
Trends
Sci.
2026;
23(6):
12188
Antimicrobial Potential and Metabolite Profiling in the Bioprospecting of Endophytic Botryosphaeria rhodina from Mangrove Xylocarpus granatum
Rudi Hendra1,4,*, Muhammad Rohim1, Ari Satia Nugraha2, Fauzan Zein Muttaqin3, Hilwan Yuda Teruna1,4 and Yuli Haryani1,4
1Department of Chemistry, Faculty of Mathematics and Natural Sciences, Universitas Riau,
Pekanbaru 28293, Indonesia
2Drug Utilisation and Discovery Research Group, Faculty of Pharmacy, University of Jember,
Jember 68121, Indonesia
3Faculty of Pharmacy, Bhakti Kencana University, Bandung 40614, Indonesia
4Centre of Biological Innovation for Regenerative and Natural Applications, Universitas Riau,
Pekanbaru 28293, Indonesia
(*Corresponding author’s e-mail: [email protected])
Received: 6 October 2025, Revised: 6 November 2025, Accepted: 13 November 2025, Published: 10 February 2026
Abstract
This research examines the antibacterial properties and metabolite diversity of Botryosphaeria rhodina, an endophytic fungus sourced from mangrove environments in Riau, Indonesia. The aim was to assess its antibacterial and antifungal properties and to characterize its metabolite composition for bioprospecting purposes. Solid-state fermentation was performed on a rice substrate, and crude extracts were partitioned into methanol and n-hexane fractions. The antimicrobial effectiveness was evaluated against eight bacterial and three fungus strains utilizing well diffusion and broth microdilution techniques. The methanol extract showed strong antibacterial activity against Listeria monocytogenes (inhibition zone: 28.2 mm; Minimum Inhibition Concentration (MIC): 31.25 ppm) and moderate antifungal effects against Candida albicans, C. glabrata, and C. krusei. The n-hexane extract exhibited minimal action. Metabolite profiling of the methanol extract via liquid chromatography-high resolution mass spectrometry identified over sixty compounds, including kojic acid, glycitein, fraxetin, deoxybrevianamide E, and usnic acid, whereas gas chromatography-mass spectrometry of the n-hexane extracts predominantly revealed fatty acids and sterols. Over twenty-five metabolites are uncharacterized, indicating the existence of unique chemical scaffolds. Partial Least Squares Discriminant Analysis established that polar metabolites were the principal contributors to antibacterial action. These findings underscore B. rhodina as a renewable microbial resource with considerable potential for sustainable drug discovery.
Keywords: Antimicrobial activity, Botryosphaeria rhodina, Endophytic fungi, Mangrove-derived natural products, Metabolite profiling, Solid-state fermentation, HRMS, GCMS
Introduction
Endophytic fungi are a polyphyletic group of highly diverse ascomycetous fungi that live within plant tissues for a while without causing apparent effects. The fungi are known to be widespread among terrestrial plants, with some plants hosting hundreds of them. Endophyte-host interactions range from mutualism to parasitism depending on endophyte transmission,
infection pattern, hostage, genetic background, and environmental factors [1]. Endophytic fungi can stimulate plant growth, increase resistance to disease-causing pathogens, suppress weeds, and increase tolerance to abiotic and biotic stresses. They also can potentially produce vast bioactive secondary metabolites with pharmaceutical importance, such as antibacterial, antifungal, and anticancer compounds [2].
Due to the unique ecosystem, endophytic fungi associated with mangrove plants are particularly interesting. Mangrove forests are a dynamic transition zone between terrestrial and marine habitats, commonly found along tropical and subtropical coastlines. Endophytic fungi from mangroves are promising sources of structurally unique natural products and drug leads with a wide range of bioactivities. The mangrove fungus may produce novel bioactive compounds like citrofulvicin, chrysogenester, brocapyrrozin A, and simpterpenoid A for antiosteoporotic, anti-inflammatory, antibacterial, and antivirus, respectively [3-6].
Mangroves are found in tropical and subtropical climates, with 41.4% of global mangroves in South and Southeast Asia. The Rhizophoraceae family includes four genera: Bruguiera, Ceriops, Kandelia, and Rhizophora [7]. Arobaya and Wanma reported Indonesia has 27% of the worldʼs mangrove forest, or 4.25 million ha, with 143 billion ha in Riau Province [8]. The isolation and characterization of endophytic fungi from mangroves in Indonesia have been reported. More precisely, eight fungal strains were obtained from the mangrove species Sonneratia griffithii Kurz, located on the coast of West Sumatra. Certain ethyl acetate extracts derived from these fungi demonstrated antibacterial efficacy against Staphylococcus aureus and Escherichia coli [9]. Moreover, the antimicrobial properties of kojic acid have been isolated from the endophytic fungus Aspergillus flavus, which was obtained from the leaves of the mangrove plant Sonneratia alba on Timor Island, Indonesia. Kojic acid is widely used in the cosmetics and pharmaceutical industries due to its antibacterial properties and ability to protect against UV B radiation [10].
Our research team isolated twenty-eight fungi from two mangrove species (Bruguiera sp. and Ceriops tagal) in Sei Pakning and Tenggayun Beach, Riau Province. Among these fungi were Trichoderma sp. and Penicillium sp. Analogous assays against S. aureus and E. coli were employed to ascertain the antibacterial properties of these fungi. The findings revealed that Trichoderma sp. and Penicillium sp. demonstrated the most potent antibacterial properties [11]. In addition, Aspergillus sp., which has been isolated from the species of the mangrove, showed anti-vibriosis activity against Vibrio alginolyticus and V. parahaemolyticus by using an antagonistic study [12].
For our ongoing investigation, we obtained 15 endophytic fungi from seven distinct mangrove species gathered from various locations in Riau Province. This study builds upon our prior research. The antagonistic activity of these fungi was assessed against various bacterial pathogens, such as E. coli, V. parahaemolyticus, V. alginolyticus, Bacillus subtilis, and S. aureus. Out of the isolated fungi, three strains displayed the highest level of effectiveness in inhibiting the growth of bacteria. By employing PCR amplification using ITS4 and ITS5 primers, we successfully identified the strains Fusarium equiseti, Aspergillus fumigatus, and Botryosphaeria rhodina based on the ribosomal region of the internal transcribed spacer (ITS).
Despite the growing acknowledgment of mangrove-associated endophytic fungi as significant sources of pharmacologically active secondary metabolites, the antibacterial properties and metabolite profile of B. rhodina remain predominantly unexamined. Prior research has predominantly concentrated on genera like Aspergillus, Trichoderma, and Penicillium, resulting in a paucity of extensive studies on B. rhodina, especially within Indonesian mangrove habitats. This knowledge deficiency constrains our comprehension of the speciesʼ metabolic capabilities and its prospective contributions to antimicrobial medication development. We propose that the secondary metabolites generated by B. rhodina have significant antibacterial and antifungal activity, due to a chemically varied assortment of polar and non-polar bioactive compounds. This work was conducted to assess the antibacterial properties of B. rhodina extracts and to execute detailed metabolite profiling utilizing liquid chromatography-tandem mass spectrometry (LC-MS/MS) and gas chromatography-mass spectrometry (GC-MS).
Materials and methods
General experimental procedure
Analytical grade solvents including methanol, n-hexane, ethyl acetate, and formic acid were obtained from Merck (Germany) and Sigma-Aldrich (USA). Culture media such as PDA (Potato Dextrose Agar), PDB (Potato Dextrose Broth), SDA (Sabouraud Dextrose Agar), SDB (Sabouraud Dextrose Broth), NB (Nutrient Broth), and MHA (Mueller-Hinton Agar) were purchased from HiMedia (India). The bacterial strains used included Staphylococcus aureus ATCC 6538, Bacillus subtilis ATCC 19659, B. cereus ATCC 11778, Listeria monocytogenes ATCC 7644, Escherichia coli ATCC 25922, Salmonella typhimurium ATCC 14028, Vibrio parahaemolyticus ATCC 17802, and V. alginolyticus ATCC 17749. The fungal strains were Candida albicans ATCC 10231, C. glabrata ATCC 15126, and C. krusei ATCC 14243. Crude extracts were concentrated using a rotary evaporator (Buchi Rotavapor R-300, Switzerland). Ultrasonic extraction was conducted using a Branson Ultrasonicator (USA). Metabolite profiling of methanol extract was performed using a UHPLC-Q Exactive Plus Orbitrap LC-MS/MS system (Thermo Scientific, USA), and n-hexane extract was analysed using a GCMS-QP2010S (Shimadzu, Japan). Data analysis was conducted using Compound Discoverer™ 3.3 (Thermo Fisher Scientific) and GraphPad Prism v9 (GraphPad Software, USA).
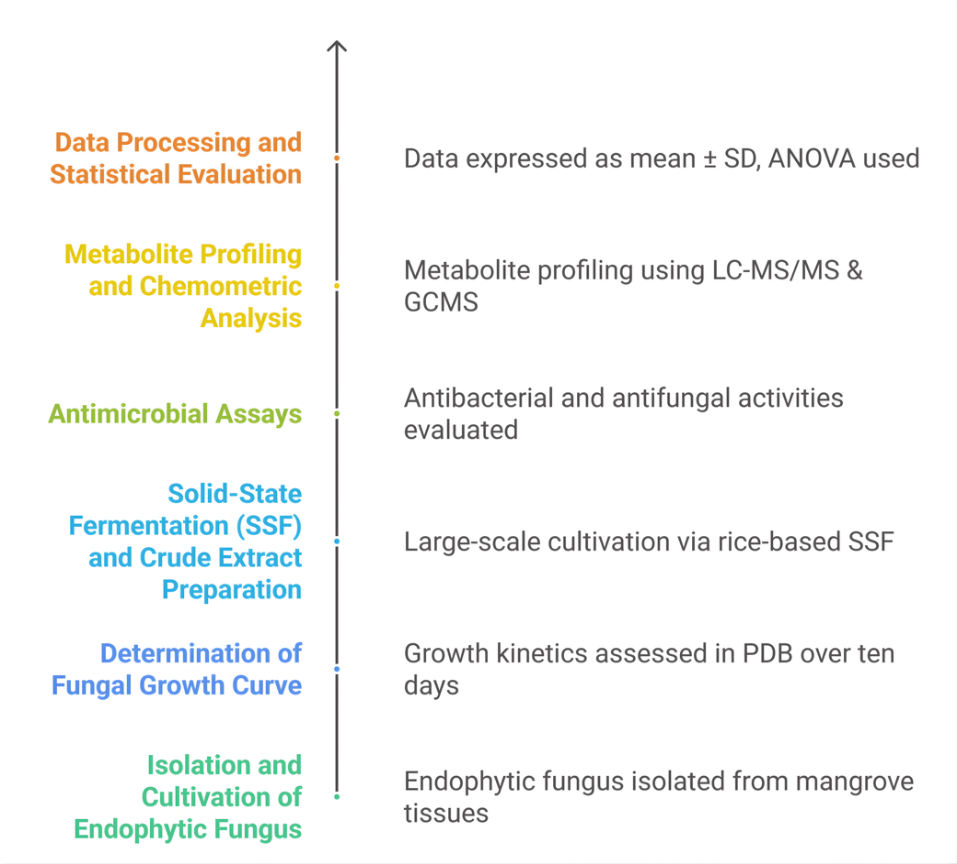
Figure 1 Experimental workflow for the bioprospecting of endophytic botryosphaeria rhodian propagation of endophytic fungi.
B. rhodina isolates were revitalized employing aseptic techniques. This study employs the direct plating technique, which entails inoculating endophytic fungus into test tubes with slanted agar media and Petri dishes to establish a stock and working culture. The PDA medium was employed. This process involves the introduction of endophytic fungus into a culture medium via the streak method and an inoculation loop. The streaked endophytic fungi were sterilized in an incubator with a 70% alcohol solution for 3 to 4 days. Certain endophytic fungi from the Petri plates and test tubes were employed for active cultivation. The remaining fraction was maintained as a stock culture at 4 °C in a refrigerator. The endophytic fungi from the Petri dishes were transferred to test tubes employing the streak method and an inoculation loop. The test tubes were incubated for a duration of 3 to 4 days. The endophytic fungi from each test tube were transferred to a Petri dish utilizing the spread method and a spreader. The Petri dish is incubated for a duration of 3 to 4 days.
Growth curve of endophytic fungi
The growth curve of B. rhodina was determined by inoculating the fungi into test tubes containing PDB medium. The inoculation was performed utilizing the plug method, in which the fungi were transported from Petri dishes to test tubes. The test tubes were subsequently incubated at room temperature (about 25 - 30 °C) in an incubator. In the preliminary incubation phase, a solitary endophytic fungus was isolated from each test tube and subsequently filtered by employing a vacuum filter with filter paper. Thereafter, the filtered fungus was placed in a desiccator, and their weight was recorded. The procedure was conducted daily (every 24 h) by observing the growth of the endophytic fungal isolates and measuring the changes in their dry cell weight. The measurement process ceased when the weight of the dry cell began to decrease. The weight data of the dried cells is later evaluated to ascertain the growth curve of B. Rhodia. The growth curve of B. rhodina was determined in triplicate, and the results are expressed as mean ± standard deviation (SD). The dry cell weight was recorded daily for ten days to evaluate biomass accumulation. The mean values and SDs were plotted to construct the growth curve. Statistical validation confirmed the consistency of replicating data.
Fermentation and crude extract preparation
The fermentation of endophytic fungus was performed on solid media utilizing thirty-one 1 L Erlenmeyer flasks. Each flask contained 200 g of rice and 200 mL of sterile water. The rice in the Erlenmeyer flasks was sterilized in an autoclave at 121 °C and 1 atm pressure for 20 min to yield cooked rice. Colonies of endophytic fungi from Petri dishes were introduced into sterile water and homogenized with an inoculation loop. The suspension was thereafter inoculated uniformly onto the rice surface utilizing a micropipette. The inoculation media were incubated at ambient temperature (about 25 - 30 °C) in a sterile incubator until the stationary phase was attained. [13]
The fermented culture attains the stationary phase and is then extracted by the maceration process with ethyl acetate as the solvent. The maceration extract was concentrated using rotary evaporator at 40 °C. The crude ethyl acetate extract was subsequently partitioned via a liquid-liquid extraction technique. The crude ethyl acetate extract was solubilized in methanol. Partitioning was sustained utilizing n-hexane as the solvent in a 1:1 ratio until no additional components were soluble in the non-polar phase. The n-hexane and methanol extracts were segregated into distinct containers, and the overall yield was determined.
Antibacterial activity using well diffusion method
The antibacterial efficacy of methanol extracts and fractions from endophytic fungus was evaluated against 4 Gram-positive pathogenic bacteria. Each pathogen was reactivated in nutritional broth and subsequently diluted to an optical density of approximately 0.1 at 600 nm. They were subsequently put to the sterile MHA medium. A cork borer was used to make 6 mm in the MHA. Subsequently, 100 μL of extracts, a positive control (chloramphenicol at a concentration of 30 ppm), and a negative control (distilled water) were added to the MHA wells. The evaluated endophytic fungal extracts included a methanol extract at a concentration of 500 ppm and a methanol fraction at 300 ppm. The test media were then incubated at 37 °C for 12 h, with variations based on the individual pathogen being studied. The resultant zones of inhibition were further measured. The procedures were executed utilizing aseptic protocols and were replicated thrice [14,15]. All experiments were performed in triplicate (n = 3), and the inhibition zones were expressed as mean ± SD. The differences between treatments were statistically analyzed by one-way ANOVA and Tukey’s post-hoc test (p < 0.05).
The Minimum Inhibitory Concentration (MIC) and Minimum Bactericidal Concentration (MBC)
The antibacterial effectiveness of pure chemicals obtained from the methanol extract of the endophytic fungus B. rhodina was evaluated using the microdilution method. The microplate wells were augmented with extracts at a concentration of 500 ppm and a positive control of chloramphenicol at 30 ppm. Each well received 50 microliters of MHB, followed by successive dilutions to achieve compound concentrations of 50 ppm and chloramphenicol concentrations of 15 ppm. Each well was administered a total of 30 microliters of medium and 10 microliters of resazurin. Each pathogen, at a concentration of roughly 0.1 OD600, was introduced into each well in a volume of 10 μL, excluding the media control. The samples were subsequently incubated at 37 °C for 12 h. Subsequent to incubation, 10 μL of each chemical concentration, along with positive and negative controls, were transferred to MHA medium. The mixture was subsequently incubated for 12 h, contingent upon the specific pathogen under examination. Bacterial proliferation was subsequently noted [14,15].
Antifungal activity using well diffusion method
The SDA was precisely dispensed into sterile petri dishes following sterilization, and the mixture was allowed to solidify. The surface of the SDA with fungi was uniformly distributed with an optical density of approximately 0.05 at 530 nm. This was accomplished with the assistance of a sterile cotton swab. The antifungal properties of the fungi-infested SDA were assessed by intentionally placing blank disc papers on its surface. The test extract was administered dropwise onto the blank discs using a micropipette containing 10 µL of the extract solution, resulting in a final concentration of 1,000 µg/disc. Ketoconazole was administered at a concentration of 30 µg/mL. The negative control discs were subjected to a comparable methodology. The solutions on the discs were then permitted to cure naturally after being exposed to them for an extended period.
Incubation of the petri dishes was conducted at a consistent temperature of 37 °C for a period of 18 to 24 h following the preparation protocol. The fungal culture and the administered substances interacted efficiently during this period. Post-incubation, the petri dishes were examined for clear zones surrounding the discs, which suggested that fungal growth had been suppressed. The level of antifungal activity was precisely measured and recorded in relation to the diameters of these transparent regions. In order to ensure the reliability and consistency of the results, the test was conducted three times for each fungal strain. The samples exhibited distinct inhibition zones, which suggested the antifungal efficacy [16].
Minimum Inhibitory Concentration (MIC) and Minimum Fungicidal Concentration (MFC)
The minimum inhibitory concentration (MIC) and minimum fungicidal concentration (MFC) were determined using the microdilution technique and a 96-well microplate. At 530 nm, the optical density of C. albicans, C. glabrata, and C. krusei cultures was adjusted to approximately 0.05 per cent. Additionally, SDA plates were implemented to identify the MFC in accordance with Hendra et al. [17] and Agustha et al. [18].
Secondary metabolites profiling
LC-MS/MS and GC-MS were employed to identify the secondary metabolites of the extracts. The LC-MS/MS was employed to analyze the methanol extract. The desiccated extract was subjected to ultrasonic extraction using a Branson Ultrasonic Corporation device located in Danbury, CT, USA. A mixture of 1.5 mL of methanol and approximately 50 mg of the extract was subjected to ultrasonic extraction at a temperature of 30 °C for a duration of 15 min. The resulting mixture was subsequently filtered through a 0.2 μm syringe filter membrane produced by SY25TF PTFE mdi. Small vials were employed to capture the liquid that passed through the filter. The metabolites were analyzed using a UHPLC-Q Exactive Plus Orbitrap High-Resolution Mass Spectrometer from Thermo Scientific™ in Waltham, MA, USA, which was coupled with an Accucore™ Phenyl Hexyl (100×2.1 mm, 2.6 μm) separation column. A UV detector was employed, with a wavelength of 254 nm. The ionization source for the MS analysis was the electrospray ionization method. The Q-Orbitrap mass analyzer was employed, and collision energies of 18, 35 and 53 eV were implemented. The spray voltage was maintained at 3.8 kV, the capillary temperature was maintained at approximately 320 °C, the sheath gas flow rate was 15 mL/min, the auxiliary gas flow rate was 3 mL/min, and the resolving power was 70,000 FWHM. The autosampler temperature was maintained at 10 °C, the sample injection volume was 0.5 μL, and the flow rate of the delivery system was adjusted to 0.3 mL/min. The mobile phase was composed of acetonitrile (B) and a solution containing 0.1% formic acid in water (A). The linear gradient elution program was implemented in the following manner: The concentration of B is 5% for the initial 0 - 1.5 min. The concentration of B increases from 5% - 10% over the course of 1.5 - 9 min. The concentration of B increases from 10% - 20% between 9 and 13 min. The concentration of B increases from 20% - 28% between 13 and 17 min. The concentration of B increases from 28% - 78% between 17 and 23 min. The concentration of B decreases from 70% - 95% between 23 and 26 min. The concentration of B remains at 95% for 26 - 29 min. Ultimately, the concentration of B decreases from 95% - 5% between 29 and 32 min. The relative abundance ranged from 0 - 100, and the exercise lasted a total of 32 min. The MS full-scan type, which operates within the mass range of 100 - 1,500 m/z, was implemented in both positive and negative ionization modes.
The Shimadzu GCMS-QP2010S instrument was used to analyze the hexane extract using gas chromatography-mass spectrometry (GC-MS). The Agilent DB-5MS UI column, which has an inner diameter of 0.25 mm and a film thickness of 0.25 µm, was used in the analysis. The column is 30 meters in length. Helium served as the carrier gas. Electron Ionization (EI) was employed to conduct the ionization process at an energy level of 70 electron volts (eV). The gas chromatography conditions were meticulously established, beginning with an initial column oven temperature of 70.0 °C and maintaining it for 5.00 min. The temperature was gradually increased at a rate of 5.00 °C per minute until it attained a final temperature of 305.0 °C. Subsequently, the temperature was maintained at this level for precisely 18 min. The injection parameters consisted of a split injection mode at a temperature of 300.0 °C and a split ratio of 49.0. The flow control was effectively maintained in pressure mode, with a total flow rate of 35.6 mL/min and a column flow rate of 0.65 mL/min. The linear velocity was 29.6 cm/s as a result of the pressure adjustment to 30.0 kilopascals. The discharge rate of the purge was established at 3.0 mL/min. In order to ensure the systemʼs readiness, the column oven, SPL1, and MS units were examined, and the SPL1 carrier and purge flow were subjected to supplementary inspections.
For mass spectrometry, the interface temperature was maintained at 305.0 °C, while the ion source temperature was maintained at 250.0 °C. The acquisition parameters were set to commence at 3.20 min and conclude at 70.00 min in scan mode, with a scan speed of 1,250 and an event time of 0.50 s. The scanning mass range was established as 28.00 to 600.00 m/z. The solvent cutting process was scheduled to last for 3 min. The detector was operated in absolute gain mode, with a detector gain of 1.50 kV and a threshold of 0. In order to ensure the precision and stability of the measurements, the system implemented a 3.0-minute equilibrium period.
Data analysis.
The mean value plus or minus the standard deviation (SD) was used to convey the antimicrobial activity results obtained using the well diffusion method. GraphPad Prism Version 9 was employed to analyse the data. The means were contrasted using a one-way analysis of variance (one-way ANOVA), followed by Tukeyʼs test. Statistically significant values were defined as those with a p < 0.05, Compound Discoverer 3.3 software (Thermo Fisher Scientific) was employed to analyze mass spectrometry data. At a mass tolerance of 5 ppm, features were identified and categorized by retention duration, with a tolerance of 0.2 min. The ChemSpider database was employed to identify the compounds. The mzCloud database was used to correlate MS2 spectra in order to improve chemical annotation. A multivariate chemometric study was conducted utilizing the online MetaboAnalyst 5.0 platform (www.metaboanalyst.ca). Partial Least Squares Discriminant Analysis (PLS-DA) was utilized to distinguish the metabolite profiles of methanol and n-hexane extracts using LC-HRMS and GC-MS datasets. Variable Importance in Projection (VIP) scores were computed to ascertain the discriminant metabolites that most significantly contribute to class differentiation. Score plots and heatmaps were created to illustrate clustering tendencies in bacterial and fungal bioassays.
Results and discussion
Growth curve of endophytic fungi
Understanding fungal growth kinetics is crucial to optimize secondary metabolite production. This work constructed the growth curve of B. rhodina, isolated from mangrove plants in Riau Province, to identify the ideal harvesting time for maximizing metabolite yield. The fungus was grown in Potato Dextrose Broth (PDB) and kept at ambient temperature (about 25 - 30 °C). Biomass accumulation was evaluated by daily measuring the dry cell weight during a ten-day duration.
The growth curve exhibited a typical sigmoidal pattern, with distinct lag, exponential, stationary, and decline phases, based on triplicate measurements of dry biomass (mean ± SD). The stationary phase (days 6 - 8) was identified as the optimal window for harvesting biomass for subsequent solid-state fermentation, where secondary metabolite biosynthesis was maximal.
As seen in Figure 2 the growth curve of B. rhodina showed a typical sigmoidal pattern with four separate phases: lag, exponential (log), stationary, and decline. During the lag phase (days 0 - 2), biomass increased slightly as the fungus adapted to the nutrient-rich medium. This phase is marked by vigorous metabolic reprogramming and enzyme production, with limited cell division.
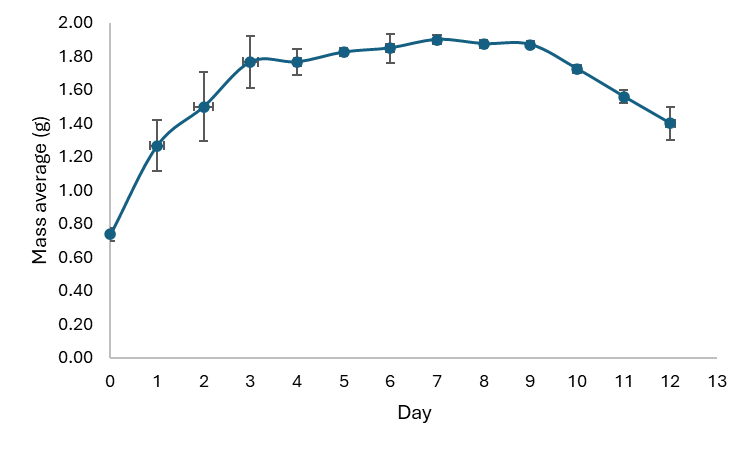
Figure 2 Growth curve of B. rhodian.
The exponential phase, occurring between days 3 and 6, exhibited a significant rise in dry biomass, signifying rapid hyphal growth and elevated metabolic activity. This phase signifies ideal circumstances for primary metabolism, encompassing the production of vital macromolecules necessary for cellular growth and division. The curveʼs slope during this phase indicates the organismʼs maximal specific growth rate (μmax) under the specified culture conditions.
The curve stabilized between days 6 and 8, indicating the start of the stationary phase. This phase is very crucial for the formation of secondary metabolites. In nutrient-deficient situations and heightened metabolic stress, fungi frequently activate secondary metabolism as a survival mechanism. In B. rhodina, this phase is posited to align with the production of antimicrobial chemicals, subsequently validated by LC-MS/MS and GC-MS analysis of the extracts. The stationary phase represents a crucial opportunity for the collection of fungal biomass for metabolite extraction, as the expression of biosynthetic gene clusters often reaches its zenith during this phase [19].
Following the stationary phase, the dry biomass declined after day 9, indicating the onset of the dying phase. This transition reflects metabolic exhaustion and cellular degradation typical of late growth stages. In this phase, autolysis and the breakdown of cellular components prevail, resulting in a decrease in biomass. Harvesting at this stage is inadequate, as both cell viability and metabolite integrity may be jeopardized [20].
The combining of the growth curve with the fermentation approach is essential. Fermentation was carried out on solid-state media (rice substrate), initiated during the stationary phase—predicted from the PDB growth curve—to optimize secondary metabolite yield. Such an approach aligns with previous reports linking the onset of the stationary phase to enhanced production of polyketides, alkaloids, and terpenoids in filamentous fungi [21].
The growth curve of B. rhodina is an essential instrument for determining the optimal period for biomass harvest and the extraction of bioactive secondary metabolites. Figure 2 distinctly demonstrates that the period from day 6 to day 8 constitutes the ideal harvest window, coinciding with the stationary phase, during which the maximum potential for secondary metabolite accumulation is anticipated. Establishing these growth dynamics improves repeatability, aids subsequent bioactivity testing, and guides future scale-up methods for the medicinal utilization of mangrove-derived endophytic fungus.
Solid state fermentation process of fungi
Following the determination of B. rhodinaʼs growth curve in liquid culture, which revealed that the stationary phase occurred between days 6 and 8 (Figure 2), the next step was to implement solid-state fermentation (SSF) to optimize the production of bioactive secondary metabolites. Compared with submerged fermentation, solid-state fermentation offers higher yields, lower contamination risk, and reduced cost. Furthermore, SSF more closely resembles the normal ecological niche of endophytic fungi, particularly those found in plant tissues, which frequently thrive on solid organic substrates.
Rice was selected as the substrate for solid-state fermentation due to its favorable physicochemical characteristics that support the growth and metabolic activity of filamentous fungi. Its balanced carbon-to-nitrogen ratio, high carbohydrate content, and porous structure provide an extensive surface area for fungal colonization and efficient oxygen transfer. The moderate moisture in rice mimics the natural habitat of endophytic fungi and promotes biosynthetic gene cluster activation. Previous studies have demonstrated that rice-based substrates enhance metabolite diversity and yield in fungal fermentations, particularly for the biosynthesis of polyketides, terpenoids, and alkaloids [21,22]. A total of about 40 Erlenmeyer flasks (1 L capacity) were filled with 200 g of sterilized rice and 210 mL of distilled water. The inoculum of B. rhodian, previously cultured on PDA, was equally distributed across the rice surface. The flasks were incubated at room temperature (25 - 30 °C) for ten days. During the incubation period, apparent mycelial development gradually colonized the rice matrix, transforming it from white to a densely coated greyish-black fungal mass (Figure 3).
All solid-state fermentations were carried out under controlled conditions to ensure reproducibility. The initial moisture content of the rice substrate was adjusted to 60 ± 2 %(w/w) and maintained by periodic humidification to prevent drying during incubation. The inoculum density was standardized to 5 %(v/w) using an actively growing mycelial suspension, ensuring uniform colonization across all flasks. Incubation was performed at 28 ± 2 °C under ambient aeration without forced airflow, reflecting optimal conditions for B. rhodina growth and metabolite synthesis. An uninoculated rice substrate (control) was processed in parallel following identical extraction procedures to exclude potential interference from the substrate matrix. Extraction yields were normalized to the dry weight of the fermented substrate (mg extract/g dry biomass) to enable quantitative comparison between replicates.
Over a 10-Day Incubation Period displays the morphological evolution of fungal biomass during the fermentation process. On day two, fungal colonization began, and by day four, there was significant surface covering. Maximum mycelial density and sporulation occurred between days 6 and 8, which corresponded to the previously observed stationary period under submerged conditions. This lends support to the hypothesis that harvest timing is an important component in synchronizing maximal biomass output with secondary metabolite accumulation.
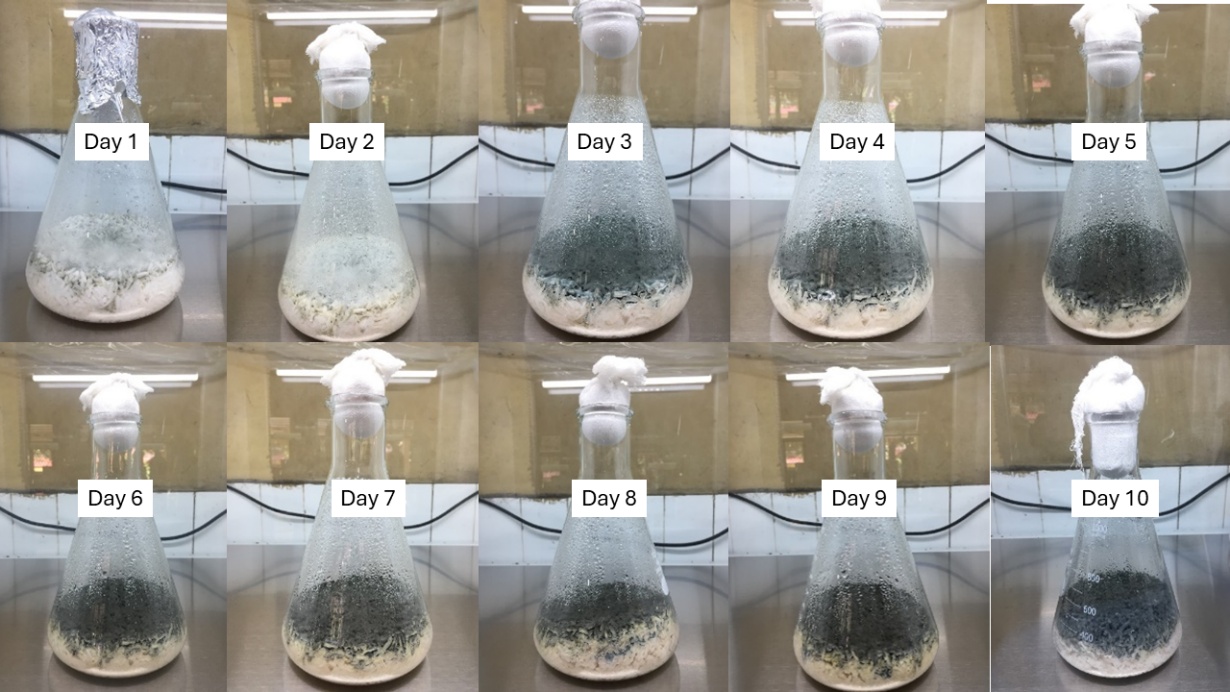

Figure 3 Solid-State fermentation of B. rhodina on rice substrate over a 10-Day incubation period.
The application of solid-state fermentation (SSF) for the synthesis of secondary metabolites is well documented in the literature. The reduced moisture levels in solid-state fermentation systems promote the synthesis of metabolites such polyketides, terpenoids, and alkaloids, which are frequently linked to antibacterial and cytotoxic properties [21,23]. The restricted diffusion of oxygen and food gradients in solid substrates may act as physiological stimuli for the activation of dormant biosynthetic gene clusters, thus increasing chemical diversity. [24].
Following fermentation, the fungal biomass, along with the colonized rice substrate, underwent extraction by the maceration process with ethyl acetate (EtOAc) as the solvent. This solvent was chosen for its moderate polarity and effectiveness in extracting a wide array of semi-polar secondary metabolites [25]. The maceration process was conducted in three consecutive cycles, each lasting 24 h. The initial extraction utilized 1,000 mL of EtOAc, succeeded by 800 mL and 600 mL in the subsequent second and third extractions, respectively. The amalgamated filtrates were subsequently concentrated under diminished pressure utilizing a rotary evaporator at 40 °C, resulting in a semi-solid crude ethyl acetate extract.
The whole crude extract was then weighed and allocated for additional processing. Approximately 1 g of the crude extract was designated for qualitative screening of secondary metabolites, indicating the presence of terpenoids, phenolic compounds, and flavonoids. The remaining extract underwent liquid-liquid partitioning to isolate components according to their polarity. The crude extract was solubilized in methanol, and partitioning was performed using n-hexane in a 1:1 ratio. The operation was reiterated until no additional components were transported to the non-polar phase, signifying complete extraction.
The partitioning method produced two parts with differing physicochemical characteristics. The n-hexane fraction, indicative of the non-polar component of the extract, had a yellowish-brown hue and released a strong odor. The final yield of the hexane fraction after evaporation was 37.17 g. This fraction probably includes lipophilic chemicals such fatty acids, terpenoids, and low polarity phenolics. These families of chemicals have been previously documented to demonstrate antibacterial, antioxidant, and anti-inflammatory effects. [26].
Conversely, the methanolic fraction, which maintained the more polar elements, displayed a dark brown to black hue and emitted a faintly fragrant odor. Upon solvent evaporation, the ultimate yield of this fraction was noted to be 56.19 g. The methanol fraction is believed to encompass polar bioactive molecules, including polyphenols, flavonoids, alkaloids, and polyketides, which are well-documented for their significant antibacterial and bioactive characteristics. [27,28]. The superior yield of the methanol fraction relative to the n-hexane extract likely signifies the prevalence of polar and semi-polar secondary metabolites in B. rhodina, highlighting the fungusʼs potential as a prolific source of various natural products for pharmacological investigation.
Several critical findings arose from the solid-state fermentation (SSF) and subsequent extraction procedures. The identification of the ideal biomass harvest date, informed by the previously established growth curve in liquid culture (Figure 2), greatly enhanced secondary metabolite production. The collection of fungal biomasses during the stationary phase (days 8 - 10) coincided with the maximum production of secondary metabolites, as demonstrated by the significant quantity of crude extracts acquired. The sequential maceration and solvent-partitioning approach enabled efficient recovery and fractionation of a wide array of metabolites. Ethyl acetate served as an effective initial solvent, encompassing a broad range of metabolites, while further partitioning with methanol and n-hexane enhanced separation according to chemical polarity.
The unique physicochemical properties, such as color, flavor, and yield of the resultant fractions, signify varying secondary metabolite profiles, which are essential for future bioassay-guided separation and structural elucidation. The integration of these fractions with sophisticated analytical platforms like liquid chromatography-tandem mass spectrometry (LC-MS/MS) and gas chromatography-mass spectrometry (GC-MS) is crucial for thorough metabolomic characterization. The effective execution of rice-based SSF for B. rhodina highlights the potential for scaling. This fermentation technique, noted for its simplicity, cost efficiency, and scalability, offers a promising strategy for future commercial applications in the pharmaceutical, agricultural, and biotechnology industries.
Antimicrobial activities
Antibacterial activity of B. rhodina extracts was tested against eight clinically relevant bacteria using well diffusion and broth microdilution methods. Table 1 demonstrates that the methanol extract displayed a wider and more potent antibacterial efficacy than the n-hexane extract. In the case of Listeria monocytogenes, the inhibition zone measured 28.20 ± 0.40 mm, signifying robust activity (≥ 16 mm), whereas the n-hexane extract exhibited negligible inhibition (6.21 ± 0.4 mm, classified as inactive). A like tendency was observed for Bacillus subtilis, Staphylococcus aureus, and Vibrio parahaemolyticus, wherein the methanol extract consistently yielded moderate to high inhibitory zones, measuring between 16.47 and 21.37 mm.
Table 1 Antibacterial activities of various extracts from B. rhodina at concentrations 1,000 ppm.
No. |
Microorganism |
Clear zone diameter (mm) |
||
n-Hexane extract |
Methanol extract |
Chloramphenicol 30 ppm |
||
1 |
B. cereus |
13.21 ± 0.5c |
18.86 ± 0.3b |
26.35 ± 2.1a |
2 |
B. subtilis |
17.62 ± 1.1c |
21.37 ± 0.81b |
31.97 ± 2.1a |
3 |
E. coli |
6.72 ± 0.31c |
9.91 ± 0.64b |
28.6 ± 0.4a |
4 |
L. monocytogenes |
6.21 ± 0.4c |
28.2 ± 0.4b |
31.2 ± 2.1a |
5 |
S. typhimurium |
13.25 ± 0.4c |
16.47 ± 0.09b |
22.8 ± 1.1a |
6 |
S. aureus |
16.08 ± 0.6c |
18.41 ± 0.4b |
28.1 ± 3.1a |
7 |
V. algynolyticus |
6.25 ± 0.4c |
19.18 ± 3.2b |
21.95 ± 3.1a |
8 |
V. parahaemolyticus |
14.36 ± 0.3c |
17.86 ± 1.0b |
25.82 ± 2.3a |
The results were expressed as mean ± Standard Deviation (SD) in the same row with different superscript letters indicates a statistically significant difference based on Tukey’s test (α = 5%).
The n-hexane extract shown moderate antibacterial activity solely against Bacillus subtilis (17.62 mm) and S. aureus (16.08 mm), while its effectiveness against other examined bacterial strains varied from inactive to weak, with inhibition zones of ≤ 15 mm. Conversely, chloramphenicol, utilized as a standard reference antibiotic, exhibited enhanced inhibition against all bacterial strains, hence validating the assayʼs trustworthiness. These results align with the established categorization standards for disc diffusion experiments, where inhibitory zones are categorized as follows: Inactive (≤ 6 mm), weak activity (7 - 10 mm), moderate activity (11 - 15 mm), and strong activity (≥ 16 mm) [28].
The MIC and MBC values (Table 3) confirmed these results. The methanol extract exhibited reduced MICs (31.25 - 250 ppm) against the majority of Gram-positive bacteria, especially L. monocytogenes (MIC: 31.25 ppm), indicating significant bacteriostatic or bactericidal efficacy. The MBC/MIC ratio for L. monocytogenes, B. subtilis, and S. aureus was ≤ 4, signifying a bactericidal mechanism of action. The ratio for E. coli and V. algynolyticus surpassed 4, indicating a bacteriostatic effect. The antibacterial effects are likely due to polar metabolites in the methanol extract, including polyphenols, alkaloids, and flavonoids, which are known to damage bacterial membranes or block metabolic processes [29].
The antifungal activity of both extracts was tested against Candida albicans, C. glabrata and C. krusei using the Kirby-Bauer disc diffusion method and MIC/MFC assays. As shown in Table 2, methanol extract exhibited moderate activity against all three strains (zones of 12.35 - 15.54 mm), whereas the n-hexane extract showed no activity against C. albicans and C. glabrata (≤ 6 mm), and weak activity (10.92 mm) against C. krusei.
Table 2 Antifungal activities of various extracts from B. rhodina at concentrations 1,000 ppm.
No. |
Microorganism |
Clear zone diameter (mm) |
||
n-Hexane extract |
Methanol extract |
Ketoconazole 30 ppm |
||
1 |
C. albicans |
6.71 ± 0.32c |
12.35 ± 0.65b |
24.86 ± 0.62a |
2 |
C. glabrata |
6.62 ± 0.26c |
13.15 ± 0.32b |
23.42 ± 0.68a |
3 |
C. krusei |
10.92 ± 0.18c |
15.54 ± 0.65b |
25.67 ± 0.63a |
The results were expressed as mean ± Standard Deviation (SD) in the same row with different superscript letters indicates a statistically significant difference based on Tukeyʼs test (α = 5%).
Table 3 The minimum inhibition concentration (MIC) and minimum bactericidal concentration (MBC)/ minimum fungicidal concentration (MFC) of various extracts from B. rhodina.
No. |
Microorganism |
MIC (ppm) |
MBC (ppm) |
||||
n-Hexane extract |
Methanol extract |
Chloramphenicol |
n-Hexane extract |
Methanol extract |
Chloramphenicol |
||
1 |
B. cereus |
500 |
250 |
31.25 |
1,000 |
500 |
62.5 |
2 |
B. subtilis |
250 |
125 |
31.25 |
500 |
500 |
62.5 |
3 |
E. coli |
>1,000 |
1000 |
31.25 |
>1,000 |
>1,000 |
62.5 |
4 |
L. monocytogenes |
>1,000 |
31.25 |
31.25 |
>1,000 |
62.5 |
62.5 |
5 |
S. typhimurium |
500 |
250 |
31.25 |
1,000 |
1,000 |
62.5 |
6 |
S. aureus |
250 |
250 |
31.25 |
500 |
500 |
62.5 |
7 |
V. algynolyticus |
>1,000 |
62.5 |
31.25 |
>1,000 |
125 |
125 |
8 |
V. parahaemolyticus |
500 |
250 |
31.25 |
1,000 |
500 |
125 |
|
|
MIC (ppm) |
MFC (ppm) |
||||
No. |
Microorganism |
n-Hexane extract |
Methanol extract |
Ketoconazole |
n-Hexane extract |
Methanol extract |
Ketoconazole |
1 |
C. albicans |
1,000 |
500 |
15.62 |
>1,000 |
1,000 |
31.25 |
2 |
C. glabrata |
1,000 |
500 |
15.62 |
>1,000 |
1,000 |
31.25 |
3 |
C. krusei |
500 |
250 |
15.62 |
>1,000 |
500 |
31.25 |
The MIC values (Table 3) corroborated these results, indicating that the methanol extract exhibited MICs between 250 and 500 ppm, and MFCs from 500 - 1,000 ppm. Notably, all MFC/MIC ratios above 4, signifying a fungistatic mechanism of action. The methanol extract had significant effectiveness, particularly against C. krusei, which frequently shows resistance to standard azoles, although it is less potent than ketoconazole (MIC: 15.62 ppm). The findings indicate that B. rhodina synthesizes secondary compounds with prospective antifungal uses, especially against opportunistic pathogenic Candida species.
Although the antifungal activity of B. rhodina extracts was moderate compared to their antibacterial potency, this difference may be attributed to fundamental structural and physiological disparities between bacterial and fungal cells. The chitin-glucan matrix and ergosterol-enriched membranes of fungi typically limit the permeability of polar metabolites, such as phenolics and flavonoids, that predominated in the methanol extract. Consequently, metabolites that act effectively on bacterial cell walls or membrane systems may exhibit reduced interaction with fungal targets. The dominance of compounds like kojic acid, glycitein, and fraxetin—well-known for strong antibacterial but only moderate antifungal activities—suggests a metabolite composition biased toward antibacterial functionality.
Nevertheless, the measurable inhibition of C. krusei, a strain often resistant to conventional azole antifungals, indicates that certain metabolites in the extract may exert selective fungistatic activity. Such effects are likely associated with interference in ergosterol biosynthesis or disruption of fungal redox balance, leading to partial growth suppression rather than fungicidal action. The high MFC/MIC ratios (≥ 4) further support this interpretation. Importantly, this outcome also implies that under modified culture conditions or co-culture stimulation, B. rhodina could potentially activate latent biosynthetic gene clusters responsible for more potent antifungal metabolites. Future bioassay-guided fractionation and transcriptomic analysis are warranted to elucidate the structures and biosynthetic pathways of these compounds.
Overall, while the antifungal effect is moderate, the observed selectivity underscores B. rhodina’s adaptive metabolic potential. The findings suggest a biosynthetic preference toward antibacterial secondary metabolism while maintaining a baseline antifungal capability that could be enhanced through fermentation optimization or elicitation strategies. This dual-spectrum potential positions B. rhodina as a valuable candidate for future antimicrobial drug discovery efforts.
The antibacterial efficacy demonstrated in this study for B. rhodina corresponds with and considerably enhances previous findings about the bioactivity of Botryosphaeria species. In the restricted yet significant literature, Abdou et al. [30] presented the inaugural complete evidence of antimicrobial metabolite synthesis from B. rhodina, isolating and assessing two depsidone compounds—Botryorhodine A and Botryorhodine B—for their antifungal and cytotoxic properties. [30]. Botryorhodine A displayed a MIC of 26.03 µM against A. terreus and 191.60 µM against Fusarium oxysporum, whereas Botryorhodine B showed MICs of 49.70 and 238.80 µM against the same fungi, respectively. [30] This study extends previous findings by showing that crude extracts, especially the methanol-soluble fraction, of B. rhodina cultivated under solid-state fermentation conditions display significant and extensive antibacterial properties. The methanol extract had significant inhibitory effects against L. monocytogenes (28.20 mm), B. subtilis (21.37 mm), and S. aureus (18.41 mm), along with modest antifungal activity against Candida species. Although our study did not isolate or quantify specific compounds like Botryorhodines, the observed antimicrobial activities—especially those demonstrating bactericidal or fungistatic properties indicated by MIC/MBC and MIC/MFC ratios—imply the existence of structurally analogous or similarly functioning metabolites.
Aside from B. rhodina, documentation regarding other Botryosphaeria species exhibiting antibacterial characteristics is scarce. Karličić et al. [31] evaluated the inhibition of mycelial growth of B. dothidea, Dothiorella sarmentorum, and Neofusicoccum parvum under the biocontrol influence of Trichoderma species and pine bark extracts. Nonetheless, this research concentrated mostly on antagonistic interactions instead of metabolite-mediated antibacterial efficacy. For example, dual-culture tests with Trichoderma achieved 67% - 85% mycelial suppression of B. dothidea, but pine bark extracts elicited only 39% - 44% inhibition. No antimicrobial chemicals were directly extracted from B. dothidea, highlighting the distinctiveness of B. rhodina as a biosynthetic reserve within the genus. [31].
The ecological significance of B. rhodina as an endophyte is likewise remarkable. Endophytic fungi are recognized for their role in chemical defense for host plants, synthesizing antimicrobial secondary metabolites that inhibit the proliferation of competing microbial diseases within the plant. This mutualistic characteristic has been suggested as a catalyst in the evolution of varied and bioactive metabolite biosynthetic gene clusters in endophytes. The presence of B. rhodina in mangrove environments, marked by variable salinity, tidal stress, and intense microbial competition, may enhance the synthesis of structurally distinct and bioactive chemicals.
Metabolites profiling of B. rhodina extracts
Metabolite profiling revealed a chemically diverse set of secondary metabolites, highlighting the fungus’s metabolic potential. A broad array of both polar and non-polar metabolites, encompassing various chemical classes, was identified using complementary analytical platforms: LC-MS/MS for the methanol extract and GC-MS for the n-hexane extract. Figure 4 depicts the Total Ion Chromatogram (TIC) of the methanol extract derived from UHPLC-Q-Orbitrap LC-MS/MS analysis, showcasing an intricate metabolite profile characterized by several high-intensity peaks over the retention time spectrum. This indicates the significant chemical variety and polarity of the extract.
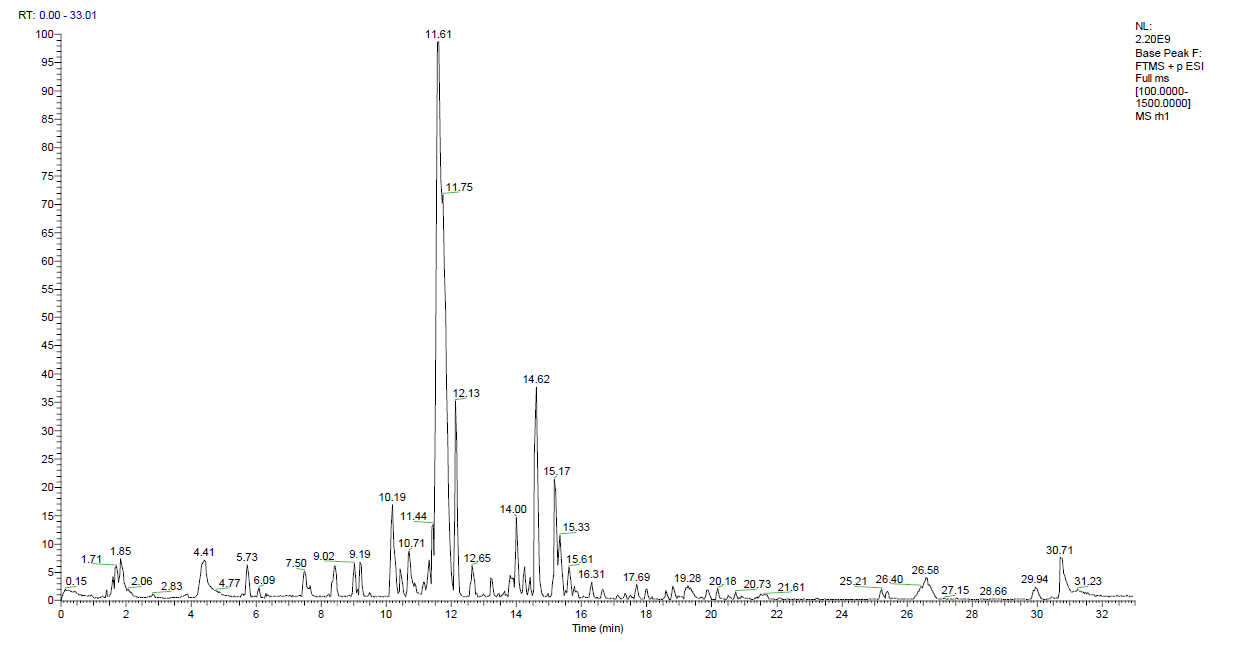
Figure 4 Total Ion Chromatogram (TIC) from a High-Resolution Mass Spectrometry (HRMS) of methanol extract.
Figure 4 presents the Total Ion Chromatogram (TIC) derived from high-resolution LC-MS/MS analysis of the methanol extract of B. rhodina. The TIC displays an intricate and concentrated distribution of peaks within the 0-32-minute retention time interval, indicating the existence of a chemically varied assortment of polar and semi-polar secondary metabolites. The elevated signals detected between 5 and 25 min indicate the elution of numerous mid-polarity chemicals, such as flavonoids, phenolics, alkaloids, and polyketides, many of which have been previously linked to antibacterial properties [27]. The numerous peaks signify a substantial biosynthetic yield under the specified solid-state fermentation conditions, underscoring the metabolic abundance of the extract. This chromatographic profile underpins the subsequent identification of compounds detailed in Table 4 and corroborates the extensive antibacterial capability demonstrated in the biological experiments.
Table 4 Metabolite profiling of methanol extract.
No |
Name |
Formula |
Annot. DeltaMass [ppm] |
Calc. MW |
Area |
1 |
Kojic acid |
C₆H₆O₄ |
‒ 1.9 |
142.02634 |
611238473.7 |
2 |
4-Acetamidobutanoic acid |
C₆H₁₁NO₃ |
‒ 1.32 |
145.0737 |
1449448289 |
3 |
Pyridoxal |
C₈H₉NO₃ |
‒ 15.52 |
167.05565 |
338653086.6 |
4 |
(Hydroxyethyl)methacrylate |
C₆H₁₀O₃ |
‒ 0.85 |
130.06288 |
1736773156 |
5 |
Sorbic acid |
C₆H₈O₂ |
0.69 |
112.05251 |
357401798.7 |
6 |
Unknown 23 |
C₈H₉NO₂ |
‒ 1.33 |
151.06313 |
274206821.4 |
7 |
N-Acetylvaline |
C₇H₁₃NO₃ |
‒ 1.36 |
159.08933 |
393009948.7 |
8 |
OI1700000 |
C₇H₁₂O₃ |
‒ 1.26 |
144.07846 |
700728707.4 |
9 |
LV1850000 |
C₇H₈O₃ |
‒ 2.14 |
140.04704 |
1058266413 |
10 |
N-(2-OXOOXOLAN-3-YL)HEXANAMIDE |
C₁₀H₁₇NO₃ |
‒ 1.49 |
199.12055 |
329035257 |
11 |
Phenylethyl alcohol |
C₈H₁₀O |
0.59 |
122.07324 |
445092318.9 |
12 |
Vanillyl alcohol |
C₈H₁₀O₃ |
‒ 1.12 |
154.06282 |
412503890.6 |
13 |
2-Acetylcyclohexanone |
C₈H₁₂O₂ |
‒ 1.87 |
140.08347 |
755270815.2 |
14 |
Ethyl 3-oxohexanoate |
C₈H₁₄O₃ |
‒ 0.82 |
158.09416 |
395148814.1 |
15 |
DL-2-(acetylamino)-3-phenylpropanoic acid |
C₁₁H₁₃NO₃ |
‒ 0.6 |
207.08942 |
471678300.2 |
16 |
N-(2-furylmethylidene)-(4-([(2-furylmethylidene)amino]methyl}cyclohexyl)methanamine |
C₁₈H₂₂N₂O₂ |
‒ 1.5 |
298.16768 |
1250462144 |
17 |
Dodecanedioic acid |
C₁₂H₂₂O₄ |
‒ 1.65 |
230.15143 |
820355426.9 |
18 |
NP-001501 |
C₁₆H₁₇N₃O₂ |
‒ 1.98 |
283.13152 |
255969460 |
19 |
NP-021844 |
C₁₂H₂₄O₅ |
88576.73 |
270.14379 |
274570720.2 |
20 |
(2S,3S)-2-hydroxytridecane-1,2,3-tricarboxylic acid |
C₁₆H₂₈O₇ |
‒ 2.06 |
332.18282 |
305562642.3 |
21 |
4,4’-Bis(diethylamino)benzophenone |
C₂₁H₂₈N₂O |
‒ 2.61 |
324.21932 |
2888445537 |
22 |
Tetraneurin A |
C₁₇H₂₂O₆ |
‒ 1.86 |
322.14104 |
259063397.3 |
23 |
Unknown 1 |
C₂₃H₃₀N₂O₃ |
‒ 1.54 |
382.22505 |
710625326.5 |
24 |
Unknown 2 |
C₂₂H₂₀N₄O₅ |
‒ 2.42 |
420.14235 |
274065787.1 |
25 |
Ofloxacin methyl ester |
C₁₉H₂₂FN₃O₄ |
‒ 5.43 |
375.1574 |
725248196.1 |
26 |
MFCD23380831 |
C₁₆H₁₃NO₆ |
‒ 1.81 |
315.07372 |
696943477.5 |
27 |
Unknown 3 |
C₁₆H₁₅NO₇ |
‒ 2.46 |
333.08403 |
365139587.8 |
28 |
Unknown 4 |
C₁₇H₁₇NO₇ |
‒ 2.43 |
347.09966 |
416699417.5 |
29 |
Erlotinib |
C₂₂H₂₃N₃O₄ |
‒ 1.75 |
393.16817 |
1011283886 |
30 |
Unknown 5 |
C₂₂H₂₈O₂S |
‒ 2.53 |
356.1801 |
332104504.3 |
31 |
Vesnarinone |
C₂₂H₂₅N₃O₄ |
‒ 1.26 |
395.18401 |
265101829.3 |
32 |
NP-021781 |
C₁₉H₃₆O₅ |
63828.58 |
366.22966 |
52316910304 |
33 |
Unknown 6 |
C₄₀H₇₂N₄S₅ |
2.11 |
768.43767 |
298424562.1 |
34 |
(3,4-Dimethoxyphenyl)((3S)-3-[5-(3-pyridinyl)-1,3,4-oxadiazol-2-yl]-1-pyrrolidinyl}methanone |
C₂₀H₂₀N₄O₄ |
57827.89 |
402.13164 |
5394895666 |
35 |
2-(3,4-Dimethoxyphenyl)-5,7,8-trihydroxy-3-methoxy-2,3-dihydro-4H-chromen-4-one |
C₁₈H₁₈O₈ |
‒ 2.59 |
362.09923 |
260749886.7 |
36 |
Unknown 7 |
C₂₁H₂₈N₂ |
‒ 2.42 |
308.2245 |
648233913.7 |
37 |
Geranyl glucoside |
C₁₆H₂₈O₆ |
‒ 2.23 |
316.18788 |
887106925.6 |
38 |
Octyl (4xi)-2-deoxy-2-[(Z)-(1-hydroxyethylidene)amino]-alpha-D-xylo-hexopyranoside |
C₁₆H₃₁NO₆ |
‒ 2.48 |
333.21431 |
386956579.3 |
39 |
Unknown 8 |
C₁₄H₂₂N₆O₄ |
‒ 1.65 |
338.16969 |
351044651.8 |
40 |
(1S,3S,12S,14R,27S)-12-Hydroxy-1,3-dimethyl-2,5,15,23,25-pentaazaheptacyclo[12.10.2.1~2,5~.0~6,11~.0~12,27~.0~15,24~.0~17,22~]heptacosa-6,8,10,17,19,21,23-heptaene-4,16,26-trione |
C₂₄H₂₁N₅O₄ |
‒ 1.66 |
443.15862 |
631153618.7 |
41 |
4-Methoxycinnamic acid |
C₁₀H₁₀O₃ |
‒ 1.65 |
178.0627 |
961957815.5 |
42 |
Deoxybrevianamide E |
C₂₁H₂₅N₃O₂ |
‒ 1.97 |
351.19399 |
263163030.8 |
43 |
9-Methoxy-12-(2-methyl-1-propen-1-yl)-1,2,3,5a,6,11,12,14a-octahydro-5H,14H-pyrrolo[1’’,2’’:4’,5’]pyrazino[1’,2’:1,6]pyrido[3,4-b]indole-5,14-dione |
C₂₂H₂₅N₃O₃ |
‒ 2.41 |
379.18868 |
1657865510 |
44 |
Unknown 9 |
C₁₈H₃₂O₆ |
‒ 1.84 |
344.21926 |
419291370.8 |
45 |
PPG Acrylate n5 |
C₁₈H₃₄O₇ |
60682.5 |
384.2115 |
254965891.2 |
46 |
Usnic acid |
C₁₈H₁₆O₇ |
‒ 2.44 |
344.08876 |
509685447.5 |
47 |
Unknown 10 |
C₃₈H₂₄N₈O₆ |
‒ 3.74 |
688.17931 |
500753787.2 |
48 |
NP-005405 |
C₁₇H₁₆O₅ |
‒ 1.41 |
300.09935 |
300686318.2 |
49 |
Unknown 11 |
C₂₄H₄₀O₈ |
‒ 1.66 |
456.27156 |
412576947.9 |
50 |
3,4-Dihydroxyphenylpropionic acid |
C₉H₁₀O₄ |
‒ 98929.48 |
164.04701 |
260039679.7 |
51 |
Fraxetin |
C₁₀H₈O₅ |
‒ 2.72 |
208.03661 |
3464812649 |
52 |
5,7-DIHYDROXY-3’,4’,5’-TRIMETHOXYFLAVANONE |
C₁₈H₁₈O₇ |
‒ 2.41 |
346.10442 |
3193661421 |
53 |
N-(4-fluorophenyl)-2-methyl-5-(morpholinosulfonyl)-3-furamide |
C₁₆H₁₇FN₂O₅S |
5.95 |
368.08641 |
341196330.6 |
54 |
Glycitein |
C₁₆H₁₂O₅ |
‒ 2.27 |
284.06783 |
2008325644 |
55 |
[(5aS,6S,7aR,8R,9S,11aS,11bS,12R)-6,9-bis(acetyloxy)-12-hydroxy-5a,8,11a-trimethyl-1-oxo-3-(pyridin-3-yl)-1,5a,6,7,7a,8,9,10,11,11a,11b,12-dodecahydro-2,5-dioxatetraphen-8-yl]methyl acetate |
C₃₁H₃₇NO₁₀ |
‒ 1.25 |
583.24101 |
345610334.8 |
56 |
Unknown 12 |
C₂₁H₁₉NO₇ |
‒ 2.04 |
397.11534 |
261918464.7 |
57 |
Unknown 13 |
C₂₂H₃₆N₆O₇ |
‒ 0.75 |
496.26418 |
327918171.7 |
58 |
Unknown 14 |
C₂₄H₄₂O₉ |
‒ 1.6 |
474.28212 |
919030640.6 |
59 |
Unknown 15 |
C₂₂H₂₁NO₇ |
‒ 2.62 |
411.13072 |
399419961.5 |
60 |
Unknown 16 |
C₂₄H₂₀N₄O₅ |
‒ 2.09 |
444.14244 |
339186996.8 |
61 |
Unknown 17 |
C₃₂H₅₆O₁₂ |
‒ 3.46 |
632.37499 |
540939073.8 |
62 |
Unknown 18 |
C₃₆H₅₆O₄P₂ |
‒ 0.96 |
614.36479 |
525531675 |
63 |
Unknown 19 |
C₄₀H₇₀O₁₅ |
‒ 3.46 |
790.46874 |
410030023.4 |
64 |
Unknown 20 |
C₂₇H₃₁N₃O₄ |
‒ 2.37 |
461.23036 |
453586050.2 |
65 |
Unknown 21 |
C₃₀H₅₅N₅O₅ |
‒ 3.1 |
565.41857 |
457024359.2 |
66 |
Unknown 22 |
C₄₂H₇₄O₁₅ |
‒ 2.43 |
818.50078 |
359907611.2 |
67 |
Oleamide |
C₁₈H₃₅NO |
‒ 1.4 |
281.27147 |
323460282.1 |
Similarly, Figure 5 illustrates the TIC obtained from GC-MS analysis of the n-hexane extract, with a more confined distribution of peaks primarily within the mid to late retention time range, signifying the presence of less polar chemicals such as fatty acids, esters, and sterols. The chromatograms show distinct chemical compositions for each extract, reflecting differences in solvent polarity and extraction efficiency.
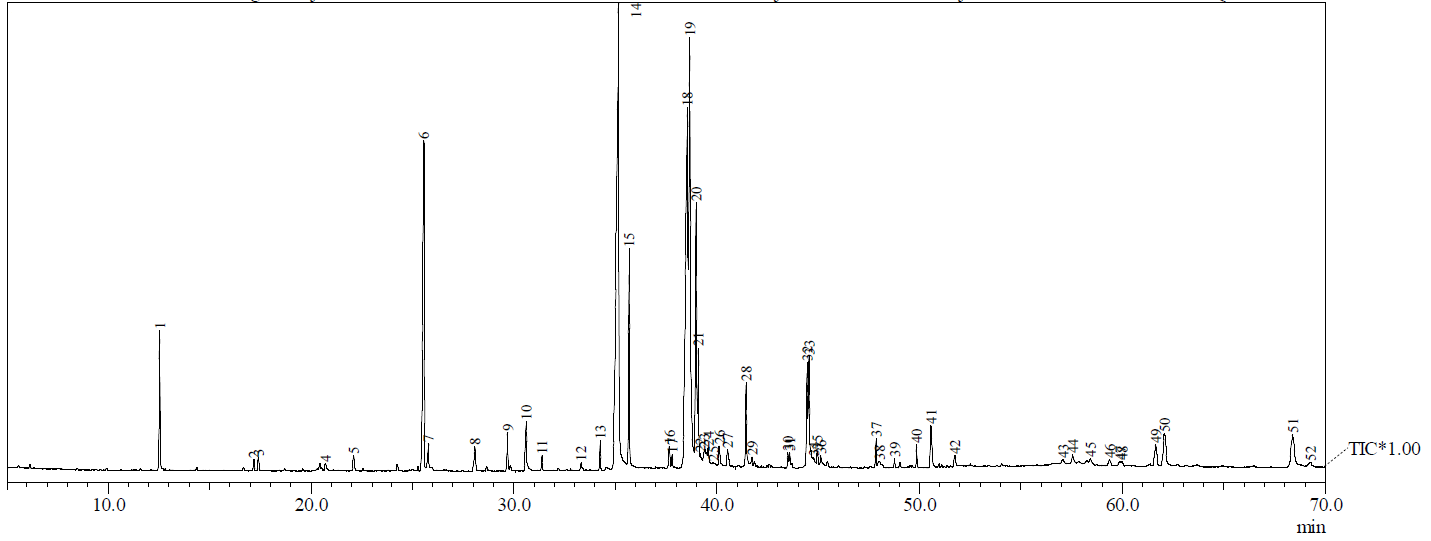
Figure 5 Total Ion Chromatogram (TIC) from GC-MS analysis of the n-hexane extract.
Figure 5 presents the Total Ion Chromatogram (TIC) derived from the GC-MS analysis of the n-hexane extract of B. rhodina, illustrating a comparatively simpler chemical profile than that of the methanol extract. The majority of peaks are concentrated in the 20-45-minute retention time range, corresponding primarily to the elution of non-polar lipophilic compounds such as saturated and unsaturated fatty acids, esters, and sterols. The intensity and distribution of these peaks suggest the extract is dominated by compounds with limited polarity, such as ethyl palmitate, linoleic acid, and clionasterol, which are known to exhibit mild to moderate antimicrobial or anti-inflammatory properties. Although less complex than the methanolic TIC, this chromatographic pattern provides valuable insight into the lipid-based metabolome of B. rhodina, as detailed in Table 5, and supports the differential antimicrobial activities observed between polar and non-polar extracts.
Table 5 Metabolite profiling of n-Hexane extract.
No |
Name |
Area |
1 |
Phenylethyl Alcohol |
16342713 |
2 |
Acetic acid, 2-phenylethyl ester |
1253967 |
3 |
5-Hydroxy-2-Decenoic Acid Lactone |
1689771 |
4 |
Benzaldehyde, 4-ethoxy- |
1020155 |
5 |
Benzeneethanol, 4-hydroxy- |
2064942 |
6 |
Dodecanoic acid |
4913178 |
7 |
Benzaldehyde, 4-ethoxy- (CAS) p-Ethoxybenzaldehyde |
4355374 |
8 |
Benzaldehyde, 3-ethoxy- |
5424632 |
9 |
Tetradecanoic acid (CAS) Myristic acid |
8398044 |
10 |
Tetradecanoic acid, ethyl ester (CAS) Ethyl myristate |
1677370 |
11 |
1H-2-Benzopyran-1-one, 3,4-dihydro-3,8-dihydroxy-3-methyl-, (-)- |
1572026 |
12 |
Hexadecanoic acid, methyl ester |
3373432 |
13 |
Hexadecanoic acid, ethyl ester (CAS) Ethyl palmitate |
25153720 |
14 |
9,12-Hexadecadienoic acid, methyl ester (CAS) METHYL-9,12-Hexadecadienoate |
3361719 |
15 |
11-Octadecenoic acid, methyl ester, (Z)- |
1833382 |
16 |
Ethyl Oleate |
13861020 |
17 |
9-Octadecenoic acid (Z)-, ethyl ester (CAS) Ethyl oleate |
2515482 |
18 |
9,12-Octadecadienoic acid (Z,Z)- (CAS) Linoleic acid |
6227145 |
19 |
Octadecanoic acid, ethyl ester |
3220091 |
20 |
9,12-Octadecadienoic acid (Z,Z)- (CAS) Linoleic acid |
1215008 |
21 |
9,12-Octadecadienoic acid (Z,Z)- |
3597971 |
22 |
Isoamyl laurate |
2702163 |
23 |
Octadecanoic acid, 2-hydroxy-1,3-propanediyl ester |
11995847 |
24 |
8,11-Octadecadienoic acid, methyl ester |
1465620 |
25 |
9,11-Octadecadienoic acid, methyl ester, (E,E)- |
2112308 |
26 |
9-Octadecene, 1-methoxy-, (E)- (CAS) 9(TRANS)-OCTADEC-1-Enyl Methyl Ether |
2199505 |
27 |
9,12-Octadecadien-1-ol (CAS) OCTADECA-9,12-DIEN-1-OL |
14524843 |
28 |
DI-(9-OCTADECENOYL)-GLYCEROL |
20306995 |
29 |
Octadecanoic acid, 3-oxo-, methyl ester (CAS) Methyl 3-oxooctadecanoate |
1831249 |
30 |
Octadecanoic acid, 2-hydroxy-1,3-propanediyl ester |
2872009 |
31 |
Hexadecanoic acid, 2-hydroxy-1-(hydroxymethyl)ethyl ester |
3338067 |
32 |
Octanoic acid, 2-phenylethyl ester |
3736257 |
33 |
9-Octadecenoic acid (Z)-, 2,3-dihydroxypropyl ester |
1965918 |
34 |
Di-n-octyl phthalate |
1192388 |
35 |
2,6,10,14,18,22-Tetracosahexaene, 2,6,10,15,19,23-hexamethyl- (CAS) Squalene |
2605876 |
36 |
Benzene, 1-(1-cyclohexen-1-yl)ethyl - |
8112199 |
37 |
9-Octadecyne |
2297577 |
38 |
Octadecanoic acid, ethenyl ester (CAS) Vinyl stearate |
2103125 |
39 |
Stigmast-5-en-3-ol, (3.beta.,24S)- (CAS) Clionasterol |
1604069 |
40 |
Stigmast-4-en-3-one |
1537555 |
41 |
Ergosta-4,7,22-trien-3-one |
1143670 |
42 |
4,22-Cholestadien-3-one |
1001862 |
43 |
Stigmast-4-en-3-one |
5911489 |
44 |
(R)-(-)-14-Methyl-8-hexadecyn-1-ol |
12119013 |
Compound identification was based on tandem mass spectrometric (MS²) data acquired using high-resolution UHPLC-Q Exactive Orbitrap LC-MS/MS and validated through comparison with mzCloud, ChemSpider, and MassBank databases at a mass tolerance of ± 5 ppm. Fragmentation spectra were matched to reference libraries to enhance annotation reliability. Therefore, metabolites such as kojic acid, usnic acid, and glycitein were putatively identified based on accurate mass, retention time, and high spectral similarity, pending further structural confirmation by NMR.
The PLS-DA models generated from LC-MS/MS (Figure 5) and GC-MS (Figure 6) datasets demonstrated clear separation between methanolic and hexane extracts, confirming distinct metabolomic profiles. VIP score plots indicated that polar metabolites, including kojic acid, fraxetin, and glycitein, were major contributors to the antibacterial activity, while several uncharacterized metabolites with high VIP values suggest the presence of novel bioactive chemical entities requiring future isolation and NMR confirmation.
The utilization of high-resolution Orbitrap-based liquid chromatography-mass spectrometry (LC-MS/MS) enabled the thorough detection and provisional identification of 67 unique secondary metabolites in the methanol extract of B. rhodina. This analytical method provided outstanding mass accuracy, elevated resolving power, and sensitive detection of a wide range of polar and semi-polar molecules, facilitating precise molecular feature extraction and annotation [32]. The utilization of tandem mass spectrometry and spectrum database matching through ChemSpider and mzCloud platforms greatly boosted chemical identification and structural prediction. Simultaneously, gas chromatography-mass spectrometry (GC-MS) was utilized to examine the n-hexane extract, which is rich in non-polar components. This method facilitated the identification of 45 volatile and semi-volatile constituents, including fatty acids, esters, and sterols, according to their retention times and fragmentation patterns. The combined orthogonal metabolomic platforms yielded a thorough chemical profile of both polar and non-polar fractions, as detailed in Tables 4 and 5, therefore clarifying the metabolic complexity and biochemical potential of this mangrove-derived endophytic fungus.
The methanol extract contained diverse compounds, including phenolics, flavonoids, alkaloids, terpenoids, and heterocyclic derivatives. Significant levels of well-documented antimicrobial metabolites, including kojic acid, usnic acid, glycitein, fraxetin, vanillyl alcohol, and deoxybrevianamide E were —were putatively identified based on accurate mass and MS/MS spectral similarity to reference spectra in the mzCloud and ChemSpider databases. These annotations are considered tentative and require further structural confirmation through isolation and NMR analysis. [33-37]. These chemicals are extensively acknowledged in the literature for their antibacterial, antifungal, or cytoprotective properties, indicating a functional role in the bioactivity identified in the extract. The existence of 25 unidentified compounds with significant relative abundance, including those with intricate nitrogen-containing or halogenated structures, underscores the potential for novel bioactive substances. Future investigations should focus on the structural elucidation of these unidentified compounds using bioassay-guided fractionation and NMR spectroscopy.
The discussion generally focuses on antibacterial action; however, numerous metabolites found in the methanol extract, including kojic acid, usnic acid, and vanillyl alcohol, are recognized for their antifungal characteristics as well [33,34,36]. These chemicals may facilitate the mild inhibition noted against Candida species, especially C. krusei, which frequently demonstrates resistance to standard antifungals. Additionally, certain undiscovered metabolites with elevated peak intensities may possess new antifungal scaffolds. The lack of isolated targeted antifungal compounds restricts accurate attribution of bioactivity; however, the findings suggest that the methanol extract encompasses multifunctional secondary metabolites with potential antibacterial and antifungal applications. Future research utilizing bioassay-guided purification and fungal-specific synergy testing is crucial to clarify these connections.
In contrast, the n-hexane extract primarily comprised saturated and unsaturated fatty acids (e.g., dodecanoic acid, myristic acid, linoleic acid), fatty acid esters (e.g., ethyl palmitate, ethyl oleate), sterols (e.g., clionasterol), and long-chain hydrocarbons. While these constituents are typically regarded as less effective direct antimicrobial agents than their polar counterparts, some have been documented to possess mild antibacterial activities and may synergistically enhance the overall activity profile. [38].
The results combined indicate a substantial segregation of metabolites that is dependent on polarity. The methanol extract, comprising a chemically varied and pharmacologically important collection of polar molecules, exhibited superior and more extensive antibacterial activity compared to the n-hexane extract, which was predominantly limited to lipid-based constituents. This distinction provides a molecular rationale for the observed variations in bioactivity and underscores the importance of solvent selection in optimizing the extraction of secondary metabolites. The observations support the view that B. rhodina is a promising source of chemically diverse secondary metabolites, with significant potential for discovering novel antimicrobial drugs from underexplored fungal endophytes in mangrove ecosystems.
An in-depth analysis of the methanol extract identified multiple well-defined bioactive chemicals exhibiting various antibacterial actions. Kojic acid, a secondary metabolite synthesized by several fungi, is recognized for its antibacterial and antifungal properties via metal ion chelation and inhibition of tyrosinase [39]. Usnic acid, a dibenzofuran derivative, demonstrates antibacterial properties through membrane disruption and the suppression of oxidative phosphorylation [34]. Flavonoid derivatives, including glycitein, fraxetin, and 5,7-dihydroxy-3’,4’,5’-trimethoxyflavanone, exhibit antibacterial, antioxidant, and anti-inflammatory activities, typically through mechanisms such as efflux pump blockage, oxidative stress induction, and cell wall disruption [40]. Their simultaneous presence in the extract indicates possible synergistic interactions. Other notable chemicals comprise phenylethyl alcohol, a quorum sensing antagonist; vanillyl alcohol, which disrupts ergosterol biosynthesis; and pyridoxal, a vitamin B6 derivative that amplifies oxidative stress in microbial cells. The inclusion of nitrogen-containing heterocycles and polyketide-like structures, such as deoxybrevianamide E and tetraneurin A, enhances the bioactive potential of the extract.
The antibacterial activity of the methanol extract was substantially linked with the concentration of these polar metabolites. Significant inhibition zones and low MIC values—31.25 ppm for L. monocytogenes and 125 ppm for B. subtilis—demonstrate bactericidal property. The detected antibacterial activity probably arises from a combination of individual and synergistic effects among these substances. Conversely, the n-hexane extract, characterized by fatty acids, esters, and sterols, demonstrated minimal antibacterial efficacy. Although certain elements, such as linoleic acid and methyl palmitate, have moderate membrane-disruptive effects, their total impact is quite insignificant.
Alongside the well-characterized chemicals, the identification of more than 25 unidentified metabolites in the methanol extract—many exhibiting intricate, high-molecular-weight structures—indicates the potential presence of novel chemical scaffolds with bioactive properties. These metabolites exhibited elevated peak intensities yet lacked conclusive matches in spectrum libraries, highlighting the constraints of existing databases and the distinctive chemical space possessed by B. rhodina.
The correlation between metabolite chemistry and biological activity suggests that several compounds act through distinct and complementary mechanisms. Kojic acid likely exerts antibacterial effects via chelation of essential metal ions and inhibition of tyrosinase-dependent redox enzymes, leading to oxidative imbalance. Usnic acid, a dibenzofuran derivative, disrupts microbial membrane integrity and inhibits oxidative phosphorylation, accounting for its broad-spectrum activity. Flavonoid derivatives such as glycitein and fraxetin may inhibit efflux pumps, interfere with nucleic acid synthesis, and induce reactive oxygen species accumulation, thereby enhancing bactericidal efficacy. The presence of deoxybrevianamide E and nitrogen-containing polyketides further supports a multi-target mode of action involving redox modulation and membrane destabilization.
These observations indicate that the antimicrobial potential of B. rhodina extracts arises from a synergistic interplay of chemically diverse metabolites rather than a single dominant compound. Although the biosynthetic pathways responsible for these metabolites were not experimentally validated in this study, their activation during the stationary growth phase likely reflects secondary metabolism associated with nutrient limitation and stress response, as widely reported in filamentous fungi.
To avoid overstatement, the discussion of sustainability has been moderated to note that solid-state fermentation represents a promising approach for eco-efficient metabolite production, aligning with principles of green chemistry, but further life-cycle analysis would be required to substantiate environmental benefits.
These findings align with and expand upon prior reports concerning B. rhodina and related Botryosphaeriaceae. Abdou et al. [30] identified Botryorhodines A-D, depsidone-type compounds exhibiting antifungal properties, from cultures of B. rhodina [30]. Despite the absence of these particular chemicals in the current investigation, the detection of structurally analogous phenolics and polyketides, including fraxetin and deoxybrevianamide E, implies similar metabolic potential. Moreover, in contrast to previous research cantered on targeted isolation, this untargeted metabolomics methodology revealed a wider and more varied chemical profile, especially under solid-state fermentation conditions that might replicate the fungusʼs natural habitat.
The genus Botryosphaeria has been inadequately represented in natural product research, with the majority of publications concentrating on its function as a plant pathogen or in co-culture inhibition experiments [41]. The present investigation identifies B. rhodina, particularly from a mangrove environment, as a significant source of chemically unique and pharmacologically advantageous metabolites. Environmental stresses typical of the mangrove ecosystem may stimulate the synthesis of adaptive secondary metabolites rarely observed in terrestrial strains, underscoring the ecological importance of endophyte-derived bioactives [42].
In conclusion, the compositional complexity, presence of structurally varied and pharmacologically active metabolites, and potential synergistic interactions elucidate the enhanced antibacterial efficacy of the methanol extract. These findings highlight the importance of metabolite profiling in understanding the role of endophytic fungi and discovering antimicrobial agents from natural sources.
Notwithstanding the thorough metabolite profiling and antimicrobial evaluations conducted in this work, certain limitations must be recognized. The identification of metabolites using LC-MS/MS and GC-MS relied mostly on spectrum library matching and in silico fragmentation predictions, lacking confirmation structural elucidation using NMR or MS/MS fragmentation for the unidentified molecules. This creates a limitation on the reliability of chemical identification, especially for metabolites with unclear or non-existent database correlations.
While this study successfully revealed a wide range of secondary metabolites from Botryosphaeria rhodina using untargeted LC-HRMS and GC-MS analyses, it is important to note that the compound identifications are putative and based solely on accurate mass and spectral database matching. Consequently, experimental evaluation of cytotoxicity or in silico pharmacokinetic assessment using tools such as SwissADME was not performed. Such predictive analyses require structurally confirmed compounds to ensure valid and reproducible results, whereas applying them to tentative chemical identities could lead to speculative or misleading interpretations. Therefore, the current findings should be viewed as a foundational step toward understanding the metabolic potential of B. rhodina. Future investigations will focus on bioassay-guided isolation and structure elucidation of the key bioactive metabolites, followed by comprehensive cytotoxicity, ADME, and drug-likeness evaluations through both experimental and computational approaches to substantiate their therapeutic relevance and safety profile.
It should be noted that all metabolite annotations presented herein are considered putative identifications (MSI level 2), derived from high-resolution MS data and spectral database matching. Definitive structural confirmation was not performed and will be addressed in future work.
Multivariate analysis (PLS-DA) of metabolite profiles
A Partial Least Squares Discriminant Analysis (PLS-DA) was conducted to quantify the connection between metabolite composition and antibacterial activity in the methanolic, and n-hexane extracts of Botryosphaeria rhodina. This chemometric method established a statistical framework to differentiate extract profiles based on metabolite composition and to find key discriminant metabolites that significantly contribute to the observed biological activities. The PLS-DA score plots (Figure 6) demonstrated a distinct separation between the methanolic (M) and n-hexane (H) extracts for all evaluated bacterial strains. The methanol extract exhibited a close clustering with L. monocytogenes, S. aureus, and B. subtilis, correlating with the most substantial inhibitory zones and the minimal MIC/MBC values seen in antimicrobial testing.
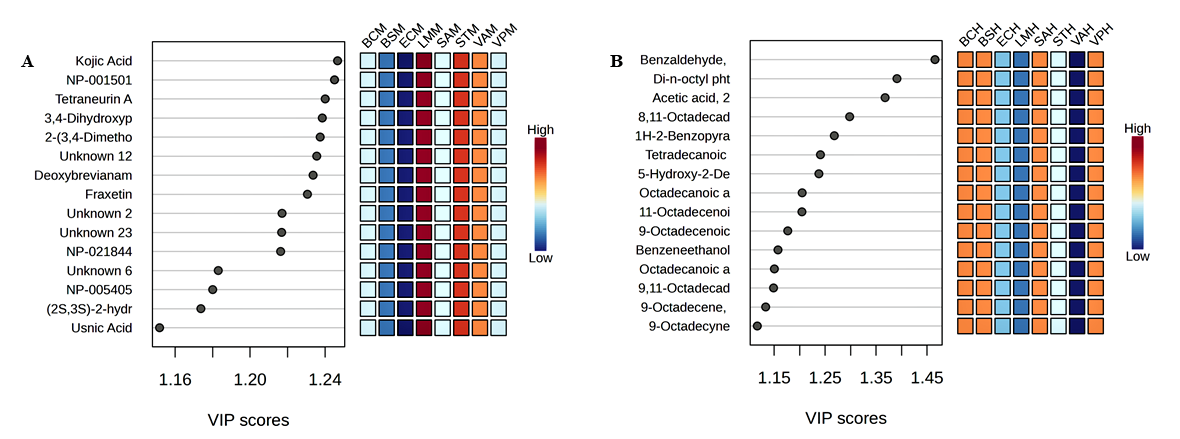
Figure 6 PLS-DA of Botryosphaeria rhodina extracts against pathogenic bacteria. (A) Methanol extract (M); (B) n-Hexane extract (H). Heatmaps illustrate the relative abundance of discriminant metabolites across bacterial strains: BC (Bacillus cereus), BS (Bacillus subtilis), EC (Escherichia coli), LM (Listeria monocytogenes), ST (Salmonella typhimurium), SA (Staphylococcus aureus), VA (Vibrio alginolyticus), and VP (Vibrio parahaemolyticus). VIP scores highlight metabolites contributing most to class separation.
The metabolites exhibiting the greatest Variable Importance in Projection (VIP) scores comprised kojic acid, glycitein, fraxetin, and deoxybrevianamide E. These polar chemicals are extensively documented for their antibacterial actions, which encompass membrane disruption, enzyme inhibition, and the generation of oxidative stress. Their robust correlation with Gram-positive bacteria indicates that they are the primary agents of the observed bactericidal activity.
Conversely, the n-hexane extract exhibited diminished clumping and a looser association with Escherichia coli and Vibrio spp., aligning with its restricted antibacterial efficacy. The distinguishing metabolites in this fraction were primarily fatty acids, sterols, and long-chain hydrocarbons, which are often noted for their poor or inconsistent antibacterial activity and contributed little to class differentiation.
PLS-DA exhibited clear grouping between methanolic, and n-hexane extracts for fungal infections (Figure 7). The methanolic extract exhibited significant antifungal activity against C. krusei, the most sensitive yeast strain evaluated. The discriminant metabolites with the highest VIP scores comprised usnic acid, vanillyl alcohol, and several flavonoid derivatives, recognized for their interference with ergosterol production and cell wall integrity. These processes are essential for antifungal effectiveness, especially in resistant Candida strains.
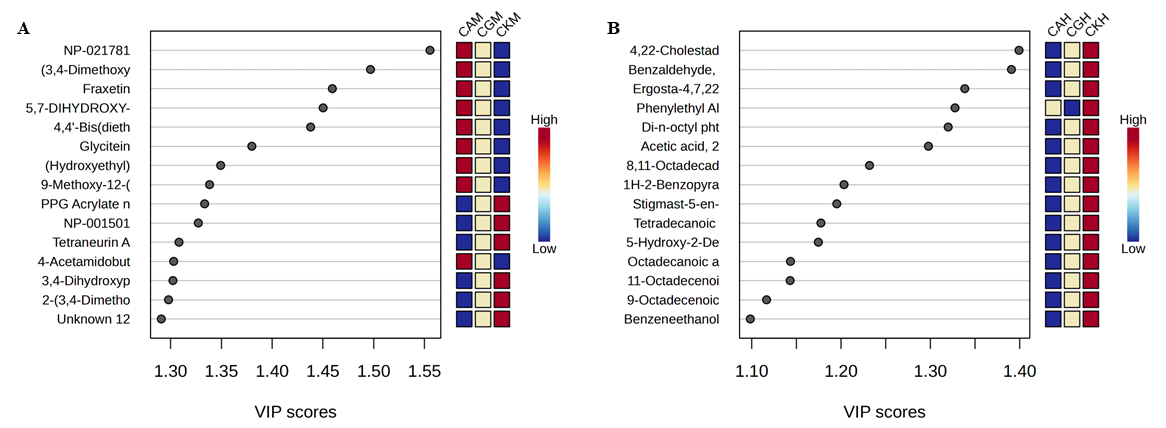
Figure 7 PLS-DA of Botryosphaeria rhodina extracts against Candida species. (A) Methanol extract (M); (B) n-Hexane extract (H). Heatmaps show the relative abundance of discriminant metabolites across Candida strains: CA (C. albicans), CG (C. glabrata), and CK (C. krusei). VIP scores indicate the most influential metabolites contributing to class separation in the PLS-DA model.
In contrast, the n-hexane extract created a distinct cluster primarily composed of lipophilic molecules. The metabolites contributed minimally to class differentiation and exhibited insignificant connection with antifungal efficacy, indicating the restricted bioactivity of the non-polar fraction. The amalgamation of PLS-DA results with antimicrobial testing substantiates that polar metabolites in the methanol extract are the primary factors influencing both antibacterial and antifungal activity. This result aligns with the enhanced inhibition zones, reduced MIC/MBC/MFC values, and the significant chemical variety identified by LC-MS/MS in the methanolic fraction. Furthermore, the findings underscore the significant impact of solvent polarity on the metabolomic profile of endophytic fungus. Methanol extracts featured a chemically diversified array of bioactive polar molecules, whereas n-hexane extracts predominantly comprised lipophilic metabolites, which contributed minimally to antibacterial activity.
The PLS-DA model not only emphasizes the distinctions across solvent fractions but also provides prediction insights by associating specific metabolites with bioactivity. Compounds exhibiting consistently elevated VIP scores—such as kojic acid, glycitein, fraxetin, deoxybrevianamide E, and usnic acid—represent potential subjects for additional research. Their consistent contribution to class differentiation in bacterial and fungal datasets highlights their potential as bioactive candidates and indicates probable synergistic effects within the methanolic extract. These discoveries offer a logical foundation for focused isolation, structural determination, and mechanistic investigations, hence enhancing the bioprospecting significance of B. rhodina.
The utilization of PLS-DA enhances our investigation beyond mere descriptive metabolite profiling by establishing a statistically validated link between metabolite identification and biological function. The discovered clustering patterns and discriminant metabolites underscore possible synergistic interactions within the methanolic extract, necessitating additional separation and combination studies.
Collectively, these data affirm that B. rhodina, especially via its methanol-derived polar metabolites, constitutes a prolific and chemically varied source of physiologically active secondary metabolites. Furthermore, the implementation of multivariate chemometric techniques like PLS-DA improves the dependability of natural product identification, hence augmenting the bioprospecting potential of endophytic fungus from mangrove habitats.
Biosynthetic origins of identified metabolites
The metabolite diversity in B. rhodina can be elucidated by associating the identified chemical classes with fungal biosynthetic pathways and gene clusters (BGCs). In filamentous fungi, secondary metabolites are frequently encoded by clustered genes, including polyketide synthases (PKSs), nonribosomal peptide synthetases (NRPSs), hybrid PKS-NRPSs, and terpene synthases (TSs). Global transcription factors and environmental signals tightly control these clusters. They make sure that metabolite production is in sync with fungal growth and adaptation to the environment [43].
The study identified compounds derived from polyketides and phenolics, including kojic acid, fraxetin, and usnic acid, which are generally biosynthesized via PKS pathways. Recent metabolomic and genomic studies have shown that PKS-driven BGCs are still very important for the production of fungal aromatic polyketides. These BGCs are often turned on when nutrients are scarce or when oxidative stress is present. Usnic acid and structurally analogous dibenzofuran polyketides are associated with aromatic PKSs in conjunction with oxidative tailoring enzymes [44]. These pathways elucidate the enrichment of bioactive phenolic compounds in the methanol extract of B. rhodina.
The identification of flavonoid-like metabolites, such as glycitein, is notably intriguing due to their traditional correlation with plants. Nonetheless, recent investigations have demonstrated that endophytic fungi may possess PKS enzymes exhibiting chalcone synthase-like activity, potentially obtained via horizontal gene transfer or ecological adaptation to plant hosts [45,46]. This may elucidate the occurrence of isoflavonoid derivatives in mangrove-derived B. rhodina, indicating adaptation to its intricate ecological niche.
Indole alkaloids and diketopiperazines, such as deoxybrevianamide E, are typically synthesized by NRPSs or PKS-NRPS hybrids. Genome mining of fungal NRPS clusters has identified conserved pathways for indole alkaloid biosynthesis, especially within genera associated with Aspergillus and Penicillium [47]. The presence of deoxybrevianamide E in B. rhodina corroborates the hypothesis that similar NRPS-mediated pathways are operational in this species.
Moreover, depsidone-type metabolites, including botryorhodines previously identified in B. rhodina, constitute an additional category of PKS-derived compounds necessitating modifications mediated by methyltransferases and oxidases [48]. Even though we didnʼt find these in this study, the metabolic capacity shown by LC-MS/MS profiling suggests that B. rhodina may still have silent or conditionally expressed PKS clusters that can make depsidone.
The nonpolar fraction, which is rich in sterols and terpenoids (like clionasterol, squalene, and ergosta-4,7,22-trien-3-one), shows that the mevalonate pathway and fungal TS clusters are working. Recent research has underscored the ecological functions of fungal terpenoids in membrane stability, oxidative stress responses, and antifungal resistance [49]. Finding them here supports the idea that making terpenoids helps with both ecological fitness and the ability to fight off bacteria.
At the regulatory level, these biosynthetic processes are regulated by global transcriptional complexes like LaeA and the Velvet family proteins. Recent research validates that these regulators amalgamate environmental signals—such as light, salinity, and nutrient stress—to simultaneously activate or inhibit various BGCs [50]. The dynamic mangrove habitat of B. rhodina, characterized by salinity variations and microbial competition, may serve as a natural catalyst for the activation of cryptic or stress-responsive gene clusters, elucidating the diverse metabolome observed during solid-state fermentation.
In summary, the metabolite classes found in B. rhodina can be directly connected to PKS, NRPS, and TS gene clusters, as well as global regulatory networks. This molecular viewpoint emphasizes the genetic and ecological factors influencing metabolite diversity and underscores the importance of combining metabolomics with genome mining to discover new antimicrobial agents.
Limitations and future perspectives
The current work offers new insights into the antibacterial properties and metabolite composition of B. rhodina; nonetheless, numerous limitations require careful attention. Initially, metabolite identification predominantly depended on LC-HRMS and GC-MS data analyzed via spectrum libraries and in silico fragmentation predictions. This method facilitated the annotation of several metabolites; nevertheless, the lack of confirmatory tests, such as NMR or targeted HR-MS/MS, limits the confidence in structural assignments. This constraint is especially significant for over 25 unidentified metabolites detected in high relative abundance, several of which may signify novel chemical scaffolds. The exact function of these chemicals in antibacterial action remains uncertain without comprehensive structural elucidation.
Secondly, while the amalgamation of antimicrobial bioassays with PLS-DA analysis enhanced the correlation between metabolite composition and biological activity, the established relationships are qualitative rather than quantitative. The utilization of crude extracts hinders the assignment of activity to individual compounds and fails to consider possible synergistic or antagonistic interactions among metabolites. Bioassay-guided fractionation and purification are crucial for validating the roles of the discriminant metabolites found using chemometric modeling. Furthermore, synergy testing and combination assays should be utilized to elucidate the interactions among various metabolites that yield the observed antibacterial and antifungal activities.
The antimicrobial assessment was confined to a specific panel of bacterial and fungal pathogens. The results indicate substantial efficacy, especially against Gram-positive bacteria and Candida species; however, the effectiveness of B. rhodina metabolites against multidrug-resistant strains or clinically significant pathogens such as Pseudomonas aeruginosa, Acinetobacter baumannii, or Cryptococcus neoformans has yet to be investigated. Broadening the range of test organisms is essential for a more thorough evaluation of the therapeutic potential of these metabolites.
This study ultimately excluded genomic or transcriptome investigations, which could yield more profound insights into the biosynthetic gene clusters responsible for metabolite production. Integrating metabolomic data with genome mining and transcriptomics would enable the identification of cryptic or silent gene clusters and assist in the strategic modification of culture conditions to optimize the production of target metabolites. Subsequent research should examine co-culture or stress-induced methodologies to activate dormant biosynthetic pathways.
Collectively, these constraints underscore significant directions for future inquiry. Comprehensive structural elucidation, bioassay-guided isolation, synergy assessment, extensive antimicrobial screening, and integrative omics methodologies will be essential for progressing B. rhodina from a potential bioprospecting candidate to a confirmed source of new antimicrobial drugs.
This study highlights the significance of metabolites generated from endophytes for sustainable chemistry and pharmacy, in addition to their therapeutic consequences. The employment of endophytic fungus as sustainable microbial resources diminishes the necessity for massive plant harvesting, therefore preserving delicate ecosystems like mangroves. Our methodology—solid-state fermentation utilizing rice substrate—exhibits a low-energy, ecologically friendly, and scalable technique for metabolite production, in stark contrast to the resource-intensive submerged fermentation process. This study identifies bioactive molecules that may operate as environmentally friendly alternatives to synthetic antimicrobials, thereby alleviating the chemical and environmental loads linked to traditional medication manufacture. Our findings align with the tenets of green chemistry and directly endorse the United Nations Sustainable Development Goals, including SDG 3 (Good Health and Well-being) and SDG 12 (Responsible Consumption and Production). Consequently, the findings of this research transcend natural product discovery, illuminating a route toward a more sustainable bio-circular economy in pharmaceutical development.
Conclusions
The compositional diversity and pharmacological potential of B. rhodina extracts underscore the importance of mangrove-derived endophytes as sustainable resources for antimicrobial drug discovery. The methanol extract, rich in polar secondary metabolites including kojic acid, usnic acid, fraxetin, and glycitein, exhibited significant antibacterial and mild antifungal properties, while chemometric tests indicated distinct correlations between metabolite profiles and bioactivity. The application of solid-state fermentation on rice substrate offers a low-energy, scalable, and ecologically friendly manufacturing method that adheres to the principles of green chemistry. This work promotes the use of endophytic fungi as sustainable microbiological sources, aiding the creation of environmentally friendly alternatives to synthetic antimicrobials, thereby advancing pharmaceutical innovation and biodiversity preservation.
Acknowledgements
This research was funded by the Directorate of Research and Community Service, Ministry of Higher Education, Science, and Technology of the Republic of Indonesia, under contract number 102/C3/DT.05.00/PL/2025. The authors also wish to express their sincere gratitude to the Institute for Research and Community Service, Universitas Riau, Pekanbaru 28293, Indonesia for their valuable support and facilitation throughout this study.
Declaration of Generative AI in Scientific Writing
The author(s) employed the QuillBot tool solely to check the grammar and sentence structure while writing this manuscript. There was no AI tools used to create content, rephrase it, or analyze data. After using this tool, the author(s) carefully edited and reviewed the whole manuscript to make sure it was correct and clear. They take full responsibility for the content and conclusions in this publication.
CRediT Author Statement
Rudi Hendra: Conceptualization; Writing - Original Draft; Funding. Muhammad Rohim: Investigation; formal analysis. Yuli Haryani: methodology; supervision. Ari Satia Nugraha: Writing - Original Draft; Writing - Review & Editing. Hilwan Yuda Terun: Writing - Review & Editing. Fauzan Zein Muttaqin: Validation; software; visualization.
References
[1] NC Baron and EC Rigobelo. Endophytic fungi: A tool for plant growth promotion and sustainable agriculture. Mycology 2022; 13(1), 39-55.
[2] R Grabka, TW d’Entremont, SJ Adams, AK Walker, JB Tanney, PA Abbasi and S Ali. Fungal endophytes and their role in agricultural plant protection against pests and pathogens. Plants 2022; 11(3), 384.
[3] GV Borkunov, EV Leshchenko, DV Berdyshev, RS Popov, EA Chingizova, NP Shlyk, AV Gerasimenko, NN Kirichuk, YV Khudyakova, VE Chausova, AS Antonov, AI Kalinovsky, AR Chingizov, EA Yurchenko, MP Isaeva and AN Yurchenko. New piperazine derivatives helvamides B-C from the marine-derived fungus Penicillium velutinum ZK-14 uncovered by OSMAC (One Strain Many Compounds) strategy. Natural Products and Bioprospecting 2024; 14, 32.
[4] MSM Al-Saleem, WHB Hassan, ZIE Sayed, MM Abdel-Aal, WM Abdel-Mageed, E Abdelsalam and S Abdelaziz. Metabolic profiling and in vitro assessment of the biological activities of the ethyl acetate extract of Penicillium chrysogenum “endozoic of Cliona sp. marine sponge” from the Red Sea (Egypt). Marine Drugs 2022; 20(5), 326.
[5] H Jia, L Wu, R Liu, J Li, L Liu, C Chen, J Li, K Zhang, J Liao and Y Long. Penifuranone A: A novel alkaloid from the mangrove endophytic fungus Penicillium crustosum SCNU-F0006. International Journal of Molecular Sciences 2024; 25(9), 5032.
[6] Y Yang, X Han, C Jiang, Y Chen, X Tang and G Li. Penisimplinoids A-K, highly oxygenated andrastin-type meroterpenoids with diverse activities from the marine-derived fungus Penicillium simplicissimum. Bioorganic Chemistry 2024; 153, 107897.
[7] A Baazeem, A Almanea, P Manikandan, M Alorabi, P Vijayaraghavan and A Abdel-Hadi. In vitro antibacterial, antifungal, nematocidal and growth promoting activities of Trichoderma hamatum FB10 and its secondary metabolites. Journal of Fungi 2021; 7(5), 331.
[8] SD Sasmito, M Basyuni, A Kridalaksana, MF Saragi-Sasmito, CE Lovelock and D Murdiyarso. Challenges and opportunities for achieving sustainable development goals through restoration of Indonesia’s mangroves. Nature Ecology & Evolution 2023; 7, 62-70.
[9] TR Nurunnabi, F Sabrin, DI Sharif, L Nahar, H Sohrab, SD Sarker, SMM Rahman and M Billah. Antimicrobial activity of endophytic fungi isolated from the mangrove plant Sonneratia apetala (Buch.-Ham) from the Sundarbans mangrove forest. Advances in Traditional Medicine 2020; 20, 419-425.
[10] ARB Ola, CAP Soa, YosephSugi, TD Cunha, HLL Belli and HJD Lalel. Antimicrobial metabolite from the endophytic fungi Aspergillus flavus isolated from Sonneratia alba, a mangrove plant of Timor-Indonesia. RASĀYAN Journal of Chemistry 2020; 13(1), 377-381.
[11] Y Haryani, R Hilma, N Delfira, T Martalinda, F Puspita, A Friska, D Juwita, A Farniga and F Ardi. Potential antibacterial activity of endophytic fungi Penicillium sp. and Trichoderma sp. derived from mangrove Ceriops tagal (Perr.) C.B.Robb and Bruguiera sp. Journal of Physics: Conference Series 2019; 1351, 012100.
[12] Y Haryani, R Hilma, N Delfira, T Martalinda, F Puspita, A Friska, D Juwita and F Ardi. Anti-vibriosis activity of endophytic fungi associated with Ceriops tagal (Perr.) C.B.Rob and Bruguiera sp., mangrove plants from Riau Province, Indonesia. AIP Conference Proceedings 2020; 2243, 030006.
[13] J Kjer, A Debbab, AH Aly and P Proksch. Methods for isolation of marine-derived endophytic fungi and their bioactive secondary products. Nature Protocols 2010; 5(3), 479-90.
[14] MK Army, R Khodijah, Y Haryani, HY Teruna and R Hendra. Antibacterial in vitro screening of Helminthostachys zeylanica (L.) Hook. root extracts. Journal of Pharmacy & Pharmacognosy Research 2023; 11(2), 291-296.
[15] R Hendra, MK Army, Y Nurulita and HY Teruna. Elucidating the phytochemical profiles and antibacterial potential of Helminthostachys zeylanica (L.) Hook. through untargeted LC-HRMS metabolomics analysis. Journal of Pharmacy & Pharmacognosy Research 2025; 13(4), 1071-109.
[16] R Khodijah, Y Haryani, HY Teruna and R Hendra. Antibacterial properties of Pyrrosia longifolia extracts. Pharmacy Education 2023; 23(2), 168-173.
[17] R Hendra, A Agustha, N Frimayanti, R Abdulah and HY Teruna. Antifungal potential of secondary metabolites derived from Arcangelisia flava (L.) Merr.: An analysis of in silico enzymatic inhibition and in vitro efficacy against Candida species. Molecules 2024; 29(10), 2373.
[18] A Agustha, S Fadhillah, HY Teruna and R Hendra. Comparative analysis of antifungal properties in diverse extracts of Arcangelisia flava (L.). Pharmacy Education 2024; 24(2), 99-103.
[19] G Nickles, I Ludwikoski, JW Bok and NP Keller. Comprehensive guide to extracting and expressing fungal secondary metabolites with Aspergillus fumigatus as a case study. Current Protocols 2021; 1(12), 321.
[20] G Camilli, M Blagojevic, JR Naglik and JP Richardson. Programmed cell death: Central player in fungal infections. Trends in Cell Biology 2021; 31(3), 179-196.
[21] JE Méndez-Hernández, LV Rodríguez-Durán, JB Páez-Lerma and NO Soto-Cruz. Strategies for supplying precursors to enhance the production of secondary metabolites in solid-state fermentation. Fermentation 2023; 9(9), 804.
[22] V Chaudhary, P Katyal, AK Puniya, H Panwar, M Arora, J Kaur, N Rokana, N Wakchaure, A Raposo, D Raheem and AK Poonia. Pilot-scale process to produce bio-pigment from Monascus purpureus using broken rice as substrate for solid-state fermentation. European Food Research and Technology 2023; 249(7), 1845-1855.
[23] K Wadhwa, N Kapoor, H Kaur, EA Abu-Seer, M Tariq, S Siddiqui, VK Yadav, P Niazi, P Kumar and S Alghamdi. A comprehensive review of the diversity of fungal secondary metabolites and their emerging applications in healthcare and environment. Mycobiology 2024; 52(6), 335-387.
[24] M Xue, X Hou, J Fu, J Zhang, J Wang, Z Zhao, D Xu, D Lai and L Zhou. Recent advances in search of bioactive secondary metabolites from fungi triggered by chemical epigenetic modifiers. Journal of Fungi 2023; 9(2), 172.
[25] AS Abdelbaky. Effect of various extraction methods and solvent types on yield, phenolic and flavonoid content and antioxidant activity of Spathodea nilotica leaves. Egyptian Journal of Chemistry 2021; 64(12), 7583-7589.
[26] I Gutiérrez-Del-Río, S López-Ibáñez, P Magadán-Corpas, L Fernández-Calleja, Á Pérez-Valero, M Tuñón-Granda, EM Miguélez, CJ Villar and F Lombó. Terpenoids and polyphenols as natural antioxidant agents in food preservation. Antioxidants 2021; 10(8), 1264.
[27] W Huang, Y Wang, W Tian, X Cui, P Tu, J Li, S Shi and X Liu. Biosynthesis investigations of terpenoid, alkaloid, and flavonoid antimicrobial agents derived from medicinal plants. Antibiotics 2022; 11(10), 1380.
[28] T Manso, M Lores and T de Miguel. Antimicrobial activity of polyphenols and natural polyphenolic extracts on clinical isolates. Antibiotics 2021; 11(1), 46.
[29] S Ghosh, S Basu, A Anbarasu and S Ramaiah. A comprehensive review of antimicrobial agents against clinically important bacterial pathogens: Prospects for phytochemicals. Phytotherapy Research 2025; 39(1), 138-161.
[30] R Abdou, K Scherlach, H-M Dahse, I Sattler and C Hertweck. Botryorhodines A-D, antifungal and cytotoxic depsidones from Botryosphaeria rhodina, an endophyte of the medicinal plant Bidens pilosa. Phytochemistry 2010; 71(1), 110-116.
[31] V Karličić, M Zlatković, J Jovičić-Petrović, MP Nikolić, S Orlović and V Raičević. Trichoderma spp. from pine bark and pine bark extracts: Potent biocontrol agents against Botryosphaeriaceae. Forests 2021; 12(12), 1731.
[32] SL Collins, I Koo, JM Peters, PB Smith and AD Patterson. Current challenges and recent developments in mass spectrometry-based metabolomics. Annual Review of Analytical Chemistry 2021; 14(1), 467-487.
[33] J-X Ye, Y-F Qian, W-Q Lan and J Xie. Evaluation of antibacterial and antibiofilm properties of kojic acid against Aeromonas sobria and Staphylococcus saprophyticus. Journal of Food Quality 2023; 2023, 2531438.
[34] T Li, Z Wang, X Zhang, Z Hao, Y Guo, J Shen, T Velkov and C Dai. Natural product usnic acid as an antibacterial therapeutic agent: Current achievements and further prospects. ACS Infectious Diseases 2025; 11(6), 1403-1415.
[35] NM Ha and NT Son. Health benefits of fraxetin: From chemistry to medicine. Archiv der Pharmazie 2024; 357(7), 2400092.
[36] B Zieniuk, K Jasińska, K Wierzchowska and A Fabiszewska. Enzymatic synthesis of flavours and fragrances, antioxidants and antimicrobials on the example of benzyl alcohol and its selected derivatives. Biology and Life Sciences Forum 2022; 18(1), 2.
[37] X Zhao, X Zhong, S Yang, K Deng, L Liu, X Song, Y Zou, L Li, X Zhou, R Jia, J Lin, H Tang, G Ye, J Yang, S Zhao, Y Lang, H Wan, Z Yin and OP Kuipers. Elucidating the mechanism of action of the Gram-negative-pathogen-selective cyclic antimicrobial lipopeptide brevicidine. Antimicrob Agents Chemother 2023; 67(5), 0001023.
[38] KT Yuyama, M Rohde, G Molinari, M Stadler and WR Abraham. Unsaturated fatty acids control biofilm formation of Staphylococcus aureus and other gram-positive bacteria. Antibiotics 2020; 9(11), 788.
[39] N Soujanya, BS Suresha, T Balasubramanian, S Rakshitha and BP Meghana. Review on kojic acid-a secondary metabolite from Aspergillus fungi. Asian Journal of Pharmaceutical and Health Sciences 2022; 12(3), 2707-2712.
[40] Y Liu, J Zhu, Z Liu, Y Zhi, C Mei and H Wang. Flavonoids as promising natural compounds for combating bacterial infections. International Journal of Molecular Sciences 2025; 26(6), 2455.
[41] CG Kumar. New and future developments in microbial biotechnology and bioengineering. Elsevier, London, 2020, p. 167-286.
[42] G Zhou, J Cai, B Wang, W Diao, Y Zhong, S Pan, W Xiong, G Huang and C Zheng. Secondary metabolites from the mangrove ecosystem-derived fungi Penicillium spp.: Chemical diversity and biological activity. Marine Drugs 2024; 23(1), 7.
[43] JM Galindo-Solís and FJ Fernández. Endophytic fungal terpenoids: Natural role and bioactivities. Microorganisms 2022; 10(2), 339.
[44] J Wang, R Zhang, X Chen, X Sun, Y Yan, X Shen and Q Yuan. Biosynthesis of aromatic polyketides in microorganisms using type II polyketide synthases. Microbial cell factories 2020; 19(1), 110.
[45] X Wu, K Zou, X Liu, S Fu, S Zhang, Z Duan, J Zhou and Y Liang. The novel distribution of intracellular and extracellular flavonoids produced by Aspergillus sp. Gbtc 2, an endophytic fungus from Ginkgo biloba root. Frontiers in Microbiology 2022; 13, 972294.
[46] P Tiwari and H Bae. Horizontal gene transfer and endophytes: An implication for the acquisition of novel traits. Plants 2020; 9(3), 305.
[47] B Wei, TT Ying, HW Lv, ZY Zhou, H Cai, GA Hu, HM Liang, WC Yu, YL Yu, AL Fan, K Hong, XN Li and H Wang. Global analysis of fungal biosynthetic gene clusters reveals the diversification of diketopiperazine biosynthesis. Bioresource Technology 2025; 422, 132218.
[48] Q Ji and Y Matsuda. Biosynthesis of depsides, depsidones, and diphenyl ethers from fungi and lichens. Asian Journal of Organic Chemistry 2025; 14(2), 202400451.
[49] RA González-Hernández, NA Valdez-Cruz, ML Macías-Rubalcava and MA Trujillo-Roldán. Overview of fungal terpene synthases and their regulation. World Journal of Microbiology and Biotechnology 2023; 39(7), 194.
[50] W Yu, R Pei, J Zhou, B Zeng, Y Tu and B He. Molecular regulation of fungal secondary metabolism. World Journal of Microbiology and Biotechnology 2023; 39(8), 204.