
Trends
Sci.
2026;
23(5):
12071
Unraveling Potential Bioactive Antimycobacterial Compounds from Endophytic Aspergillus tubingensis and Syncephalastrum racemosum: An Integrated Bioassay, Metabolite Profiling, and Molecular Mechanism Approach
Syariful Anam1,*, Armini Syamsidi1, Ni Luh Putu Dwijayanti1, Zidan Saputra1,
Rizqi Hulul Amalia1, Thylka Mawadha Tamagola1, Yuliet1, Abd. Rahman Razak2,
Agnes Dwi Sis Perwitasari3 and Saipul Maulana1
1Department of Pharmacy, Faculty of Mathematics and Natural Sciences, Tadulako University,
Central Sulawesi 94148, Indonesia
2Department of Chemistry, Faculty of Mathematics and Natural Sciences, Tadulako University,
Central Sulawesi 94148, Indonesia
3Tropical Disease Diagnostic Center, Universitas Airlangga, Surabaya, East Java 60115, Indonesia
(*Corresponding author’s e-mail: [email protected])
Received: 26 September 2025, Revised: 13 October 2025, Accepted: 1 November 2025, Published: 10 January 2026
Abstract
Tuberculosis (TB), a significant global health threat, ranks third in the world, with 67% of cases occurring in productive age groups. Natural compounds, such as fungal secondary metabolites, exhibit various pharmaceutically relevant activities, including antimycobacterial activity. Our ongoing research investigated the ethyl acetate extract of endophytic fungi Aspergillus tubingensis and Syncephalastrum racemosum from Rui (Harrisonia perforata) against Mycobacterium tuberculosis. The extract was obtained by fermenting the fungi on PDB and extracting the liquid culture using ethyl acetate. It showed antimycobacterial activity against Mycobacterium tuberculosis H37Rv by culturing the crude extract and bacteria in Middlebrook 7H9 Broth and Middlebrook 7H10 Agar. UHPLC-MS/MS profiling revealed about 85 compounds from Aspergillus tubingensis and 105 from Syncephalastrum racemosum. Molecular docking analysis revealed potential dual-action mechanisms by targeting two crucial enzymes in M. tuberculosis: KatG and KasA. Potential metabolites include bis(methylbenzylidene)sorbitol, sphinganine, and a chromenone derivative. In conclusion, the crude extract of Aspergillus tubingensis and Syncephalastrum racemosum demonstrates potential as an antimycobacterial agent.
Keywords: Antimycobacterial, Aspergillus tubingensis, Syncephalastrum racemosum, Ethyl acetate extract, Harrisonia perforata, Molecular docking, Mass Spectrometry
Introduction
The AMR organization reported in 2014 that resistant bacterial strains, HIV, tuberculosis, and malaria cause up to 700,000 deaths annually. They estimate antibiotic resistance will increase to 10 million cases in 2050, with cumulative costs of around 100 trillion USD [1]. WHO reports that multidrug-resistant tuberculosis remains a global public health threat, as many as 10 million people suffer from TB, and 1.2 million people die every year. An estimated 484,000 new rifampicin-resistant cases occurred in 2020 [2]. Indonesia accounts for two-thirds of the world’s TB cases, with most cases occurring in West Java, East Java, Central Java, DKI Jakarta, and North Sumatra in 2019 [3]. Indonesia’s 824,000 TB cases ranked third globally, after India and China, with 67% occurring at a productive age [4].
In efforts to overcome antimicrobial resistance, primarily Mycobacterium tuberculosis, several treatment approaches have been used. Nevertheless, antimicrobials are the first treatment choice for Tuberculosis (TB). However, the first line-up of antimicrobials has been resistant, causing the treatment time to be longer [5-7]. In addition, drug resistance cases increased, resulting in the discovery of new antimicrobials that are becoming essential.
Antimicrobial-based plants/herbs to treat infectious diseases are one approach that has been applied. According to the WHO, 80% of the worldʼs population still relies on traditional medicine, including plants/herbs. Until now, a quarter of modern medicines circulating worldwide come from active ingredients isolated and developed from plants [8]. One of the plants that is empirically used for antidiarrhea in the Palu is Rui (Harrisonia perforata (Blanco) Merr.) [9]. Scientifically, Rui has been proven to inhibit the growth of bacteria such as Escherichia coli, Salmonella typhi, and Shigella dysentriae [10]. Ethanolic extract of branches from H. perforata showed anti-TB activity [11].
Plant-based antimicrobials faced limited factors, including the level of production of medicines due to the limitation of herbal raw materials, conservation issues, and cultivation problems [12,13]. Antimicrobial compounds from endophytic fungi are becoming potential alternatives for secondary metabolite sources. Endophytic fungi have been proven to be a source of antimycobacterial compounds [14-17].
Endophytic fungi are prominent sources of antimicrobial metabolites. Mohamed et al. [18] reported that compounds of Aspergillus tubingensis extract showed antibacterial activities against Pseudomonas aeruginosa and Escherichia coli. Tapfuma et al. [19] reported that methanol crude extract from Clonostachys rogersoniana MGK33 showed antimicrobial activity against M. smegmatis mc2155 and M. tuberculosis H37Rv. In silico studies revealed bionectin F as a potential inhibitor of M. tuberculosis β-ketoacyl-acyl carrier protein reductase (MabA). Andrioli et al. [20] studied the antimycobacterial activity of six fungal metabolites: Austdiol (1), austdiol diacetate (2), mycoleptone A (3), eugenitin (4) from Mycoleptodiscus indicus, emodin (5) from Penicillium citrinum and δ-lactam (6) from Humicola grisea. Emodin was the most active against M. tuberculosis and M. kansasii strains, followed by austdiol, austdiol diacetate, and mycoleptone A [20]. Marine-derived fungi exhibit antimycobacterial activity. A study found 19 genera of marine fungi with 139 metabolites, 131 of which are antimycobacterial. These metabolites include polyketides (57.58%), alkaloids and peptides (25.76%), terpenoids and steroids (13.64%), and miscellaneous compounds (3.03%). Polyketides are diverse and have distinct bioactivities. Between 2002 and 2023, half of all marine-fungal antimycobacterial compounds reported each year were novel. Novel compounds were more common in alkaloids and peptides than in other groups. Polyketides, the largest group, have more known compounds than novel ones. This is partly due to their abundance in fungi, leading to more frequent discovery [21].
Our previous study revealed that the fungal extract of A. tubingensis and S. racemosum from H. perforata showed antibacterial activity against Staphylococcus aureus and Salmonella typhi, with the MIC and MBC starting from 24 to 32 mg/mL. Fungal extract exhibits cytotoxic activity with an LC 30.83 ± 0.39 µg/mL [22]. Therefore, as part of our ongoing research into the source of antimycobacterial compounds, we are investigating the antimycobacterial activity of the ethyl acetate extract of A. tubingensis and S. racemosum isolated from Rui (H. perforata (Blanco) Merr.).
Materials and methods
Materials
Bacteria Mycobacterium tuberculosis H37Rv, Middlebrook 7H10 Agar, Middlebrook 7H9 Broth, Middlebrook Oleic Albumin Dextrose Catalase (OADC) Growth Supplement, Potato Dextrose Agar (PDA) (Merck), Potato Dextrose Broth (PDB) (Merck), distilled water, ethyl acetate (Brataco Chem.), Biosafety Cabinet (BSC), Autoclave, Incubator, Laminar Air Flow (LAF), and Liquid Chromatography-High Resolution Mass Spectrometry (LC-HRMS). Computational studies in this research were performed on a Dell workstation running Ubuntu 20.04.3 LTS, featuring an Intel® Xeon® W-2223 CPU @ 3.60 GHz, 16 GB of RAM, and an NVIDIA RTX 4060 Ti GPU.
Fermentation and extraction of endophytic fungi
A. tubingensis was inoculated at three points on a petri dish containing PDA media and then incubated at 30 °C for seven days. Five mL of NaCl 0.9% was transferred to a petri dish; then, the fungal biomass was homogenized in the tube and subsequently measured, the absorbance on a spectrophotometer at 600 nm to obtain OD 0.3. One mL of fungal suspension was inoculated into 100 mL of PDB (Potato Dextrose Broth) media (KgaA Darmstadt) in an Erlenmeyer flask, incubated at 30 °C for ten days with shaking at 120 rpm. The liquid culture was filtered using a Whatmann paper, and the filtrate was extracted with a 1:1 ratio of ethyl acetate (Brataco Chem.) twice. The concentrated extract was obtained using a rotary vacuum evaporator [22-24].
Antimycobacterial assay
This antimycobacterial assay started with the first step: Prepare Middlebrook 7H9 broth media (1.6 g in 300 mL of distilled water) and homogenize it in the Erlenmeyer flask. The media was transferred 4.5 mL into a screw-top glass tube and sterilized using an autoclave at 121°C for 15 min. Perform a 1×24-hour sterility test, and then the Middlebrook 7H9 broth media is ready for use. Transferred 500 µl of the 0.5 McFarland H37Rv bacterial suspension into a screw-top glass tube and added 4.5 mL of 7H9 media. Each sample with a series of dilution concentrations (4%, 2%, 1%, 0.5%, 0.25% and 0.125%) was transferred 500 µl into a screw glass tube, and 500 µl of H37Rv 0.5McFarland bacteria was added. Homogenized and incubated at 37 °C for one week. The second step used the solid dilution method by making Middlebrook 7H10 agar media (22 g in 1,000 mL of distilled water), homogenized using a microwave for 1 min, then autoclaved at 121 °C for 10 min. At a temperature of 50 - 55 °C, add 2.5 mL of the OADC media supplement. The media is poured into a petri dish ± 25 - 30 mL. Please wait until it solidifies and carry out a 1×24-hour sterility test. Media that does not contain contamination is ready to be used for testing. Take the culture suspension/liquid incubated for one week and inoculate 100 µl on agar media. Incubate for 1 - 3 weeks at 37 °C and observe growth. All series concentrations were performed twice [25,26].
HPLC-HRMS experiments
Mass spectra were acquired on a Thermo Scientific™ Vanquish™ UHPLC Binary Pump equipped with a reversed-phase column Thermo Scientific™ Accucore™ Phenyl-Hexyl 100 mm×2.1 mm ID×2.6 µm maintained at 40 °C with volume injection at 3 µL. The mobile phases used were MS-grade water containing 0.1% formic acid (A) and MS-grade methanol containing 0.1% formic acid (B), employing a gradient technique with a 0.3 mL/min flow rate. First, the mobile phase B was set at 5% and increased gradually to 90% in 16 min. Then, it was held at 90% for 4 min and continued to the initial condition (5% B) until 25 min. The HPLC was connected to an Orbitrap high-resolution mass spectrometry (Thermo Scientific™ Q Exactive™ Hybrid Quadrupole-Orbitrap™ High-Resolution Mass Spectrometer) equipped with an electrospray ionization (ESI) source. Mass spectra were acquired in positive-ion mode with a capillary voltage of 3.30 kV, a drying temperature of 320 °C, and a scan range of 66.7 - 1,000 m/z [27-29].
Ligand preparation
The secondary metabolites identified in the ethyl acetate extracts of Aspergillus tubingensis and Syncephalastrum racemosum through Ultra-High Performance Liquid Chromatography-High Resolution Mass Spectrometry (UHPLC-HRMS) were used as test ligands. These compoundsʼ two-dimensional (2D) structures were generated using ChemDraw 19.0. Subsequently, the structures were optimized to 3D structures by energy minimization, adding hydrogens, and protonated at a physiological pH of 7.4 ± 0.2 using the Epik tool in the LigPrep module within the Schrödinger Suite 2020-3.
Receptor preparation
A set of multiple protein targets involved in the pathogenesis of Mycobacterium tuberculosis was selected for molecular docking simulations. The crystal structures of these receptors were obtained from the Protein Data Bank (PDB), specifically isoniazid-bound KatG catalase-peroxidase (PDB ID: 3WXO), KasA β-ketoacyl synthase (PDB ID: 2WGE), the protein kinase domain PknB (PDB ID: 6B2P), the transcription initiation complex (PDB ID: 5UHB), and quinolinic-acid phosphoribosyltransferase (QAPRTase; PDB ID: 1QPQ). Before docking, each receptor was prepared using the Protein Preparation Wizard in Maestro (Schrödinger Suite 2020-3), which involved the removal of residual solvents and co-crystallized ligands, adding hydrogen atoms, assigning bond orders, and a restrained energy minimization with the OPLS3e force field to relieve steric clashes and optimize protein geometry.
Molecular docking
A molecular docking study was carried out using the Glide module of the Schrödinger Suite. For each receptor, the docking grid was generated around the binding site based on the coordinates of its co-crystallized ligand, which was isoniazid for KatG catalase-peroxidase (PDB ID: 3WXO), thiolactomycin for KasA β-ketoacyl synthase (PDB ID: 2WGE), and a synthetic inhibitor (5-{5-chloro-4-[(5-cyclopropyl-1H-pyrazol-3-yl)amino]pyrimidin-2-yl}thiophene-2-sulfonamide) for the protein kinase domain PknB (PDB ID: 6B2P), rifampicin for the transcription initiation complex (RNA polymerase; PDB ID: 5UHB), and quinolinic acid for quinolinic-acid phosphoribosyltransferase (QAPRTase; PDB ID: 1QPQ).
The prepared ligands were subsequently docked into the active sites using the Extra Precision (XP) mode in Glide. The docking poses were ranked according to their GlideScore for further evaluation. To complement docking results and predict the binding free energy, the best poses were further analyzed using the Molecular Mechanics Generalized Born Surface Area (MM-GBSA) approach as implemented in the Prime module of Schrödinger.
Results and discussion
A. tubingensis (Figure 1(A)) is part of the phylum Ascomycota and has become one of the numerous existing endophytic fungi. Aspergillus strains constitute one of the most prolific sources of secondary metabolites with diverse chemical classes and engaging biological activities. The isolated metabolites were chemically varied and exhibited various biological activities such as antibacterial, anti-cancer, anti-plasmodial, anti-inflammatory, antioxidant, immunosuppressive, and antifungal activities [30].
S. racemosum (Figure 1(B)) is part of the phylum Mucoromycota and is also well known as an endophytic fungus from plants. The previous study reveals that metabolites from ethyl acetate extract of S. racemosum exhibited antioxidant activity with 41.84 ± 0.38 g/mL and cytotoxic activity against T47D cells at 420.06 ± 12.98 g/mL [31].
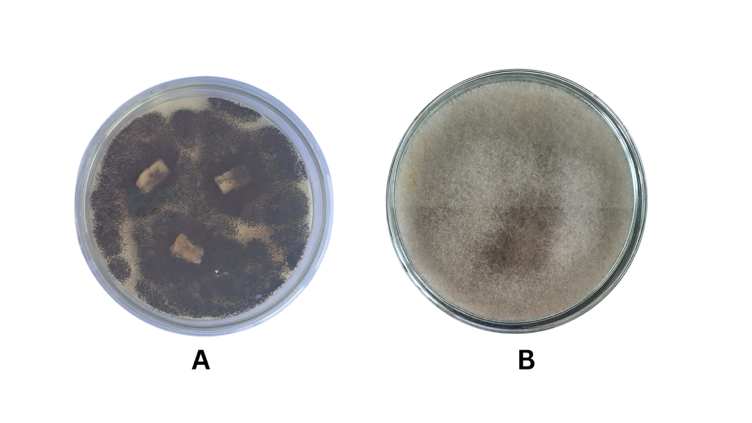
Figure 1 (A) A. tubingensis culture in media PDA incubation at 30 °C for seven days. (B) S. racemosum culture in media PDA incubation at 30 °C for 7 days.
Our study determined the antimycobacterial activity of the ethyl acetate extract of A. tubingensis and S. racemosum using M. tuberculosis H37Rv. Table 1 represents the results of the antimycobacterial assay. The ethyl acetate extract showed inhibition, as indicated by the absence of colonies, which had a white color in the medium (Figures 2 and 3). In this assay, 6 series concentrations were applied, and all concentrations showed inhibition from the highest concentration, 40 mg/mL, to the lowest concentration of 1.25 mg/mL. However, the extract from fungus S. racemosum showed colonies of Mycobacterium in the lowest concentration, 1.25 mg/mL (Figure 3). This result indicated that the ethyl acetate extract from 2 endophytic fungi has potency as an antimycobacterial against M. tuberculosis H37Rv.
Table 1 Antimycobacterial assay of ethyl acetate extract of A. tubingensis and S. racemosum.
Concentration (mg/mL) |
Colony M. tuberculosis (CFU/mL) |
|||||
Extract from A. tubingensis |
Extract from S. racemosum |
|||||
Replicate 1 |
Replicate 2 |
Mean |
Replicate 1 |
Replicate 2 |
Mean |
|
40.0 |
0 |
0 |
0 |
0 |
0 |
0 |
20.0 |
0 |
0 |
0 |
0 |
0 |
0 |
10.0 |
0 |
0 |
0 |
0 |
0 |
0 |
5.0 |
0 |
0 |
0 |
0 |
0 |
0 |
2.5 |
0 |
0 |
0 |
0 |
0 |
0 |
1.25 |
0 |
0 |
0 |
> 105 |
> 105 |
> 105 |
In a previous study, fungal endophytes showed inhibition of Mycobacterium bacteria. Ferreira 2017 reported that Trichoderma effusum endophytic fungi from the plant Vellozia gigantea displayed selective antibacterial activity against methicillin-resistant Staphylococcus aureus and Mycobacterium intracellulare [32]. Andrioli 2022 reported that Azaphilone compounds and emodin from endophytic fungi Mycoleptodiscus indicus and Humicola grisea exhibited antimycobacterial activity [20].
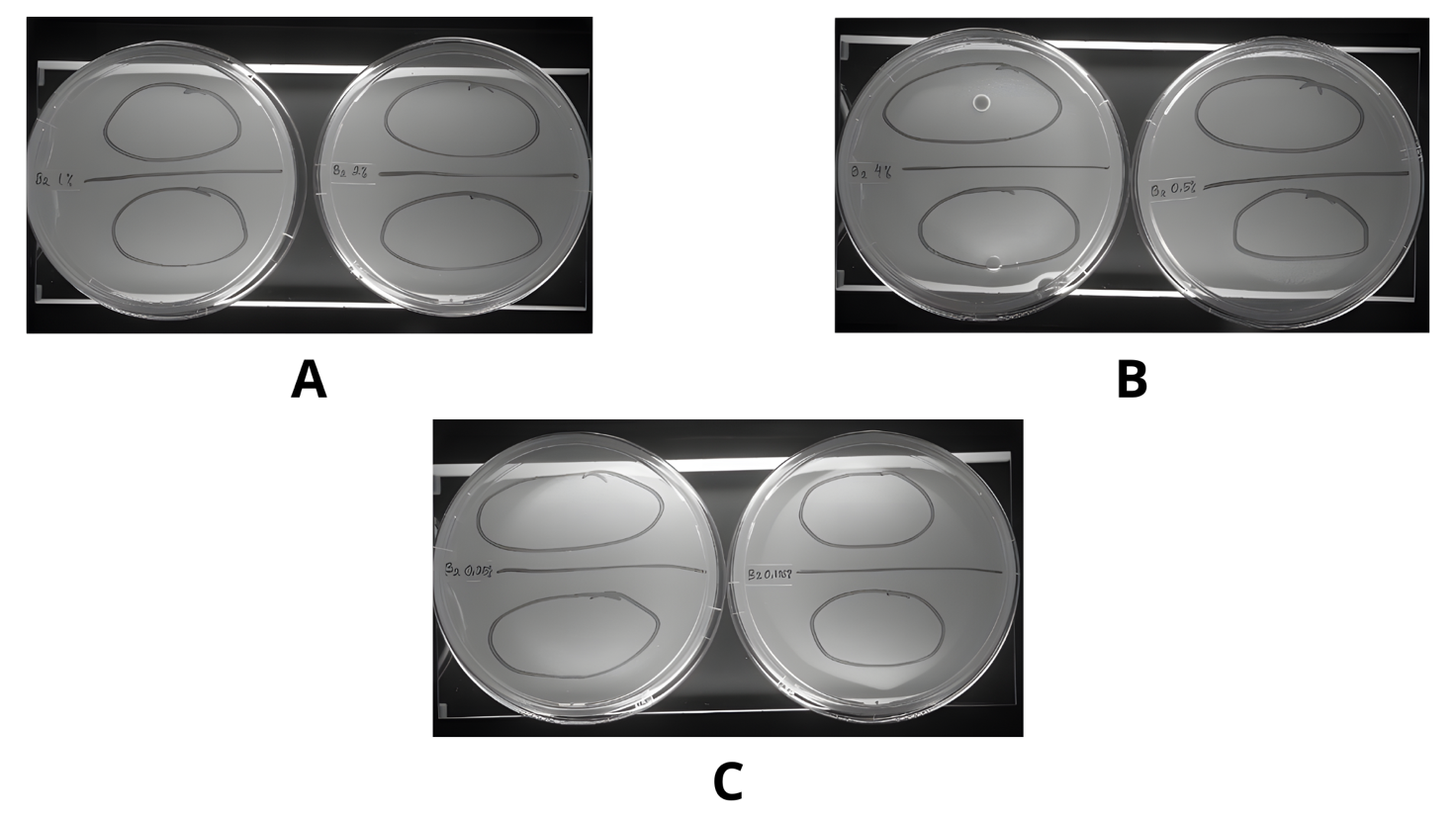
Figure 2 Antimycobacterial assay of ethyl acetate extract of A. tubingensis on middlebrook 7H10 agar media. (A) Concentration 10.0 and 20.0 mg/mL. (B) Concentration 5.0 and 40.0 mg/mL. (C) Concentration 2.5 and 1.25 mg/mL.
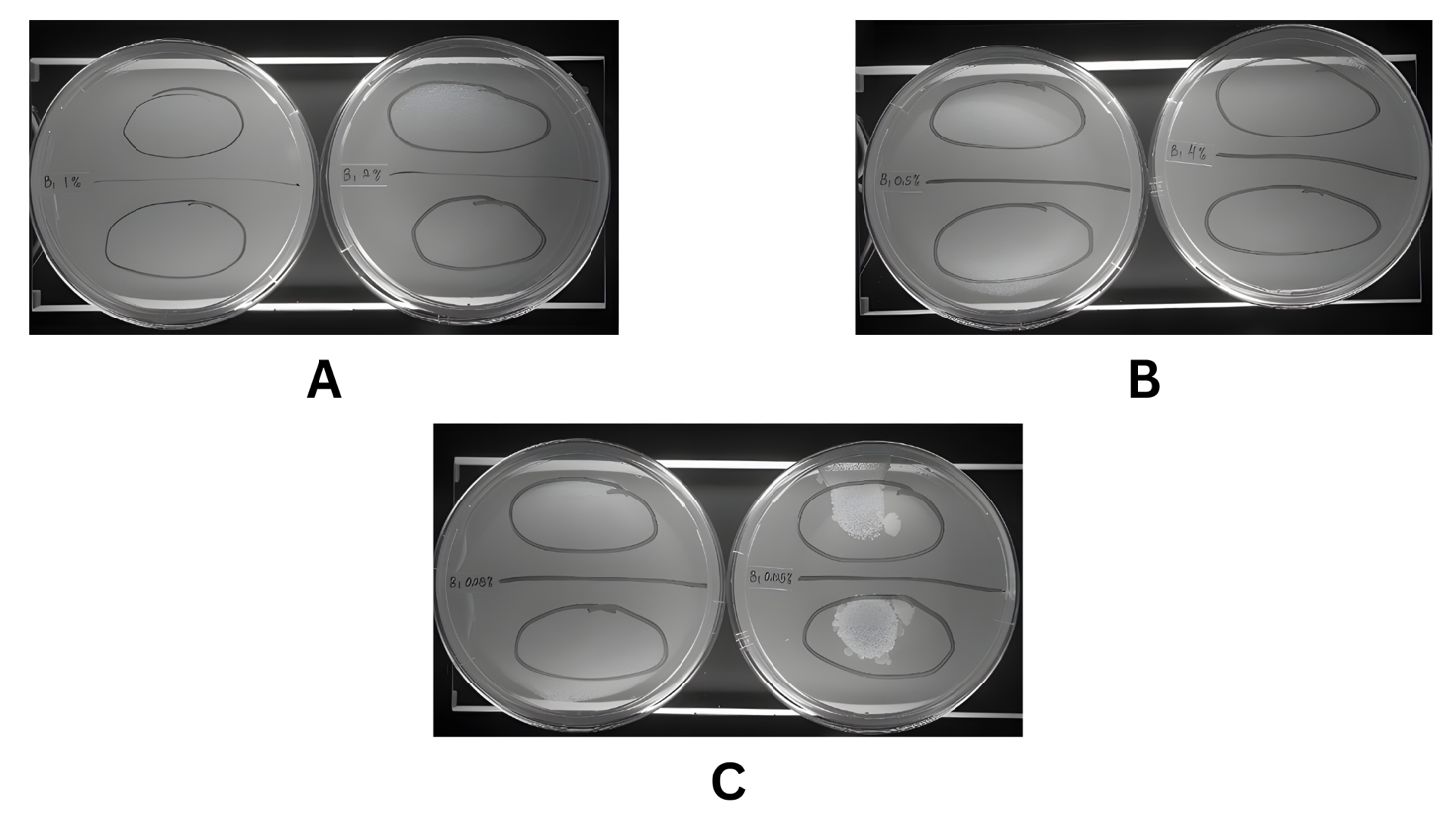
Figure 3 Antimycobacterial assay of ethyl acetate extract of S. racemosum on middlebrook 7H10 agar media. (A) Concentration 10.0 and 20.0 mg/mL. (B) Concentration 5.0 and 40.0 mg/mL. (C) Concentration 2.5 and 1.25 mg/mL.
Table 2 LC-HRMS/MS Profile of ethyl acetate extract of A. tubingensis.
Name |
Formula |
Calc. MW |
RT [min] |
Area (Max.) |
2-Mercaptoethanol |
C2 H6 O S |
78.0142 |
0.889 |
22082042776 |
2-Mercaptoethanol |
C2 H6 O S |
78.0141 |
1.098 |
8058535539 |
Oleamide |
C18 H35 N O |
281.271 |
14.88 |
502810832,8 |
5-Benzyldihydro-2(3H)-furanone |
C11 H12 O2 |
176.084 |
5.57 |
424169701.2 |
N,N-Bis(2-hydroxyethyl)dodecanamide |
C16 H33 N O3 |
287.246 |
10.1 |
252365912.3 |
5-Benzyldihydro-2(3H)-furanone |
C11 H12 O2 |
176.084 |
5.844 |
186611290 |
Cuminaldehyde |
C10 H12 O |
148.089 |
5.57 |
180250687.6 |
Hydroxy-1,4-benzoquinone |
C6 H4 O3 |
124.016 |
1.584 |
172017098.2 |
4-methyl-5-oxo-2-pentyl-2,5-dihydrofuran-3-carboxylic acid |
C11 H16 O4 |
212.105 |
7.258 |
137838223.1 |
EL9600000 |
C10 H12 O |
148.089 |
5.844 |
135899950.7 |
2-Amino-1,3,4-octadecanetriol |
C18 H39 N O3 |
317.292 |
9.939 |
114290409.9 |
5-Benzyldihydro-2(3H)-furanone |
C11 H12 O2 |
176.084 |
8.056 |
107848234.7 |
Bis(methylbenzylidene)sorbitol |
C22 H26 O6 |
386.172 |
10.62 |
100771380.9 |
5-Benzyldihydro-2(3H)-furanone |
C11 H12 O2 |
176.084 |
8.146 |
85570859.29 |
(1S,4S,7R,10S,13S)-4,7-bis(butan-2-yl)-10-(propan-2-yl)-15,16-dithia-2,5,8,11,19-pentaazabicyclo[11.4.2]nonadecane-3,6,9,12,18-pentone |
C23 H39 N5 O5 S2 |
529.238 |
10.58 |
71539516.7 |
Bis(4-ethylbenzylidene)sorbitol |
C24 H30 O6 |
414.204 |
11.57 |
63819500.32 |
5-Benzyldihydro-2(3H)-furanone |
C11 H12 O2 |
176.084 |
7.01 |
57820313.09 |
1-Phenyl-2-butanone |
C10 H12 O |
148.089 |
8.143 |
57656328.14 |
sphinganine |
C18 H39 N O2 |
301.298 |
10.97 |
53263590.48 |
(10S)-Juvenile hormone III diol |
C16 H28 O4 |
284.198 |
9.804 |
50298306.24 |
Linoleamide |
C18 H33 N O |
279.256 |
14.23 |
46322763.91 |
Hydroxy-1,4-benzoquinone |
C6 H4 O3 |
124.016 |
5.001 |
44662639.2 |
nylon cyclic dimer |
C12 H22 N2 O2 |
226.168 |
2.962 |
44067510.78 |
2-(2H-Benzotriazol-2-yl)-4,6-bis(1-methyl-1-phenylethyl)phenol |
C30 H29 N3 O |
447.231 |
17.64 |
41545223.38 |
Cuminaldehyde |
C10 H12 O |
148.089 |
8.051 |
40025299.83 |
3-Allylcyclohexene |
C9 H14 |
122.11 |
9.413 |
39369640.74 |
3,4-Dimethoxycinnamic acid |
C11 H12 O4 |
208.074 |
5.702 |
36567348.39 |
Bicine |
C6 H13 N O4 |
163.084 |
0.779 |
35290440.34 |
Dodecylamine |
C12 H27 N |
185.214 |
9.871 |
35040379.34 |
6-Hydroxy-5-methyl-4,11-dioxoundecanoic acid |
C12 H20 O5 |
244.131 |
7.01 |
32619661.12 |
n-propylbenzene |
C9 H12 |
120.094 |
5.569 |
31806376.74 |
(2E,4E,7E)-2,4,7-Decatrienoic acid |
C10 H14 O2 |
166.099 |
5.844 |
30586210.01 |
Sorbic acid |
C6 H8 O2 |
112.053 |
2.795 |
29149825.28 |
3-[(4-hydroxyphenyl)methyl]-octahydropyrrolo[1,2-a]pyrazine-1,4-dione |
C14 H16 N2 O3 |
260.116 |
3.842 |
28819539.31 |
5-Ethyl-4-hydroxy-3(2H)-furanone |
C6 H8 O3 |
128.047 |
1.291 |
28021248.66 |
Lauramide |
C12 H25 N O |
199.194 |
11.9 |
27865395.2 |
(1S)-1,5-Anhydro-1-benzyl-D-galactitol |
C13 H18 O5 |
254.115 |
9.647 |
27850710.4 |
NP-011220 |
C11 H18 N2 O2 |
210.137 |
5.131 |
25503586.17 |
n-propylbenzene |
C9 H12 |
120.094 |
5.844 |
23736034.52 |
NSC 622050 |
C16 H31 N O3 |
285.23 |
12.53 |
23502911.22 |
Diethanolamine |
C4 H11 N O2 |
105.079 |
0.759 |
22072745.06 |
QL5300000 |
C10 H12 O |
148.089 |
7.009 |
22005584.29 |
{4-Ethyl-6-[(3E)-2-ethyl-3-hexen-1-yl]-6-methyl-1,2-dioxan-3-yl}acetic acid |
C17 H30 O4 |
298.214 |
11.61 |
21476957.29 |
(10S)-Juvenile hormone III diol |
C16 H28 O4 |
284.198 |
9.362 |
20774355.95 |
Glycocholic acid |
C26 H43 N O6 |
465.309 |
10.72 |
20461003.45 |
(2E,4E,7E)-2,4,7-Decatrienoic acid |
C10 H14 O2 |
166.099 |
5.57 |
20201238.89 |
Carvone |
C10 H14 O |
150.104 |
5.844 |
19952466.47 |
1-Methyl-1-benzylidene-acetone |
C11 H12 O |
160.089 |
10.64 |
19740602.57 |
Myristamide |
C14 H29 N O |
227.225 |
13.23 |
17635113.91 |
5-[(5-Hydroxytetrahydro-2-furanyl)methyl]-1,3-benzenediol |
C11 H14 O4 |
210.089 |
5.694 |
16661226.88 |
N,N-Bis(2-hydroxyethyl)dodecanamide |
C16 H33 N O3 |
287.246 |
11.05 |
14809643.71 |
DEET |
C12 H17 N O |
191.131 |
9.305 |
14573505.4 |
4'-Methoxyacetophenone |
C9 H10 O2 |
150.068 |
5.694 |
14500402.81 |
Benzaldehyde |
C7 H6 O |
106.042 |
5.57 |
13981436 |
C16-Dihydroceramide |
C34 H69 N O3 |
539.526 |
17.56 |
13727370.88 |
EL9600000 |
C10 H12 O |
148.089 |
7.248 |
13149145.82 |
Styrene |
C8 H8 |
104.063 |
5.845 |
12388073.67 |
5-(3,4-Dihydroxyphenyl)-4-hydroxypentanoic acid |
C11 H14 O5 |
226.084 |
5.702 |
11549860.99 |
(2S)-4-Methyl-2-({[(3S,4S,5R)-2,3,4-trihydroxy-5-(hydroxymethyl)tetrahydro-2-furanyl]methyl}amino)pentanoic acid (non-preferred name) |
C12 H23 N O7 |
293.147 |
7.29 |
11224035.02 |
NP-021018 |
C12 H18 O4 |
226.121 |
9.558 |
10934746.17 |
(2E,4E,7E)-2,4,7-Decatrienoic acid |
C10 H14 O2 |
166.099 |
8.144 |
10407772.98 |
Styrene |
C8 H8 |
104.063 |
5.57 |
10301303.61 |
n-propylbenzene |
C9 H12 |
120.094 |
8.143 |
10156792.55 |
1-Methyl-1-benzylidene-acetone |
C11 H12 O |
160.089 |
13.44 |
9896860.355 |
butyrin |
C15 H26 O6 |
302.173 |
13.41 |
9423518.394 |
Maculosin |
C14 H16 N2 O3 |
260.116 |
5.355 |
8788314.047 |
N,N-Dimethyldecylamine N-oxide |
C12 H27 N O |
201.209 |
8.467 |
8214689.508 |
Anisole |
C7 H8 O |
108.058 |
5.844 |
7662268.941 |
3-(propan-2-yl)-octahydropyrrolo[1,2-a]pyrazine-1,4-dione |
C10 H16 N2 O2 |
196.121 |
3.863 |
7152893.693 |
4-Phenyl-3-buten-2-one |
C10 H10 O |
146.073 |
5.57 |
7143183.705 |
Decanamide |
C10 H21 N O |
171.162 |
10.31 |
6882174.212 |
n-propylbenzene |
C9 H12 |
120.094 |
8.051 |
6873776.058 |
Indole |
C8 H7 N |
117.058 |
5.569 |
6659469.822 |
Indole |
C8 H7 N |
117.058 |
5.844 |
6199741.338 |
(1R,2R,3S,5R,6R,7R,8S,9S,12R)-2,8-Dihydroxy-12-isopropenyl-7-methyl-11H-spiro[4,10-dioxatetracyclo[7.2.1.0~2,7 ~.0~3,5~]dodecane-6,2'-oxiran]-11-one |
C15 H18 O6 |
294.11 |
7.486 |
6140429.408 |
4-methyl-5-oxo-2-pentyl-2,5-dihydrofuran-3-carboxylic acid |
C11 H16 O4 |
212.104 |
5.188 |
5610847.348 |
Styrene |
C8 H8 |
104.063 |
8.144 |
5560314.699 |
2,5-Dipropyl-4-methylthiazole |
C10 H17 N S |
183.108 |
7.064 |
4973296.97 |
Acetophenone |
C8 H8 O |
120.058 |
5.844 |
4967946.796 |
Acetophenone |
C8 H8 O |
120.058 |
5.569 |
4914014.542 |
(1E)-1-(4-Methoxyphenyl)-4-methyl-1,4-pentadien-3-one |
C13 H14 O2 |
202.099 |
10.66 |
4827803.826 |
(2E,4E,7E)-2,4,7-Decatrienoic acid |
C10 H14 O2 |
166.099 |
8.051 |
4716464.241 |
p-Hydroxybenzalacetone |
C10 H10 O2 |
162.068 |
5.702 |
4654149.4 |
Tetralin |
C10 H12 |
132.094 |
8.145 |
4538414.898 |
Choline |
C5 H13 N O |
103.1 |
0.765 |
3909058.756 |
Table 2 represents the LC-HRMS/MS profile of the crude ethyl acetate extract of A. tubingensis, showing about 85 tentative compounds.
Table 3 LC-HRMS/MS Profile of ethyl acetate extract of S. racemosum.
Name |
Formula |
Calc. MW |
RT [min] |
Area (Max.) |
2-Mercaptoethanol |
C2 H6 O S |
78.01422 |
0.884 |
26758497357 |
2-Mercaptoethanol |
C2 H6 O S |
78.01422 |
1.096 |
13338910231 |
Methylimidazoleacetic acid |
C6 H8 N2 O2 |
140.05847 |
1.905 |
414201289.7 |
DA9185000 |
C12 H10 O S |
202.04502 |
0.761 |
380815795.7 |
3-[(1S)-2-Cyclohexen-1-yl]-L-alanine |
C9 H15 N O2 |
169.11007 |
5.125 |
309450975.7 |
Cyclo(phenylalanyl-prolyl) |
C14 H16 N2 O2 |
244.12096 |
6.056 |
284560619.9 |
NP-011220 |
C11 H18 N2 O2 |
210.13661 |
5.409 |
260133092.1 |
3-(propan-2-yl)-octahydropyrrolo[1,2-a]pyrazine-1,4-dione |
C10 H16 N2 O2 |
196.12107 |
3.829 |
216420394.8 |
N,N-Bis(2-hydroxyethyl)dodecanamide |
C16 H33 N O3 |
287.24562 |
10.089 |
212555706 |
MFCD18695608 |
C11 H15 N O4 |
225.09993 |
5.244 |
147184252.8 |
Butyl 4-aminobenzoate |
C11 H15 N O2 |
193.11009 |
7.297 |
136887695.8 |
2-Amino-1,3,4-octadecanetriol |
C18 H39 N O3 |
317.29236 |
9.935 |
134753248.8 |
1-Hydroxy-3-methoxy-10-methyl-9(10H)-acridinone |
C15 H13 N O3 |
255.08937 |
7.662 |
120455703.1 |
Bis(methylbenzylidene)sorbitol |
C22 H26 O6 |
386.17222 |
10.622 |
117157589 |
3-[(4-hydroxyphenyl)methyl]-octahydropyrrolo[1,2-a]pyrazine-1,4-dione |
C14 H16 N2 O3 |
260.116 |
3.916 |
107646109.9 |
Hydrocotarnine |
C12 H15 N O3 |
221.10495 |
7.297 |
104608936.7 |
(10S)-Juvenile hormone III diol |
C16 H28 O4 |
284.19811 |
9.8 |
9243187763 |
3-[(4-hydroxyphenyl)methyl]-octahydropyrrolo[1,2-a]pyrazine-1,4-dione |
C14 H16 N2 O3 |
260.11601 |
3.793 |
91656018.59 |
4-Oxo-4-(3-pyridyl)butyric acid |
C9 H9 N O3 |
179.05826 |
3.266 |
83027388.62 |
Hydrocotarnine |
C12 H15 N O3 |
221.10495 |
9.577 |
78983616.46 |
8-Methoxy-9H-carbazole-3-methanol |
C14 H13 N O2 |
227.09445 |
7.661 |
70775640.73 |
3-Hydroxy-2-methylpyridine |
C6 H7 N O |
109.0529 |
0.805 |
70199911.57 |
Bis(4-ethylbenzylidene)sorbitol |
C24 H30 O6 |
414.20365 |
11.572 |
68542901.73 |
Bicine |
C6 H13 N O4 |
163.08446 |
0.778 |
55780829.01 |
Betaine |
C5 H11 N O2 |
117.07907 |
0.787 |
52575061.91 |
DOG |
C19 H36 O5 |
344.25578 |
11.178 |
51824769.22 |
Nicotinic acid |
C6 H5 N O2 |
123.03208 |
1.041 |
50886283.96 |
Bis(7-methyloctyl) adipate |
C24 H46 O4 |
398.33885 |
17.975 |
48973391.95 |
sphinganine |
C18 H39 N O2 |
301.29781 |
10.963 |
47780018.97 |
2-(2H-Benzotriazol-2-yl)-4,6-bis(1-methyl-1-phenylethyl)phenol |
C30 H29 N3 O |
447.23055 |
17.635 |
46464220.32 |
Lauramide |
C12 H25 N O |
199.19353 |
11.894 |
44614081.14 |
NP-019811 |
C6 H7 N O2 |
125.0477 |
0.801 |
41231549.37 |
DOG |
C19 H36 O5 |
344.25573 |
12.065 |
38907325.61 |
NPPB |
C16 H16 N2 O4 |
300.11089 |
4.707 |
38197719.92 |
nylon cyclic dimer |
C12 H22 N2 O2 |
226.16808 |
2.96 |
35659163.85 |
Hydrocotarnine |
C12 H15 N O3 |
221.10505 |
5.991 |
32460753.93 |
(10S)-Juvenile hormone III diol |
C16 H28 O4 |
284.19806 |
9.35 |
32267169.01 |
Dodecylamine |
C12 H27 N |
185.21408 |
9.871 |
30473362.48 |
4-(5,6-Dihydroxy-6-methyl-1-hepten-2-yl)-1-methyl-1,2-cyclohexanediol |
C15 H28 O4 |
272.19869 |
12.647 |
30243223.53 |
Carbofuran |
C12 H15 N O3 |
221.10503 |
5.701 |
28761077.6 |
Linoleamide |
C18 H33 N O |
279.25612 |
14.229 |
28269440.94 |
1-Hydroxy-3-methoxy-10-methyl-9(10H)-acridinone |
C15 H13 N O3 |
255.08922 |
9.865 |
27738706.24 |
Nicotinic acid |
C6 H5 N O2 |
123.03208 |
0.863 |
27658406.99 |
Diethanolamine |
C4 H11 N O2 |
105.07912 |
0.755 |
27401375.74 |
(R)-Salsolinol |
C10 H13 N O2 |
179.09456 |
5.245 |
25425975.76 |
C14-Dihydroceramide |
C32 H65 N O3 |
511.49523 |
16.707 |
24986254.75 |
Norharman |
C11 H8 N2 |
168.06886 |
4.533 |
24937015.76 |
Pro-Hyp |
C10 H16 N2 O4 |
228.11101 |
2.824 |
24423909.77 |
Methylone |
C11 H13 N O3 |
207.08933 |
5.043 |
22590711.92 |
perlolyrine |
C16 H12 N2 O2 |
264.08983 |
6.031 |
22569112.9 |
MFCD03011415 |
C11 H13 N O3 |
207.08933 |
8.138 |
20960167.54 |
Azobenzene |
C12 H10 N2 |
182.08437 |
5.035 |
20841972.25 |
N-(3-acetamidopropyl)pyrrolidin-2-one |
C9 H16 N2 O2 |
184.12119 |
4.196 |
19311925.8 |
N-(3-Carboxypropanoyl)-5-hydroxynorvaline |
C9 H15 N O6 |
233.08982 |
0.771 |
18537696.93 |
Pro-tyr |
C14 H18 N2 O4 |
278.12667 |
6.116 |
18252490.47 |
Butyl 4-aminobenzoate |
C11 H15 N O2 |
193.11009 |
9.578 |
18099495.19 |
1,6-Dihydroxy-3-methoxy-10-methyl-9(10H)-acridinone |
C15 H13 N O4 |
271.08428 |
5.369 |
17408901.89 |
NSC 622050 |
C16 H31 N O3 |
285.23041 |
12.529 |
17378542.04 |
Cyclo(phenylalanyl-prolyl) |
C14 H16 N2 O2 |
244.12096 |
5.732 |
17330513.18 |
1-Methyl-1-benzylidene-acetone |
C11 H12 O |
160.08882 |
10.627 |
17308557.91 |
8-Hydroxyquinoline |
C9 H7 N O |
145.05272 |
3.371 |
16860931.4 |
Myristamide |
C14 H29 N O |
227.22493 |
13.232 |
16385290.7 |
Methylhippuric acid |
C10 H11 N O3 |
193.07395 |
4.434 |
14897657.5 |
Glycocholic acid |
C26 H43 N O6 |
465.30861 |
10.718 |
14566437.09 |
Hydrocotarnine |
C12 H15 N O3 |
221.10495 |
9.314 |
14424611.79 |
5-Hexyltetrahydro-2-furanoctanoic acid |
C18 H34 O3 |
298.25074 |
11.771 |
13879765.66 |
UX9350000 |
C6 H7 N O |
109.0529 |
7.298 |
13722770.79 |
DiisopropylSulfate |
C6 H14 O4 S |
182.06047 |
6.056 |
13213397.24 |
N-Methylanhalonine |
C13 H17 N O3 |
235.12078 |
8.53 |
12775300.85 |
propionylcarnitine |
C10 H19 N O4 |
217.13153 |
6.72 |
12160126.75 |
5-Ethyl-4-hydroxy-3(2H)-furanone |
C6 H8 O3 |
128.0475 |
1.301 |
11794106.77 |
C16-Dihydroceramide |
C34 H69 N O3 |
539.5269 |
17.555 |
11671825.44 |
L(-)-Carnitine |
C7 H15 N O3 |
161.10526 |
0.775 |
11497467.04 |
Talatizamine |
C24 H39 N O5 |
421.28227 |
10.601 |
11440823.77 |
NP-008521 |
C14 H18 N2 O2 |
246.13695 |
6.713 |
11206716.94 |
alpha-(4-methoxyphenyl)-6-methyl-2-pyridineacrylic acid |
C16 H15 N O3 |
269.10521 |
8.816 |
10847601.78 |
muramic acid |
C9 H17 N O7 |
251.10044 |
0.772 |
10840376.26 |
N,N-Bis(2-hydroxyethyl)dodecanamide |
C16 H33 N O3 |
287.24605 |
11.048 |
10651099.22 |
Maculosin |
C14 H16 N2 O3 |
260.11601 |
5.351 |
10484071.95 |
Pyrogallol |
C6 H6 O3 |
126.03181 |
0.772 |
10268268.46 |
MFCD00195543 |
C12 H17 N O2 |
207.12587 |
8.529 |
9873273.98 |
Fenspiride |
C15 H20 N2 O2 |
260.15233 |
7.457 |
9559205.054 |
1-Methyl-1-benzylidene-acetone |
C11 H12 O |
160.08882 |
13.434 |
9042713.297 |
6-Quinolinecarboxylic acid |
C10 H7 N O2 |
173.04776 |
1.193 |
9011612.395 |
N-Acetylserotonin |
C12 H14 N2 O2 |
218.10556 |
4.702 |
8931917.693 |
4-Vinylcyclohexene |
C8 H12 |
108.09405 |
7.901 |
8759114.337 |
Aplindore |
C18 H18 N2 O3 |
310.13143 |
6.162 |
8721169.072 |
3-{[4-(1,3-Benzodioxol-5-ylmethyl)piperazino]carbonyl}-6,7-dimethoxy-2H-chromen-2-one |
C24 H24 N2 O7 |
452.15953 |
11.917 |
8662511.632 |
8-Methoxy-9H-carbazole-3-methanol |
C14 H13 N O2 |
227.09477 |
9.867 |
8613086.433 |
(1E)-1-(4-Methoxyphenyl)-4-methyl-1,4-pentadien-3-one |
C13 H14 O2 |
202.09934 |
10.627 |
8092494.48 |
N,N-Dimethyldecylamine N-oxide |
C12 H27 N O |
201.20931 |
8.464 |
7884672.547 |
Crotetamide |
C12 H22 N2 O2 |
226.16799 |
7.096 |
7738760.393 |
Phenylac-gln-OH |
C13 H16 N2 O4 |
264.11095 |
5.021 |
7344209.322 |
Decanamide |
C10 H21 N O |
171.1624 |
10.314 |
6950450.272 |
MFCD03011415 |
C11 H13 N O3 |
207.08933 |
6.111 |
6550808.386 |
4-Indolecarbaldehyde |
C9 H7 N O |
145.05272 |
6.354 |
6502548.849 |
(1R)-6-Hydroxy-7-methoxy-1-methyl-1,2,3,4-tetrahydro-1-isoquinoliniumcarboxylate |
C12 H15 N O4 |
237.10005 |
4.622 |
5926711.017 |
4-tert Butylbenzoic acid |
C11 H14 O2 |
178.09949 |
6.001 |
5369139.292 |
MFCD01320532 |
C20 H37 N O3 |
339.27681 |
8.773 |
5172747.603 |
4-(2-Aminopropoxy)-3,5-dimethylphenol |
C11 H17 N O2 |
195.12609 |
8.605 |
4950057.718 |
N-Acetyl-L-phenylalanine |
C11 H13 N O3 |
207.08933 |
7.233 |
4754834.639 |
1,4-dihydroxy-1,4-dimethyl-7-(propan-2-ylidene)-decahydroazulen-6-one |
C15 H24 O3 |
252.1725 |
10.564 |
4579759.255 |
4-Vinylcyclohexene |
C8 H12 |
108.09405 |
6.729 |
4369575.34 |
4-Aminobenzoic acid |
C7 H7 N O2 |
137.04769 |
7.297 |
4332698.757 |
Crotetamide |
C12 H22 N2 O2 |
226.16799 |
7.273 |
4256784.382 |
Table 3 represents the LC-HRMS/MS profile of the crude ethyl acetate extract of S. racemosum, showing about 105 tentative compounds.
Table 4 Validation of the docking protocol and binding energies of control ligands for each target receptor.
Target Protein (PDB ID) |
Native Ligand |
Binding Energy (kcal/mol) |
RMSD (Å) |
KatG (3WXO) |
Isoniazid (prodrug) |
‒ 18.75 |
1.1464 |
KasA (2WGE) |
Thiolactomycin |
‒ 50.6 |
0.4328
|
PknB (6B2P) |
Synthetic Inhibitor |
‒ 58.95 |
0.6234
|
RNAP (5UHB) |
Rifampicin |
‒ 58.37 |
1.7937 |
QAPRTase (1QPQ) |
Quinolinic Acid |
‒ 35.8 |
1.2311 |
Table 5 Docking result of compounds identified from Syncephalastrum racemosum against selected Mycobacterium tuberculosis target proteins.
Name |
Binding Energy (kcal/mol) |
||||
KatG |
KasA |
PknB |
RNAP |
QAPRTase |
|
2-Mercaptoethanol |
‒ 10.73 |
‒ 16.98 |
‒ 14.07 |
‒ 14.95 |
‒ 4.37 |
Methylimidazoleacetic acid |
‒ 20.80 |
‒ 25.93 |
‒ 8.89 |
‒ 13.54 |
‒ 21.62 |
DA9185000 |
‒ 28.49 |
‒ 46.26 |
‒ 23.26 |
‒ 27.57 |
‒ 6.27 |
3-[(1S)-2-Cyclohexen-1-yl]-L-alanine |
‒ 25.17 |
‒ 26.02 |
‒ 21.71 |
‒ 0.63 |
‒ 3.27 |
Cyclo(phenylalanyl-prolyl) |
‒ 22.35 |
‒ 42.20 |
‒ 22.68 |
‒ 27.52 |
‒ 15.46 |
3-(propan-2-yl)-octahydropyrrolo[1,2-a]pyrazine-1,4-dione |
‒ 21.35 |
‒ 37.29 |
‒ 22.20 |
‒ 26.07 |
‒ 9.99 |
N,N-Bis(2-hydroxyethyl)dodecanamide |
‒ 25.25 |
‒ 51.86 |
‒ 34.05 |
‒ 34.10 |
‒ 15.39 |
MFCD18695608 |
‒ 23.36 |
‒ 26.21 |
‒ 33.31 |
‒ 26.69 |
‒ 18.11 |
Butyl 4-aminobenzoate |
‒ 20.66 |
‒ 39.39 |
‒ 38.66 |
‒ 27.34 |
‒ 8.72 |
2-Amino-1,3,4-octadecanetriol |
‒ 34.33 |
‒ 62.59 |
‒ 35.59 |
‒ 44.84 |
‒ 34.43 |
1-Hydroxy-3-methoxy-10-methyl-9(10H)-acridinone |
‒ 21.35 |
‒ 31.45 |
‒ 31.90 |
‒ 30.95 |
‒ 11.59 |
Bis(methylbenzylidene)sorbitol |
‒ 49.92 |
‒ 60.01 |
‒ 33.27 |
‒ 49.93 |
‒ 17.12 |
3-[(4-hydroxyphenyl)methyl]-octahydropyrrolo[1,2-a]pyrazine-1,4-dione |
‒ 28.90 |
‒ 44.86 |
‒ 32.51 |
‒ 38.57 |
‒ 31.35 |
Hydrocotarnine |
‒ 34.49 |
‒ 46.54 |
‒ 46.86 |
‒ 31.53 |
‒ 12.60 |
(10S)-Juvenile hormone III diol |
‒ 31.24 |
‒ 58.87 |
‒ 31.12 |
‒ 33.27 |
‒ 16.76 |
4-Oxo-4-(3-pyridyl)butyric acid |
3.86 |
‒ 27.75 |
‒ 25.48 |
‒ 16.92 |
‒ 17.24 |
8-Methoxy-9H-carbazole-3-methanol |
‒ 16.69 |
‒ 38.49 |
‒ 33.94 |
‒ 32.98 |
‒ 23.33 |
3-Hydroxy-2-methylpyridine |
‒ 25.24 |
‒ 41.53 |
‒ 29.59 |
‒ 29.58 |
‒ 24.18 |
Bis(4-ethylbenzylidene)sorbitol |
‒ 16.11 |
‒ 47.43 |
‒ 39.88 |
‒ 44.99 |
‒ 14.29 |
Bicine |
‒ 16.43 |
‒ 26.39 |
‒ 24.86 |
‒ 16.40 |
‒ 29.79 |
Betaine |
‒ 17.51 |
‒ 23.34 |
‒ 14.54 |
‒ 10.05 |
‒ 3.05 |
Nicotinic acid |
‒ 17.33 |
‒ 26.44 |
‒ 11.33 |
‒ 16.67 |
‒ 22.49 |
Bis(7-methyloctyl) adipate |
‒ 37.00 |
‒ 31.51 |
‒ 47.52 |
‒ 37.32 |
‒ 18.01 |
sphinganine |
‒ 41.78 |
‒ 60.05 |
‒ 37.85 |
‒ 40.27 |
‒ 23.61 |
Lauramide |
‒ 24.70 |
‒ 40.76 |
‒ 38.75 |
‒ 34.75 |
‒ 1.26 |
NPPB |
‒ 13.97 |
‒ 29.98 |
‒ 37.29 |
‒ 26.16 |
‒ 14.04 |
nylon cyclic dimer |
‒ 17.88 |
‒ 30.44 |
‒ 16.98 |
‒ 18.65 |
‒ 9.47 |
Dodecylamine |
‒ 25.16 |
‒ 37.60 |
‒ 39.42 |
‒ 31.47 |
‒ 14.85 |
4-(5,6-Dihydroxy-6-methyl-1-hepten-2-yl)-1-methyl-1,2-cyclohexanediol |
‒ 31.71 |
‒ 49.86 |
‒ 27.49 |
‒ 34.21 |
‒ 20.28 |
Carbofuran |
‒ 18.74 |
‒ 39.41 |
‒ 27.19 |
‒ 22.90 |
‒ 10.76 |
Linoleamide |
‒ 24.10 |
‒ 50.08 |
‒ 34.34 |
‒ 32.31 |
32.12 |
Diethanolamine |
‒ 30.08 |
‒ 35.51 |
‒ 23.90 |
‒ 30.36 |
‒ 24.57 |
(R)-Salsolinol |
‒ 30.21 |
‒ 48.81 |
‒ 42.66 |
‒ 31.95 |
‒ 20.59 |
Norharman |
‒ 29.11 |
‒ 48.01 |
‒ 32.02 |
‒ 24.03 |
‒ 19.26 |
Pro-Hyp |
‒ 17.64 |
‒ 35.66 |
‒ 1.79 |
‒ 20.94 |
‒ 22.22 |
perlolyrine |
‒ 31.15 |
‒ 41.93 |
‒ 44.19 |
‒ 39.01 |
‒ 22.50 |
MFCD03011415 |
‒ 6.72 |
‒ 25.66 |
‒ 22.38 |
‒ 19.24 |
‒ 14.83 |
Azobenzene |
‒ 18.00 |
‒ 42.62 |
‒ 32.02 |
‒ 28.88 |
‒ 18.73 |
N-(3-acetamidopropyl)pyrrolidin-2-one |
‒ 24.59 |
‒ 31.28 |
‒ 23.94 |
‒ 22.65 |
‒ 24.51 |
N-(3-Carboxypropanoyl)-5-hydroxynorvaline |
‒ 19.98 |
‒ 24.15 |
‒ 13.55 |
‒ 22.87 |
‒ 21.96 |
Pro-tyr |
‒ 13.20 |
‒ 32.50 |
‒ 22.75 |
‒ 23.96 |
‒ 20.82 |
1,6-Dihydroxy-3-methoxy-10-methyl-9(10H)-acridinone |
‒ 19.70 |
‒ 29.32 |
‒ 33.88 |
‒ 29.46 |
‒ 1.71 |
NSC 622050 |
‒ 2.22 |
‒ 25.24 |
‒ 29.31 |
‒ 14.42 |
9.33 |
1-Methyl-1-benzylidene-acetone |
‒ 24.29 |
‒ 45.87 |
‒ 27.70 |
‒ 27.27 |
‒ 23.50 |
8-Hydroxyquinoline |
‒ 20.35 |
‒ 36.58 |
‒ 28.78 |
‒ 25.87 |
‒ 22.55 |
Myristamide |
‒ 30.50 |
‒ 47.69 |
‒ 43.07 |
‒ 31.64 |
‒ 11.99 |
Methylhippuric acid |
5.29 |
‒ 27.02 |
‒ 18.11 |
‒ 11.08 |
‒ 27.47 |
5-Hexyltetrahydro-2-furanoctanoic acid |
‒ 9.07 |
‒ 40.29 |
‒ 28.06 |
‒ 29.38 |
‒ 25.64 |
UX9350000 |
‒ 8.49 |
‒ 25.51 |
‒ 17.48 |
‒ 16.78 |
‒ 25.97 |
DiisopropylSulfate |
‒ 18.65 |
‒ 32.73 |
‒ 26.89 |
‒ 18.12 |
‒ 17.44 |
propionylcarnitine |
‒ 23.44 |
‒ 9.38 |
‒ 9.34 |
‒ 19.17 |
1.19 |
5-Ethyl-4-hydroxy-3(2H)-furanone |
‒ 14.79 |
‒ 33.55 |
‒ 26.97 |
‒ 29.37 |
‒ 20.73 |
L(-)-Carnitine |
‒ 22.93 |
‒ 17.70 |
4.24 |
‒ 14.82 |
‒ 0.84 |
alpha-(4-methoxyphenyl)-6-methyl-2-pyridineacrylic acid |
‒ 29.88 |
‒ 38.60 |
‒ 33.96 |
‒ 31.41 |
‒ 3.67 |
muramic acid |
‒ 7.74 |
‒ 24.54 |
‒ 11.40 |
‒ 22.86 |
‒ 8.76 |
Maculosin |
‒ 26.00 |
‒ 42.69 |
‒ 33.99 |
‒ 31.90 |
9.04 |
Pyrogallol |
‒ 18.45 |
‒ 32.77 |
‒ 29.74 |
‒ 27.18 |
‒ 30.62 |
MFCD00195543 |
‒ 39.50 |
‒ 46.94 |
‒ 46.15 |
‒ 31.18 |
‒ 15.14 |
Fenspiride |
‒ 32.41 |
‒ 46.03 |
‒ 43.64 |
‒ 35.52 |
‒ 21.21 |
6-Quinolinecarboxylic acid |
‒ 8.64 |
‒ 30.58 |
‒ 14.38 |
‒ 18.86 |
‒ 17.32 |
N-Acetylserotonin |
‒ 21.07 |
‒ 21.92 |
‒ 26.40 |
‒ 29.85 |
‒ 20.04 |
4-Vinylcyclohexene |
‒ 8.72 |
‒ 27.91 |
‒ 18.37 |
‒ 22.02 |
‒ 14.82 |
Aplindore |
‒ 30.06 |
‒ 55.28 |
‒ 27.45 |
‒ 35.38 |
‒ 24.11 |
3-{[4-(1,3-Benzodioxol-5-ylmethyl)piperazino]carbonyl}-6,7-dimethoxy-2H-chromen-2-one |
‒ 42.71 |
‒ 63.08 |
‒ 51.01 |
‒ 37.63 |
‒ 29.59 |
(1E)-1-(4-Methoxyphenyl)-4-methyl-1,4-pentadien-3-one |
‒ 15.59 |
‒ 36.81 |
‒ 41.73 |
‒ 28.26 |
‒ 24.69 |
N,N-Dimethyldecylamine N-oxide |
‒ 31.71 |
‒ 30.03 |
‒ 33.33 |
‒ 28.33 |
‒ 13.69 |
Crotetamide |
‒ 27.13 |
‒ 39.95 |
‒ 22.47 |
‒ 25.55 |
‒ 16.59 |
Phenylac-gln-OH |
‒ 13.93 |
‒ 31.09 |
0.84 |
‒ 23.28 |
‒ 11.44 |
Decanamide |
‒ 20.27 |
‒ 36.75 |
‒ 43.07 |
‒ 24.50 |
‒ 9.46 |
4-Indolecarbaldehyde |
‒ 11.51 |
‒ 24.55 |
‒ 23.03 |
‒ 28.36 |
‒ 25.46 |
(1R)-6-Hydroxy-7-methoxy-1-methyl-1,2,3,4-tetrahydro-1-isoquinoliniumcarboxylate |
‒ 23.96 |
‒ 39.22 |
‒ 29.65 |
‒ 32.96 |
‒ 10.19 |
4-tert Butylbenzoic acid |
10.49 |
‒ 9.48 |
‒ 11.70 |
‒ 9.89 |
‒ 6.91 |
MFCD01320532 |
‒ 12.72 |
‒ 32.69 |
‒ 28.72 |
‒ 30.53 |
‒ 14.53 |
4-(2-Aminopropoxy)-3,5-dimethylphenol |
‒ 34.52 |
‒ 43.16 |
‒ 34.19 |
‒ 34.45 |
‒ 35.49 |
N-Acetyl-L-phenylalanine |
1.10 |
‒ 32.55 |
‒ 0.96 |
‒ 14.22 |
‒ 23.41 |
1,4-dihydroxy-1,4-dimethyl-7-(propan-2-ylidene)-decahydroazulen-6-one |
‒ 26.68 |
‒ 35.73 |
‒ 34.84 |
‒ 33.19 |
‒ 10.85 |
4-Aminobenzoic acid |
1.39 |
‒ 14.44 |
‒ 9.44 |
‒ 10.09 |
‒ 24.31 |
Table 6 Docking result of compounds identified from A. tubingensis against selected Mycobacterium tuberculosis target proteins.
Name |
Binding Energy (kcal/mol) |
||||
KatG |
KasA |
PknB |
RNAP |
QAPRTase |
|
2-Mercaptoethanol |
‒ 10.73 |
‒ 16.98 |
‒ 14.07 |
‒ 14.95 |
‒ 4.37 |
Oleamide |
‒ 24.40 |
‒ 56.13 |
‒ 52.86 |
‒ 36.89 |
4.19 |
5-Benzyldihydro-2(3H)-furanone |
‒ 28.49 |
‒ 42.59 |
‒ 29.78 |
‒ 30.95 |
‒ 26.83 |
N,N-Bis(2-hydroxyethyl)dodecanamide |
‒ 25.25 |
‒ 51.86 |
‒ 34.05 |
‒ 34.10 |
‒ 15.39 |
Cuminaldehyde |
‒ 9.12 |
‒ 39.75 |
‒ 27.53 |
‒ 26.86 |
‒ 22.73 |
Hydroxy-1,4-benzoquinone |
3.36 |
‒ 20.38 |
‒ 5.81 |
‒ 9.73 |
‒ 23.85 |
4-methyl-5-oxo-2-pentyl-2,5-dihydrofuran-3-carboxylic acid |
9.54 |
‒ 38.01 |
‒ 21.03 |
‒ 17.52 |
‒ 16.82 |
EL9600000 |
‒ 24.97 |
‒ 37.22 |
‒ 21.25 |
‒ 25.90 |
‒ 18.02 |
2-Amino-1,3,4-octadecanetriol |
‒ 34.33 |
‒ 62.59 |
‒ 35.59 |
‒ 44.84 |
‒ 34.43 |
Bis(methylbenzylidene)sorbitol |
‒ 49.92 |
‒ 60.01 |
‒ 33.27 |
‒ 49.93 |
‒ 17.12 |
Bis(4-ethylbenzylidene)sorbitol |
‒ 16.11 |
‒ 47.43 |
‒ 39.88 |
‒ 44.99 |
‒ 14.29 |
sphinganine |
‒ 41.78 |
‒ 60.05 |
‒ 37.85 |
‒ 40.27 |
‒ 23.61 |
(10S)-Juvenile hormone III diol |
‒ 31.24 |
‒ 58.87 |
‒ 31.12 |
‒ 33.27 |
‒ 16.76 |
Linoleamide |
‒ 24.10 |
‒ 50.08 |
‒ 34.34 |
‒ 32.31 |
32.12 |
nylon cyclic dimer |
‒ 17.88 |
‒ 30.44 |
‒ 16.98 |
‒ 18.65 |
‒ 9.47 |
3-Allylcyclohexene |
‒ 14.69 |
‒ 31.31 |
‒ 21.31 |
‒ 9.43 |
‒ 8.40 |
3,4-Dimethoxycinnamic acid |
‒ 2.49 |
‒ 14.89 |
‒ 22.71 |
‒ 18.94 |
‒ 17.06 |
Bicine |
‒ 16.43 |
‒ 26.39 |
‒ 24.86 |
‒ 16.40 |
‒ 29.79 |
Dodecylamine |
‒ 25.16 |
‒ 37.60 |
‒ 39.42 |
‒ 31.47 |
‒ 14.85 |
6-Hydroxy-5-methyl-4,11-dioxoundecanoic acid |
‒ 5.35 |
‒ 46.50 |
‒ 22.95 |
‒ 18.66 |
‒ 21.44 |
n-propylbenzene |
‒ 15.95 |
‒ 31.07 |
‒ 29.13 |
‒ 21.06 |
‒ 14.93 |
(2E,4E,7E)-2,4,7-Decatrienoic acid |
1.96 |
‒ 26.41 |
‒ 20.98 |
‒ 18.98 |
‒ 8.52 |
Sorbic acid |
6.90 |
‒ 12.42 |
‒ 9.69 |
‒ 3.58 |
‒ 13.23 |
3-[(4-hydroxyphenyl)methyl]-octahydropyrrolo[1,2-a]pyrazine-1,4-dione |
‒ 28.90 |
‒ 44.86 |
‒ 32.51 |
‒ 38.57 |
‒ 31.35 |
5-Ethyl-4-hydroxy-3(2H)-furanone |
‒ 14.79 |
‒ 33.55 |
‒ 26.97 |
‒ 29.37 |
‒ 20.73 |
Lauramide |
‒ 24.70 |
‒ 40.76 |
‒ 38.75 |
‒ 34.75 |
‒ 1.26 |
NSC 622050 |
‒ 2.22 |
‒ 25.24 |
‒ 29.31 |
‒ 14.42 |
9.33 |
Diethanolamine |
‒ 30.08 |
‒ 35.51 |
‒ 23.90 |
‒ 30.36 |
‒ 24.57 |
QL5300000 |
‒ 17.62 |
‒ 37.42 |
‒ 34.83 |
‒ 28.01 |
‒ 21.01 |
{4-Ethyl-6-[(3E)-2-ethyl-3-hexen-1-yl]-6-methyl-1,2-dioxan-3-yl}acetic acid |
‒ 12.91 |
‒ 37.87 |
‒ 18.45 |
‒ 21.73 |
‒ 3.40 |
Carvone |
‒ 20.63 |
‒ 47.46 |
‒ 28.69 |
‒ 25.71 |
‒ 20.54 |
1-Methyl-1-benzylidene-acetone |
‒ 24.29 |
‒ 45.87 |
‒ 27.70 |
‒ 27.27 |
‒ 23.50 |
Myristamide |
‒ 30.50 |
‒ 47.69 |
‒ 43.07 |
‒ 31.64 |
‒ 11.99 |
5-[(5-Hydroxytetrahydro-2-furanyl)methyl]-1,3-benzenediol |
‒ 24.64 |
‒ 39.11 |
‒ 39.02 |
‒ 31.04 |
‒ 28.98 |
DEET |
‒ 21.32 |
‒ 44.43 |
‒ 22.66 |
‒ 14.31 |
‒ 17.79 |
4'-Methoxyacetophenone |
‒ 11.34 |
‒ 38.97 |
‒ 27.82 |
‒ 28.62 |
‒ 27.13 |
Benzaldehyde |
‒ 11.94 |
‒ 32.14 |
‒ 24.38 |
‒ 22.71 |
‒ 27.38 |
Styrene |
‒ 11.12 |
‒ 29.69 |
‒ 28.36 |
‒ 25.43 |
‒ 16.29 |
5-(3,4-Dihydroxyphenyl)-4-hydroxypentanoic acid |
‒ 13.43 |
‒ 29.51 |
‒ 22.51 |
‒ 23.63 |
‒ 14.43 |
(2S)-4-Methyl-2-({[(3S,4S,5R)-2,3,4-trihydroxy-5-(hydroxymethyl)tetrahydro-2-furanyl]methyl}amino)pentanoic acid (non-preferred name) |
‒ 22.23 |
‒ 37.32 |
‒ 29.13 |
‒ 29.63 |
‒ 20.37 |
NP-021018 |
‒ 19.45 |
‒ 28.81 |
‒ 2.69 |
‒ 29.92 |
‒ 6.44 |
butyrin |
‒ 23.39 |
‒ 36.55 |
‒ 35.52 |
‒ 28.36 |
‒ 18.09 |
Maculosin |
‒ 26.00 |
‒ 42.69 |
‒ 33.99 |
‒ 31.90 |
9.04 |
N,N-Dimethyldecylamine N-oxide |
‒ 31.71 |
‒ 30.03 |
‒ 33.33 |
‒ 28.33 |
‒ 13.69 |
Anisole |
‒ 12.02 |
‒ 30.59 |
‒ 22.92 |
‒ 27.22 |
‒ 20.42 |
3-(propan-2-yl)-octahydropyrrolo[1,2-a]pyrazine-1,4-dione |
‒ 21.35 |
‒ 37.29 |
‒ 22.20 |
‒ 26.07 |
‒ 9.99 |
4-Phenyl-3-buten-2-one |
‒ 14.40 |
‒ 42.19 |
‒ 32.94 |
‒ 26.15 |
‒ 21.96 |
Decanamide |
‒ 20.27 |
‒ 36.75 |
‒ 43.07 |
‒ 24.50 |
‒ 9.46 |
Indole |
‒ 6.49 |
‒ 22.26 |
‒ 15.82 |
‒ 24.31 |
‒ 19.27 |
2,5-Dipropyl-4-methylthiazole |
‒ 24.95 |
‒ 31.01 |
‒ 27.95 |
‒ 23.44 |
‒ 8.69 |
Acetophenone |
‒ 9.86 |
‒ 29.81 |
‒ 26.61 |
‒ 23.78 |
‒ 26.13 |
(1E)-1-(4-Methoxyphenyl)-4-methyl-1,4-pentadien-3-one |
‒ 15.59 |
‒ 36.81 |
‒ 41.73 |
‒ 28.26 |
‒ 24.69 |
p-Hydroxybenzalacetone |
‒ 23.87 |
‒ 35.55 |
‒ 34.84 |
‒ 31.01 |
‒ 22.59 |
Tetralin |
‒ 12.92 |
‒ 36.18 |
‒ 27.53 |
‒ 21.04 |
‒ 15.77 |
Choline |
‒ 19.49 |
‒ 26.42 |
‒ 15.94 |
‒ 13.02 |
‒ 12.90 |
The tentatively identified compounds from both endophytes were subjected to molecular docking against several M. tuberculosis protein targets to understand the potential mechanisms of action of the secondary metabolites. This multi-target in silico strategy was designed to predict inhibitory pathways and identify specific receptors responsible for the observed antimycobacterial effects at the molecular level. The selected receptors in this study were KatG (PDB ID: 3WXO), KasA (PDB ID: 2WGE), PknB (PDB ID: 6B2P), RNA polymerase (RNAP; PDB ID: 5UHB), and quinolinic acid phosphoribosyltransferase (QAPRTase; PDB ID: 1QPQ).
KatG is a bifunctional catalase-peroxidase enzyme for neutralizing reactive oxygen species, i.e., hydrogen peroxide, and activating the pro-drug isoniazid (INH) [33]. KasA is a β-ketoacyl synthase involved in mycolic acid biosynthesis. Inhibition of KasA has been shown to synergize with INH, which disrupts the mycobacterial cell wall [34]. PknB, a serine/threonine protein kinase, regulates cell wall synthesis, growth, and stress adaptation, and is indispensable for bacterial viability [35]. RNAP is the transcriptional machinery of M. tuberculosis, and despite being targeted by rifampicin, it remains a clinical issue [36]. Finally, QAPRTase is an essential enzyme in the NAD biosynthesis pathway, representing a key metabolic vulnerability in M. tuberculosis.
Foregoing docking simulations with the test compounds, the protocol was validated based on the root mean squared value. In this research, the RMSD values below 2.0 Å on multiple receptors consistently confirm the reliability of the docking procedure. Specifically, isoniazid in KatG showed an RMSD of 0.40 Å with a binding energy of ‒18.75 kcal/mol, while thiolactomycin in KasA (PDB ID: 2WGE) delivered an RMSD of 0.43 Å with ‒50.60 kcal/mol. Likewise, the synthetic inhibitor of PknB (6B2P) produced an RMSD of 0.62 Å with a binding energy of ‒58.95 kcal/mol, rifampicin in RNAP (5UHB) gave 1.79 Å with ‒58.37 kcal/mol, and quinolinic acid in QAPRTase (1QPQ) achieved 1.23 Å with
‒35.80 kcal/mol. Since all RMSD values remained well below the accepted threshold of 2.0 Å, the docking protocol was considered valid and suitable for subsequent analyses [37] (Table 4).
The docking of endophytic fungal metabolites against the validated receptors revealed the target-dependent inhibitory potential. The promising candidates were chosen based on the ability to achieve binding energies superior to those of the native ligands or reference inhibitors, which means a lower binding energy, indicating better stability [37]. Based on it, the KatG and KasA emerged as the most compelling molecular targets.
Metabolites from Syncephalastrum racemosum showed bis(methylbenzylidene)sorbitol (‒49.92 kcal/mol), sphinganine (‒41.78 kcal/mol), and the chromenone derivative 3-{[4-(1,3-benzodioxol-5-ylmethyl)piperazino]carbonyl}-6,7-dimethoxy-2H-chromen-2-one (‒42.71 kcal/mol) are the top 3 compounds with the best binding affinity on the KatG receptor (Table 5). Similarly, the compounds deriving out of Aspergillus tubingensis comprehend bis(methylbenzylidene)sorbitol (‒49.92 kcal/mol), sphinganine (‒41.78 kcal/mol), and 2-amino-1,3,4-octadecanetriol (‒34.33 kcal/mol) (Table 6) showed the strongest interaction, surpassing the standard drug isoniazid (‒18.75 kcal/mol).
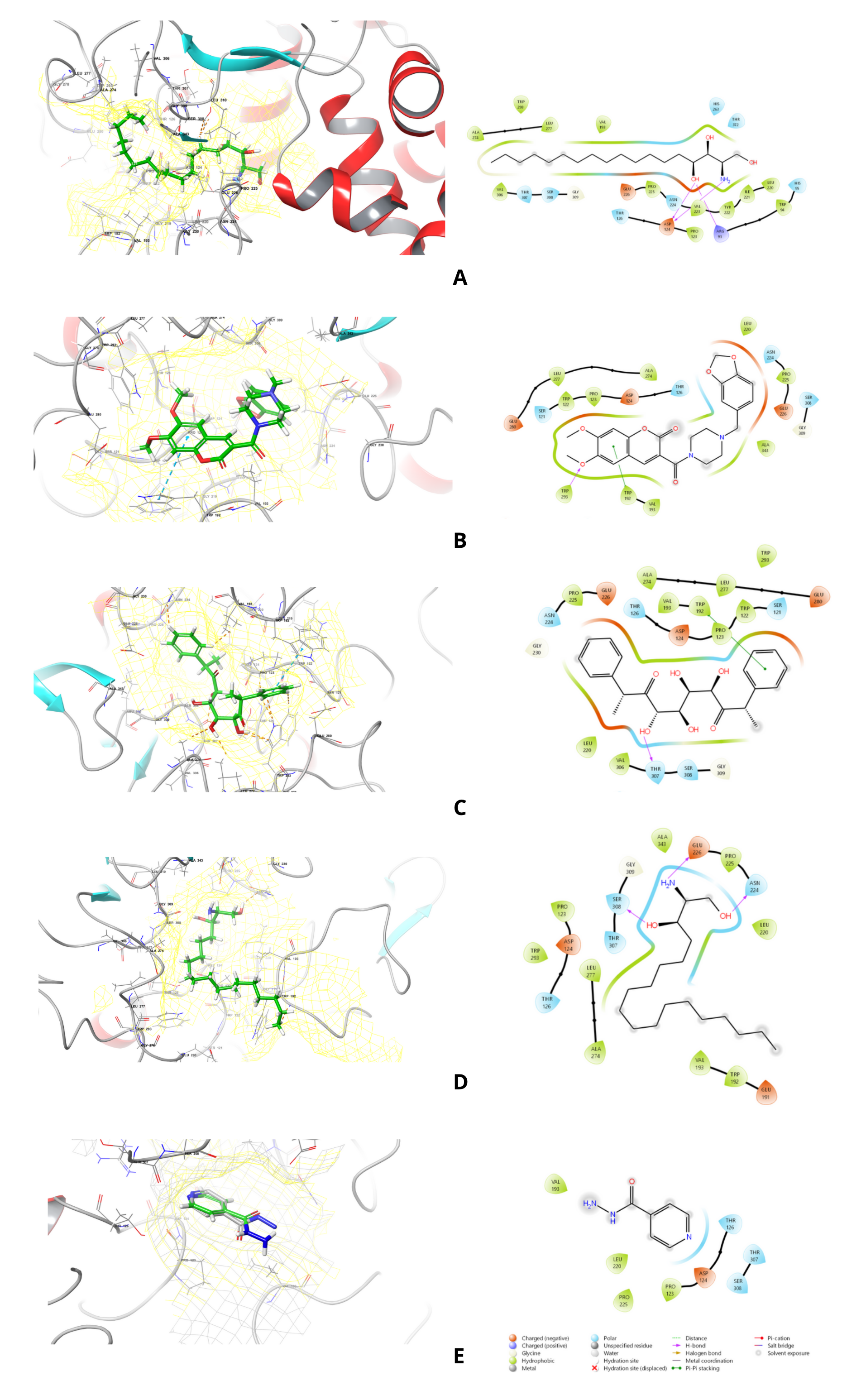
Figure 4 The binding poses of 2-amino-1,3,4-octadecanetriol (A), 3-{[4-(1,3-benzodioxol-5-ylmethyl)piperazino] carbonyl}-6,7-dimethoxy-2H-chromen-2-one (B), bis(methylbenzylidene)sorbitol (C), sphinganine (D), and the reference drug isoniazid (E) within the active site of the KatG receptor.
As shown in Figure 4(A), the binding mode of 2-amino-1,3,4-octadecanetriol in KatG is defined by 2 key features. Its polar headgroup is anchored by strong hydrogen bonds between its hydroxyl groups and residues ASP124 and ARG91. The amine moiety also contributes to hydrogen bond and electrostatic interactions with ASP124. Meanwhile, the ligand’s long aliphatic chain settles into a large hydrophobic pocket formed by VAL193, LEU220, ILE221, TYR222, VAL223, PRO225, ALA274, LEU277, TRP293, and VAL306. Π-alkyl interactions with TRP94 and TRP293 enhance this fit, while a polar contact with HIS95, HIS263, THR126, THR307, SER308, and THR372 helps to lock in its orientation. While hydrophobic packing appears to be the primary driver of affinity, the numerous polar interactions are critical for specificity, making this compound a promising candidate for KatG inhibition.
A different binding profile was observed for the 3-{[4-(1,3-benzodioxol-5-ylmethyl)piperazino]carbonyl} -6,7-dimethoxy-2H-chromen-2-one compound. This chromenone derivative achieved stable binding through a well-coordinated balance of polar and hydrophobic interactions (Figure 4(B)). The oxygen atoms of the methylenedioxy or ether bridge group formed hydrogen bonds with TRP293, while the aromatic group stands interact in π-π stacking with TRP192. Besides, another inter-electrostatic interaction (ASP124 and GLU226) stabilized the ligand via polar contacts with the ether and carbonyl oxygen atoms. The heteroaryl-piperazine ring contributed advanced interactions with ASN224 and SER308, ensuring better binding orientation. Hence, Hydrophobic residues (TRP122, VAL193, LEU220, and LEU277) encapsulate the aromatic system, which enhances the van der Waals stabilization. Synergistic π-π stacking, hydrogen bonding, and hydrophobic interactions of the ligand suggest the polycyclic scaffold may be utilized as a versatile pharmacophore for KatG-targeted inhibition.
The polyhydroxylated scaffold of bis(methylbenzylidene)sorbitol allows broader molecular interactions (Figure 4(C)). Hydrogen bonding with THR307 and additional polar contacts involving ASP124, GLU226, and GLU280 were observed at the polar site. Simultaneously, π-π stacking with TRP192 stabilizes the aromatic rings of its. Beyond that, surrounding hydrophobic residues (VAL193, LEU220, TRP122, PRO123, LEU277, TRP293, and ALA274) induce a hydrophobic enclosure that complements the aromatic moiety. Other nearby residues, including SER121, THR126, ASN224, and SER308, encourage polar contacts, which help fine-tune binding orientation. These robust molecular interactions account for its low calculated binding energy, positioning it as one of the most potent binders among the studied metabolites.
Sphinganine binds to the KatG active site using a two-model stabilization mechanism. The polar headgroup is formed by hydrogen bonds between the terminal amine and GLU226 and the hydroxyl groups with ASN224 and SER308. Since the structure of these consisted of a long aliphatic chain, the hydrophobic tail extends into a greasy pocket where van der Waals forces stabilize it from a cluster of residues (VAL193, LEU220, LEU277, ALA274, ALA343) (Figure 4(D)). The fit of this aliphatic chain is further secured by π-alkyl interactions with the aromatic residues TRP192 and TRP293, which drive strong and stable binding of sphinganine to KatG.
Consistent with its role as a drug, isoniazid displayed a weaker binding energy than the fungal metabolites. Our model shows that hydrogen bonds primarily stabilize its binding: the carbonyl oxygen interacts with ASP124, while the terminal hydrazide moiety forms contact with THR307 and SER308 (Figure 4 (E)). The position of the pyridine ring is within a pocket lined by hydrophobic residues, such as PRO123, LEU220, and VAL193, which provide additional stabilization. Notably, we observed a cation-π interaction between the pyridine ring and THR126. However, the overall number of these stabilizing hydrophobic and π-system interactions is limited compared to what we saw with the fungal metabolites, which likely explains isoniazidʼs lower binding affinity. This finding is consistent with the known mechanism of KatG, which must bind isoniazid effectively enough for activation but not so tightly as to be inhibited. Since isoniazid relies predominantly on hydrogen bonding and a single aromatic stacking interaction, fungal metabolites exploit a combination of extensive interaction types. The ability of compounds, bis(methylbenzylidene)sorbitol and sphinganine, to achieve superior stabilization suggests their potential as inhibitors that may interfere with KatG’s catalytic functions. Importantly, these interactions indicate that natural metabolites could disrupt the enzyme’s protective role in oxidative stress and activate isoniazid, offering a novel strategy for therapeutic intervention against drug-resistant Mycobacterium tuberculosis.
Broader potential of these fungal compounds from the docking result observed with KasA target showed the compound belonging to S. racemosum, 2-amino-1,3,4-octadecanetriol (‒62.59 kcal/mol), sphinganine (‒60.05 kcal/mol), and the chromenone derivative 3-{[4-(1,3-benzodioxol-5-ylmethyl)piperazino]carbonyl} -6,7-dimethoxy-2H-chromen-2-one (‒63.08 kcal/mol) (Table 4) all exhibited powerful binding affinities. Likewise, A. tubingensis metabolites 2-amino-1,3,4-octadecanetriol (‒62.59 kcal/mol), bis(methylbenzylidene)sorbitol (‒60.01 kcal/mol), and sphinganine (‒60.05 kcal/mol) (Table 5) also outperformed thiolactomycin as a native ligand (‒50.60 kcal/mol). These results indicate highly stable interactions within the KasA that inhibit mycolic acid biosynthesis in the lifecycle of M. tuberculosis.
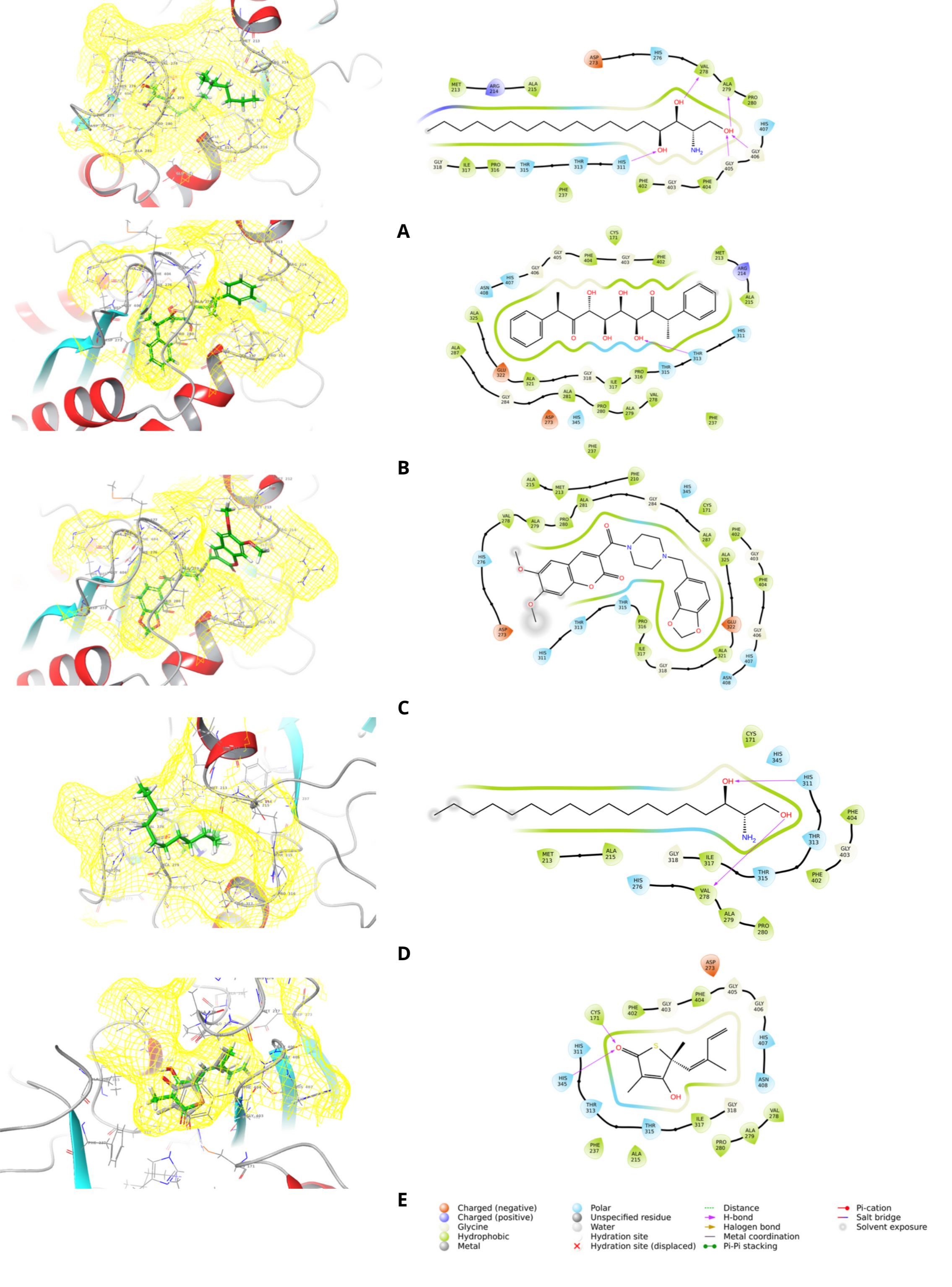
Figure 5 The binding poses of 2-amino-1,3,4-octadecanetriol (A), bis(methylbenzylidene)sorbitol (B), 3-{[4-(1,3-benzodioxol-5-ylmethyl)piperazino]carbonyl}-6,7-dimethoxy-2H-chromen-2-one (C), sphinganine (D), and the native ligand thiolactomycin (E) within the active site of the KasA receptor.
Among the ligands, 2-amino-1,3,4-octadecanetriol demonstrated a binding stabilization mechanism mediated by polar and hydrophobic interactions. Its hydroxyl groups formed directional hydrogen bonds with VAL278, ALA279, GLY405, and GLY406, while the primary amine in polar contacts with HIS311 and HIS407 residues. The expansion of the hydrophobic chain is deeply embedded among nonpolar residues (MET213, ALA215, PHE237, ILE317, PHE402, and PHE404) (Figure 5(A)). The closeness of ARG214 is facilitated through cation-π shielding. This amphiphilic binding mode, which combines polar and hydrophobic encapsulation, provides a clear structural basis for the ligandʼs binding to the KasA receptor.
The binding mode of Bis(methylbenzylidene) sorbitol is stabilized through various molecular interactions. A terminal hydroxyl group forms a hydrogen bond with THR315 within the polar entrance of the cavity. Additional stabilization was provided by HIS311, THR313, HIS345, ASN408 and HIS407, which created a complementary polar environment. The conjugated aromatic rings interact in π-π stacking and π-alkyl interactions with PHE237, PHE402, and PHE404, while also embedding within hydrophobic residues by ALA287, ALA321, ALA325, VAL278 and ILE317 (Figure 5(B)). Electrostatic steering from ASP273 and GLU322, although indirect, likely enhanced orientation. This combination of polar and aromatic contacts conferred strong affinity, suggesting that polyphenolic scaffolds could serve as potent KasA inhibitors.
3-{[4-(1,3-benzodioxol-5-ylmethyl)piperazino] carbonyl}-6,7-dimethoxy-2H-chromen-2-one, as the chromenone derivative, exhibited one of the most favorable binding energies, which was carried up by a rich interaction profile. A strong hydrogen bond with THR315 served as a principal anchoring point, while surrounding residues HIS311, THR313, HIS345, HIS407 and ASN408 contributed to a stabilizing polar environment. The extended polyaromatic system was encapsulated by residues including ALA287, ALA321, ALA325 and ILE317, with key aromatic interactions from PHE237, PHE402 and PHE404 reinforcing π-π stacking (Figure 5(C)). Although indirect, electrostatic contributions from ASP273 and GLU322 likely improved the ligand and pocket complementarity. The combination of hydrophobic encapsulation, polar stabilization, and aromatic stacking supports the interpretation of this ligand as a highly stable and promising candidate for KasA inhibition.
Sphinganine exhibited strong stabilization through its hydrophobic and polar headgroups. Its hydroxyl and amino groups formed hydrogen bonds with HIS311, VAL278 and THR315, while additional polar residues (THR313 and HIS345) enhanced orientation. The aliphatic chain was accommodated in a hydrophobic groove lined by MET213, ALA215, VAL278, ALA279 and ILE317. Although π-π stacking was absent due to its non-aromatic structure, flanking aromatic residues PHE402 and PHE404 provided shielding to the hydrophobic tail (Figure 5(D)). This amphiphilic interaction strategy underlies the strong binding of sphinganine, albeit with slightly less specificity than planar aromatic ligands.
Finally, thialactomycin, as a heteroaryl-thioether derivative, exhibited a hybrid stabilization mechanism. First, Hydrogen bonds between its hydroxyl group and HIS311/CYS171 and additional polar contacts with THR313, THR315, HIS345 and HIS407 anchored the ligand within the polar cavity. Then, its aromatic heterocyclic ring engaged in weak π interactions with PHE402 and PHE404, while the hydrophobic scaffold was embedded among ALA215, PHE237, ILE317 and PRO280. ASP273 contributed electrostatic steering, subtly enhancing orientation (Figure 5 (E)).
Taken together, the docking results demonstrate that fungal metabolites from S. racemosum and A. tubingensis display superior affinities toward KasA compared to thiolactomycin, with some ligands showing complementary interaction, which stabilizes their binding. For instance, Amphiphilic ligands 2-amino-1,3,4-octadecanetriol and sphinganine utilized dual mechanisms of hydrophobic embedding and polar anchoring, while aromatic scaffolds such as bis(methylbenzylidene)sorbitol and the chromenone derivative achieved stabilization through hydrogen bonding and π-π stacking. These findings highlight the pharmacological potential of fungal metabolites as novel inhibitors of KasA, which suggests disrupting mycolic acid biosynthesis and warrants further exploration through experimental validation and structure-activity relationship studies.
While the present investigation highlights KatG and KasA as highly promising molecular targets, it is essential to acknowledge a key limitation. We also screened the metabolites against other targets, including PknB, RNAP, and QAPRTase; however, none of these interactions achieved binding affinities superior to their respective native ligands. For this reason, our detailed interaction analyses were focused on KatG and KasA, emphasizing them as the most relevant molecular targets for the antimycobacterial potential of these fungal metabolites.
Molecular docking analysis showed that metabolites from S. racemosum and A. tubingensis have potential as promising candidates, such as sphinganine, bis(methylbenzylidene)sorbitol, 2-amino-1,3,4-octadecanetriol, and chromenone derivative 3-{[4-(1,3- benzodioxol-5-ylmethyl)piperazino]carbonyl}-6,7- dimethoxy-2H-chromen-2-one. Sphinganine, or dihydrosphingosine, is a sphingoid base that’s a key intermediate in sphingolipid metabolism. It’s structurally related to sphingosine, but lacks the trans double bond at the C4 - C5 position [38]. A previous study demonstrated that sphinganine is antibacterial against Gram-positive and Gram-negative bacteria [39,40]. As mentioned above, it is structurally related to sphingosine, which, in a previous study, was reported to be effective against Staphylococcus aureus, Streptococcus pyogenes, Micrococcus luteus, Propionibacterium acnes, Brevibacterium epidermidis, and Candida albicans, and moderately active against Pseudomonas aeruginosa [41,42]. Sphinganine lacks documented direct antimycobacterial activity, but its derivative, sphingosine, and metabolite, sphingosine 1-phosphate (S1P), have shown direct killing and immune-modulating effects against Mycobacterium tuberculosis. In a previous study, Garg et al. [43] reported that S1P showed antimycobacterial activity in vitro and in vivo against M. tuberculosis and M. smegmatis [43]. Wu et al. [44] demonstrated that sphingosine kills mycobacteria, including M. tuberculosis and M. smegmatis, and prevents human ex vivo bronchi infection with mycobacteria [44]. Our findings suggest Sphinganine may be a novel antimycobacterial agent that binds to the molecular targets KatG and KasA.
Limited scientific evidence supports using bis(methylbenzylidene)sorbitol as an antimycobacterial agent. Sorbitol derivatives are used in specific applications, while bis(methylbenzylidene)sorbitol is mainly used as a nucleating and clarifying agent for polyolefins and a stabilizer in specific pharmaceutical formulations [45]. Our findings suggest that substances can also induce antimycobacterial activity, not just as excipients.
Indraningrat et al. [46] reported that 2-amino-1,3,4-octadecanetriol exhibits antibacterial activity. This compound, a significant antibacterial component in an extract of Pseudomonas aeruginosa ISP1RL4, showed activity against multidrug-resistant bacteria, including MRSA, ESBL-producing E. coli, and ESBL-producing K. pneumoniae [46]. Our study enhances the compounds’ antimicrobial properties, including their anti-mycobacterium effects.
The chromenone derivative 3-{[4-(1,3-benzodioxol-5-ylmethyl)piperazino]carbonyl}-6,7-dimethoxy-2H-chromen-2-one shows promise as an anti-mycobacterial agent. Studies have explored similar chromone and chromanone structures for inhibiting M. tuberculosis [47,48]. Research on chromone derivatives, including those with triazole-embedded chromones, has identified compounds with significant in vitro activity against M. tuberculosis, with MIC values as low as 1.56 µg/mL [49]. A series of structurally novel spirooxindolopyrrolidine-tethered chromanones was synthesized and evaluated for their antitubercular activity against M. tuberculosis H37Rv, a clinical isolate resistant to isoniazid due to mutations in the katG and inhA promoters. The spirooxindolopyrrolidine-integrated chromanones exhibited significant activity against the tested strains [50]. Our findings indicate that the metabolites derived from both extracts of endophytic fungi with this scaffold exhibit potential as anti-tuberculosis agents.
The present study was performed in vivo and in silico; in vitro and in silico studies have limitations because they fail to replicate the intricate cellular environments, systemic interactions, and physiological rhythms of whole organisms. Consequently, they can oversimplify biological pathways. To ensure the validity of these models, future in vivo validation is essential. This involves assessing toxicity and confirming the relevance of these models to human and animal health outcomes [51,52]. Moreover, although the molecular docking analysis provided insights into potential compound-target interactions at the molecular level, these computational predictions require biochemical confirmation. Therefore, enzyme inhibition assays targeting key proteins (e.g., KatA, KasG) will be performed in future studies to validate the docking results and substantiate the computational findings experimentally.
Conclusions
Our study demonstrates the potential of an endophytic fungus, A. tubingensis and S. racemosum, isolated from H. perforata, as an antimycobacterial agent. Molecular docking analysis revealed that these compounds possess a potential dual-action mechanism by targeting two crucial enzymes in M. tuberculosis: KatG and KasA. Potential metabolites, including bis(methylbenzylidene)sorbitol, sphinganine, and a chromenone derivative, demonstrated significantly stronger binding affinities than isoniazid for KatG and thiolactomycin for KasA. These findings indicate that these fungal metabolites could disrupt the protective functions of the KatG enzyme and simultaneously inhibit the essential mycolic acid biosynthesis pathway.
Acknowledgements
The authors would like to thank The National Research and Innovation Agency (BRIN) and the Indonesia Endowment Fund for Education (LPDP) for providing the funding, Tadulako University, Tropical Disease Diagnostic Center, Universitas Airlangga, for the antimycobacterial assays, and Computational Chemistry Laboratory, Department of Pharmacy, Faculty of Mathematics and Natural Sciences, Tadulako University, for the docking experiments.
Declaration of generative AI in scientific writing
The authors acknowledge using generative AI tools (e.g., Grammarly and Writing Tools by Apple Intelligence) in preparing this manuscript, specifically for language editing and grammar correction. AI performed no content generation or data interpretation. The authors take full responsibility for the content and conclusions of this work.
CRediT author statement
Syariful Anam: Conceptualization, Methodology, Supervision, Validation, Funding acquisition, and Writing-original draft-review and editing. Armini Syamsidi: Supervision, Validation, Writing, original draft, review, and editing. Ni Luh Putu Dwijayanti: Investigation. Zidan Saputra: Investigation, Visualization. Rizqi Hulul Amalia: Investigation. Thylka Mawadha Tamagola: Investigation, Visualization. Yuliet: Supervision, writing review, and editing. Abd. Rahman Razak: Supervision, writing review, and editing. Agnes Dwi Sis Perwitasari: Methodology, Investigation, Visualization. Saipul Maulana: Methodology, Data curation, Software, Formal analysis, Writing-original draft-review and editing.
References
[1] Amr-review.org, Available at: https://amr-review.org/sites/default/files/160518_Final%20paper_with%20cover.pdf, accessed October 2025.
[2] JA Priyanto, ME Prastya, ENW Hening and RI Astuti. Exploring antimycobacterial potential and profiling secondary metabolite gene clusters in the whole genome of Bacillus isolated from dogfruit (Archidendron pauciflorum). Journal of Applied Pharmaceutical Science 2025; 15(3), 194-205.
[3] Tbindonesia, Available at: https://www.tbindonesia.or.id, accessed October 2025.
[4] Kompas, Available at: https://www.kompas.com/sains/read/2022/02/11/170500823/kemenkes-sebut-tuberkulosis-di-indonesia-masuk-3-besar-kasus-terbanyak-di?page=all, accessed October 2025.
[5] CF Deacon. Physiology and pharmacology of DPP-4 in glucose homeostasis and the treatment of Type 2 diabetes. Front Endocrinol (Lausanne) 2019; 10, 80.
[6] DJ Payne, MN Gwynn, DJ Holmes and DL Pompliano. Drugs for bad bugs: Confronting the challenges of antibacterial discovery. Nature Reviews Drug Discovery 2007; 6, 29-40.
[7] World Health Organization, Available at: https://www.who.int/publications/i/item/9789241548809, accessed October 2025.
[8] DJ Newman and GM Cragg. Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019. Journal of Natural Products 2020; 83, 770-803.
[9] D Permatasari, R Pitopang, S Anam and dan Ivan. Uji daya hambat ekstrak batang tumbuhan harrisonia perforatamerr. terhadap pertumbuhan bakteri shigella dysentriae. Biocelebes 2015; 9(1), 1-7.
[10] IO Aimang, Ramadhanil, S Anam and Ivan. Uji daya hambat daun harrisonia perforatamerr. terhadap pertumbuhan bakteri escherichia coli dan salmonella typhi. Biocelebes 2015; 9, 20-27.
[11] P Tuntiwachwuttikul, P Phansa, Y Pootaeng-On and WC Taylor. Chromones from the branches of Harrisonia perforata. Chemical and Pharmaceutical Bulletin (Tokyo) 2006; 54(1), 44-47.
[12] S Anam, A Syamsidi, Musyahidah, N Ambianti, A Widodo and MS Zubair. Isolation of endophytic fungi from benalu batu (Begonia Medicinalis) and their toxicity on Artemia Salina. Jurnal Ilmiah Farmasi (Scientific Journal of Pharmacy) 2022; 2022, 20-30.
[13] RCG Corrêa, SA Rhoden, TR Mota, JL Azevedo, JA Pamphile, CGM de Souza, M de Lourdes Teixeira de Moraes Polizeli, A Bracht and RM Peralta. Endophytic fungi: Expanding the arsenal of industrial enzyme producers. Journal of Industrial Microbiology and Biotechnology 2014; 41(10), 1467-1478.
[14] A Alvin, KI Miller and BA Neilan. Exploring the potential of endophytes from medicinal plants as sources of antimycobacterial compounds. Microbiological Research 2014; 169(7-8), 483-495.
[15] CR de Carvalho, MQ Maia, M Sobral, GMD Pereira, K da Silva, MJS Vital, JÉ Zilli, CA Rosa and LH Rosa. Diversity and antimicrobial activity of culturable endophytic fungi associated with the neotropical ethnomedicinal plants Copaifera langsdorffii and Copaifera pubiflora. South African Journal of Botany 2021; 142, 305-315.
[16] S Dettrakul, P Kittakoop, M Isaka, S Nopichai, C Suyarnsestakorn, M Tanticharoen and Y Thebtaranonth. Antimycobacterial pimarane diterpenes from the fungus Diaporthe sp. Bioorganic & Medicinal Chemistry Letters 2003; 13(7), 1253-1255.
[17] A Zerroug, N Sadrati, R Demirel, B Sabrina and D Harzallah. Antibacterial activity of endophytic fungus, Penicillium griseofulvum MPR1 isolated from medicinal plant, Mentha pulegium L. African Journal of Microbiology Research 2018; 12(48), 1056-1066.
[18] H Mohamed, W Ebrahim, M El-Neketi, MF Awad, H Zhang, Y Zhang and Y Song. In vitro phytobiological investigation of bioactive secondary metabolites from the Malus domestica-Derived endophytic fungus Aspergillus tubingensis strain AN103. Molecules 2022; 27(12), 3762.
[19] KI Tapfuma, K Nyambo, F Adu-Amankwaah, L Baatjies, L Smith, N Allie, M Keyster, AG Loxton, M Ngxande, R Malgas-Enus and V Mavumengwana. Antimycobacterial activity and molecular docking of methanolic extracts and compounds of marine fungi from Saldanha and False Bays, South Africa. Heliyon 2022; 8, 12406.
[20] WJ Andrioli, TLBV Simão, DP Ferreira, MH Araújo, SD Calixto, JK Bastos, L Seldin, E Lasunskaia and MF Muzitano. Antimycobacterial and anti-inflammatory activities of metabolites from endophytic and soil fungi. Phytomedicine Plus 2022; 2(3), 100312.
[21] M Azhari, N Merliani, M Singgih, M Arai and E Julianti. Insights into natural products from marine-derived fungi with antimycobacterial properties: Opportunities and challenges. Marine Drugs 2025; 23, 279.
[22] S Anam, A Syamsidi, MFU Tunreng, HF Djaleha, WN Arisca, G Syaputra, Iklima, MF Indriani, A Widodo,
R Pratiwi, Yuliet and AR Razak. Isolation of endophytic fungi from rui (Harrisonia perforata (Blanco) Merr.) and determining their antibacterial, antioxidant, and cytotoxic activity. HAYATI Journal of Biosciences 2024; 31(3), 443-456.
[23] TNT Hamzah, SY Lee, A Hidayat, R Terhem, I Faridah-Hanum and R Mohamed. Diversity and characterization of endophytic fungi isolated from the tropical mangrove species, Rhizophora mucronata, and identification of potential antagonists against the soil-borne fungus, Fusarium solani. Frontiers Microbiology 2018; 9, 1707.
[24] J Wang, X Wei, X Qin, X Lin, X Zhou, S Liao, B Yang, J Liu, Z Tu and Y Liu. Arthpyrones A-C, pyridone alkaloids from a sponge-derived fungus Arthrinium arundinis ZSDS1-F3. Organic Letters 2015; 17, 656-659.
[25] AN Garmana, EY Sukandar and I Fidrianny. Uji aktivitas ekstrak beberapa tumbuhan terhadap Mycobacterium tuberculosis galur sensitif dan resisten. Acta Pharmaceutica Indonesia 2011; 36(3-4), 35-39.
[26] Tuberculosis Cluster Bureau of AIDS, T., and SITs. Standard Operating Procedure for Drug Susceptibility Testing in Tuberculosis Laboratory, International Training Workshop on “Laboratory Methods for Drug Susceptibility Testing inTuberculosis”. Bangkok, 2007.
[27] H Ardalani, S Anam, KJK Kromphardt, D Staerk and KT Kongstad. Coupling microplate-based antibacterial assay with liquid chromatography for high-resolution growth inhibition profiling of crude extracts: Validation and proof-of-concept study with Staphylococcus aureus. Molecules 2021; 26(6), 1550.
[28] KT Kongstad, SG Wubshet, A Johannesen, L Kjellerup, A-ML Winther, AK Jäger and D Staerk. High-resolution screening combined with HPLC-HRMS-SPE-NMR for identification of fungal plasma membrane H+-ATPase inhibitors from plants. Journal of Agricultural and Food Chemistry 2014; 62(24), 5595-5602.
[29] N Fabre, I Rustan, E de Hoffmann and J Quetin-Leclercq. Determination of flavone, flavonol, and flavanone aglycones by negative ion liquid chromatography electrospray ion trap mass spectrometry. Journal of the American Society for Mass Spectrometry 2001; 12, 707-715.
[30] A Hagag, MF Abdelwahab, AMA El-Kader and MA Fouad. The endophytic aspergillus strains: A bountiful source of natural products. Journal of Applied Microbiology 2022; 132, 4150-4169.
[31] AE Rahmawati, F Romadhonsyah, BM Gemantari, A Nurrochmad, S Wahyuono and P Astuti. Effect of light exposure on secondary metabolite production and bioactivities of Syncephalastrum racemosum endophyte. Tropical Journal of Natural Product Research 2021; 5, 312-318.
[32] MC Ferreira, CL Cantrell, DE Wedge, VN Gonçalves, MR Jacob, S Khan, CA Rosa and LH Rosa. Antimycobacterial and antimalarial activities of endophytic fungi associated with the ancient and narrowly endemic neotropical plant Vellozia gigantea from Brazil. Memórias do Instituto Oswaldo Cruz 2017; 112(10), 692-697.
[33] J Li, R Duan, ES Traore, RC Nguyen, I Davis, WP Griffth, DC Goodwin, AA Jarzecki and A Liu. Indole N-Linked hydroperoxyl adduct of protein-derived cofactor modulating catalase-peroxidase functions. Angewandte Chemie International Edition 2024; 63, 202407018.
[34] P Kumar, GC Capodagli, D Awasthi, R Shrestha, K Maharaja, P Sukheja, S-G Li, D Inoyama, M Zimmerman, HPH Liang, J Sarathy, M Mina, G Rasic, R Russo, AL Perryman, T Richmann, A Gupta, E Singleton, S Verma, S Husain, ... , D Alland. Synergistic lethality of a binary inhibitor of mycobacterium tuberculosis KasA. mBio 2018; 9(6), 02101-02117.
[35] O Burastero, M Cabrera, ED Lopez, LA Defelipe, JP Arcon, R Durán, MA Marti and AG Turjanski. Specificity and reactivity of Mycobacterium tuberculosis serine/threonine kinases PknG and PknB. Journal of Chemical Information and Modeling 2022; 62(7), 1723-1733.
[36] AL Garner, J Rammohan, JP Huynh, LM Onder, J Chen, B Bae, D Jensen, LA Weiss, AR Manzano, SA Darst, EA Campbell, BE Nickels, EA Galburt and CL Stallings. Effects of increasing the affinity of CarD for RNA polymerase on mycobacterium tuberculosis growth, rRNA transcription, and virulence. Journal of Bacteriology 2017; 199(4), 0069816
[37] J Ekowati, BA Tejo, S Maulana, WA Kusuma, R Fatriani, NS Ramadhanti, N Norhayati, KA Nofianti,
MI Sulistyowaty, MS Zubair, T Yamauchi and IS Hamid. potential utilization of phenolic acid compounds as anti-inflammatory agents through TNF-α convertase inhibition mechanisms: A network pharmacology, docking, and molecular dynamics approach. ACS Omega 2023; 8(49), 46851-46868.
[38] J Fantini and FJ Barrantes. Sphingolipid/cholesterol regulation of neurotransmitter receptor conformation and function. Biochimica et Biophysica Acta (BBA) - Biomembranes 2009; 1788(11), 2345-2361.
[39] TC Kunz and V Kozjak-Pavlovic. Diverse facets of sphingolipid involvement in bacterial infections. Frontiers in Cell and Developmental Biology 2019; 7, 203.
[40] S Sholekha, S Budiarti, AEZ Hasan, NPRA Krishanti and AT Wahyudi. Antimicrobial potential of an actinomycete Gordonia terrae JSN1.9-Derived orange pigment extrac. HAYATI Journal of Biosciences 2024; 31(1), 161-170.
[41] DJ Bibel, R Aly and HR Shinefield. Antimicrobial activity of sphingosines. Journal of Investigative Dermatology 1992; 98(3), 269-273.
[42] Y Wu, Y Liu, E Gulbins and H Grassmé. The anti-infectious role of sphingosine in microbial diseases. Cells 2021; 10(5), 1105.
[43] SK Garg, E Volpe, G Palmieri, M Mattei, D Galati, A Martino, MS Piccioni, E Valente, E Bonanno, PD Vito, PM Baldini, LG Spagnoli, V Colizzi and M Fraziano. Sphingosine 1-phosphate induces antimicrobial activity both in vitro and in vivo. The Journal of Infectious Diseases 2004; 189(11), 2129-2138.
[44] Y Wu, F Schnitker, Y Liu, S Keitsch, F Caicci, F Schumacher, A Riehle, B Pollmeier, J Kehrmann, B Kleuser, M Kamler, I Szabo, H Grassmé and E Gulbins. Sphingosine kills Mycobacteria and suppresses mycobacterial lung infections. Journal of Molecular Medicine 2025; 103(5), 547-558.
[45] Bis(methylbenzylidene)sorbitol, Available at: https://pubchem.ncbi.nlm.nih.gov/compound/Bis_methylbenzylidene_sorbitol, accessed October 2025.
[46] AAG Indraningrat, PPCP Purnami, E Damayanti, MD Wijaya, DAPS Masyeni and NLPEK Sari. Antibacterial potential of pseudomonas aeruginosa ISP1RL4 isolated from seaweed eucheuma cottonii against multidrug-resistant bacteria. Biomedical & Pharmacology Journal 2024; 17(4), 2341-2354.
[47] RS Keri, S Budagumpi, RK Pai and RG Balakrishna. Chromones as a privileged scaffold in drug discovery: A review. European Journal of Medicinal Chemistry 2014; 78, 340-374.
[48] S Kamboj and R Singh. Chromanone-A prerogative therapeutic scaffold: An overview. Arabian Journal for Science and Engineering 2022; 47(1), 75-111.
[49] V Nalla, A Shaikh, S Bapat, R Vyas, M Karthikeyan, P Yogeeswari, D Sriram and M Muthukrishnan. Identification of potent chromone embedded [1,2,3]-triazoles as novel anti-tubercular agents. Royal Society Open Science 2018; 5(4), 171750.
[50] MF Alkaltham, AI Almansour, N Arumugam, SK Vagolu, T Tønjum, SI Alaqeel, S Rajaratnam and
V Sivaramakrishnan. Activity against Mycobacterium tuberculosis of a new class of spirooxindolopyrrolidine embedded chromanone hybrid heterocycles. RSC Advances 2024; 14(17), 11604-11613.
[51] O Graudejus, RDP Wong, N Varghese, S Wagner and B Morrison. Bridging the gap between in vivo and in vitro research: Reproducing in vitro the mechanical and electrical environment of cells in vivo. Frontiers in Cellular Neuroscience 2018; 12.
[52] E Madorran, A Stožer, S Bevc and U Maver. In vitro toxicity model: Upgrades to bridge the gap between preclinical and clinical research. Bosnian Journal of Basic Medical Sciences 2020; 20(2), 157-168.