targeted Therapy 2025; 10(1), 282.
DD Rosario-Gilabert, A Valenzuela-Miralles and G Esquiva. Advances in mechanotransduction and sonobiology: Effects of audible acoustic waves and low-vibration stimulations on mammalian cells. Biophysical Reviews 2024; 16(6), 783-812.
W Sun, X Gao, H Lei, W Wang and Y Cao. Biophysical approaches for applying and measuring biological forces. Advanced Science 2022; 9(5), e2105254.
O Padovan-Merhar and A Raj. Using variability in gene expression as a tool for studying gene regulation. Wiley Interdisciplinary Reviews: Systems Biology and Medicine 2013; 5(6), 751-759.
K Haase and AE Pelling. Investigating cell mechanics with atomic force microscopy. Journal of the Royal Society Interface 2015; 12(104), 20140970.
DH Cho, S Aguayo and AX Cartagena-Rivera. Atomic force microscopy-mediated mechanobiological profiling of complex human tissues. Biomaterials 2023; 303(1), 122389.
X Hu and M Bao. Advances in micropatterning technology for mechanotransduction research. Mechanobiology in Medicine 2024; 2(3), 100066.
A Zancla, P Mozetic, M Orsini, G Forte and A Rainer. A primer to traction force microscopy. Journal of Biological Chemistry 2022; 298(5), 101867.
AK Denisin, H Kim, IH Riedel-Kruse and BL Pruitt. Field guide to traction force microscopy. Cellular and Molecular Bioengineering 2024; 17(2), 87-106.
GD Marconi, A Mazzone, YD Rocca, O Trubiani, J Pizzicannella and F Diomede. Three-Dimensional culture system: A new frontier in cancer research, drug discovery, and stem cell-based therapy. Biology 2025; 14(7), 875.
WH Abuwatfa, WG Pitt and GA Husseini. Scaffold-based 3D cell culture models in cancer research. Journal of Biomedical Science 2024; 31(1), 7.
L An, Y Liu and Y Liu. Organ-on-a-Chip applications in microfluidic platforms. Micromachines 2025; 16(2), 201.
AG Monteduro, S Rizzato, G Caragnano, A Trapani, G Giannelli and G Maruccio. Organs-on-chips technologies - A guide from disease models to opportunities for drug development. Biosensors and Bioelectronics 2023; 231(1), 115271.
KR Kukurba and SB Montgomery. RNA sequencing and analysis. Cold Spring Harbor Protocols 2015; 2015(11), 951-969.
H Tekin, S Simmons, B Cummings, L Gao, X Adiconis, CC Hession, A Ghoshal, D Dionne, SR Choudhury, V Yesilyurt, NE Sanjana, X Shi, C Lu, M Heidenreich, JQ Pan, JZ Levin and F Zhang. Effects of 3D culturing conditions on the transcriptomic profile of stem-cell-derived neurons. Nature Biomedical Engineering 2018; 2(7), 540-554.
DL Jones, GF Hallström, X Jiang, RC Locke, MK Evans, ED Bonnevie, A Srikumar, TP Leahy, MP Nijsure, JD Boerckel, RL Mauck and NA Dyment. Mechanoepigenetic regulation of extracellular matrix homeostasis via Yap and Taz. Proceedings of the National Academy of Sciences of the United States of America 2023; 120(22), e2211947120.
BV Fearing, L Jing L, MN Barcellona, SE Witte, JM Buchowski, LP Zebala, MP Kelly, S Luhmann, MC Gupta, A Pathak and LA Setton. Mechanosensitive transcriptional coactivators MRTF-A and YAP/TAZ regulate nucleus pulposus cell phenotype through cell shape. The FASEB Journal 2019; 33(12), 14022-14035.
J Isaacson and M Brotto. Physiology of mechanotransduction: How do muscle and bone “Talk” to one another? Clinical Reviews in Bone and Mineral Metabolism 2014; 12(2), 77-85.
PF Lenne, E Munro, I Heemskerk, A Warmflash, L Bocanegra-Moreno, K Kishi, A Kicheva, Y Long, A Fruleux, A Boudaoud, TE Saunders, P Caldarelli, A Michaut, J Gros, Y Maroudas-Sacks, K Keren, E Hannezo, ZJ Gartner, B Stormo, A Gladfelter, A Rodrigues, A Shyer, N Minc, JL Maître, S Di Talia, B Khamaisi, D Sprinzak and S Tlili. Roadmap for the multiscale coupling of biochemical and mechanical signals during development. Physical Biology 2021; 18(4), 041501.
J Wang, D Lü, D Mao and M Long. Mechanomics: An emerging field between biology and biomechanics. Protein and Cell 2014; 5(7), 518-531.
PHG Sanches, NCD Melo, AM Porcari and LMD Carvalho. Integrating molecular perspectives: Strategies for comprehensive multi-omics integrative data analysis and machine learning applications in transcriptomics, proteomics, and metabolomics. Biology 2024; 13(11), 848.
O Ntousi, M Roumpi, PK Siogkas, D Polyzos, I Kakkos, GK Matsopoulos and DI Fotiadis. Advances in computational modeling of scaffolds for bone tissue engineering: A narrative review of the current approaches and challenges. Biomechanics 2025; 5(4), 76.
R Oria, K Jain and VM Weaver. Exploring the intersection of mechanobiology and artificial intelligence. Npj Biological Physics and Mechanics 2025; 2(1), 9.
L Donati, ML Valicenti, S Giannoni, F Morena and S Martino. Biomaterials mimicking mechanobiology: A Specific design for a specific biological application. International Journal of Molecular Sciences 2024; 25(19), 10386.
W Tiskratok, N Chuinsiri, P Limraksasin, M Kyawsoewin and P Jitprasertwong. Extracellular matrix stiffness: mechanotransduction and mechanobiological response-driven strategies for biomedical applications targeting fibroblast inflammation. Polymers 2025; 17(6), 822.
LE Charelli, JPD Ferreira, CP Naveira-Cotta and TA Balbino. Engineering mechanobiology through organoids-on-chip: A strategy to boost therapeutics. Journal of Tissue Engineering and Regenerative Medicine 2021; 15(11), 883-899.