
Trends
Sci.
2026; 23(6): 12653
Stem Cell Physiology and Cellular Differentiation: Molecular Insights and Therapeutic Potential
Aswin
Rafif Khairullah1,
![]() ,
Saifur
Rehman2,
,
Saifur
Rehman2,
![]() ,
Imam
Mustofa3,
,
Imam
Mustofa3,
![]() ,
Bima Putra Pratama4,
,
Bima Putra Pratama4,
![]() ,
,
Eny
Martindah1,
![]() ,
Arif Nur Muhammad Ansori5,6,7,
,
Arif Nur Muhammad Ansori5,6,7,
![]() ,
Mohammad Sukmanadi8,*,
,
Mohammad Sukmanadi8,*,
![]() ,
,
Ilma
Fauziah Ma’ruf9,
![]() ,
Analinasari Analinasari10,
,
Analinasari Analinasari10,
![]() ,
Riza Zainuddin Ahmad1,
,
Riza Zainuddin Ahmad1,
![]() ,
,
Sri
Suryatmiati Prihandani1,
![]() ,
Bantari
Wisynu Kusuma Wardhani9,
,
Bantari
Wisynu Kusuma Wardhani9,
![]() ,
Siti Hamidatul Aliyah12,
,
Siti Hamidatul Aliyah12,
![]() ,
Wita Yulianti13,
,
Wita Yulianti13,
![]() ,
Dea Anita Ariani Kurniasih14,
,
Dea Anita Ariani Kurniasih14,
![]() and Fadhila Utari9,
and Fadhila Utari9,
![]()
1Research Center for Veterinary Science, National Research and Innovation Agency (BRIN), Bogor,
West Java 16911, Indonesia
2Department of Pathobiology, Faculty of Veterinary and Animal Sciences, Gomal University,
Dera Ismail Khan 27000, Pakistan
3Division of Veterinary Reproduction, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya,
East Java 60115, Indonesia
4Research Center for Process Technology, National Research and Innovation Agency (BRIN), South Tengrang,
Banten 15310, Indonesia
5Postgraduate School, Universitas Airlangga, Surabaya, East Java 60286, Indonesia
6Uttaranchal Institute of Pharmaceutical Sciences, Uttaranchal University, Uttarakhand 248007, India
7Medical Biotechnology Research Group, Virtual Research Center for Bioinformatics and Biotechnology,
Surabaya, East Java 60493, Indonesia
8Division of Basic Veterinary Medicine, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya,
East Java 60115, Indonesia
9Research Center for Pharmaceutical Ingredients and Traditional Medicine, National Research and Innovation Agency (BRIN), Bogor, West Java 16911, Indonesia
10Postgraduate Program in Applied Master of Food Security, Lampung State Polytechnic (Polinela), Bandar Lampung, Lampung 35144, Indonesia
12Center for Biomedical Research, National Research and Innovation Agency (BRIN), Bogor,
West Java 16911, Indonesia
13Research Center for Biota Systems, National Research and Innovation Agency (BRIN), Bogor,
West Java 16911, Indonesia
14Research Center for Public Health and Nutrition, National Research and Innovation Agency (BRIN), Bogor,
West Java 16911, Indonesia
(*Corresponding author’s e-mail: [email protected])
Received: 13 November 2025, Revised: 20 December 2025, Accepted: 30 December 2025, Published: 30 January 2026
Abstract
Stem cells are unique cells capable of maintaining self-renewal and differentiating into various specialized cell types. These properties make them crucial components in tissue physiology, homeostasis, and regeneration. Stem cell differentiation is regulated through a complex interplay of molecular signaling pathways, transcription factors, microRNAs, and epigenetic mechanisms. Key pathways such as Wnt, Notch, Hedgehog, and BMP/TGF-β play a role in determining cell fate, regulating proliferation, and balancing self-renewal and specialization. Proper regulation ensures that stem cells transition to specific phenotypes in a directed, stable, and physiologically appropriate manner, while disruption of regulation can lead to tissue degeneration or tumors. Understanding these molecular mechanisms paves the way for innovative regenerative therapies. Pluripotent and multipotent cells are used for cell replacement strategies in degenerative diseases, organ injuries, and hematopoietic disorders. iPSCs allow the engineering of a patientʼs own cells to minimize the risk of immunoreactions, while MSCs possess immunomodulatory capabilities and secrete paracrine factors that support endogenous tissue repair. Technologies such as 3D organoid culture, biomaterial scaffolds, and gene editing improve control over differentiation, tissue integration, and therapeutic safety before clinical application. Stem cells and cellular differentiation mechanisms form a solid scientific foundation for the development of modern regenerative therapies. Ongoing research in this area is expected to open new opportunities for tissue function restoration, the treatment of degenerative diseases, and the implementation of safe, effective, and precision therapeutic strategies.
Keywords: Stem cells, Cellular differentiation, Disease, Molecular signaling, Regenerative medicine
Introduction
Stem cells are specialized cells with the ability to maintain self-renewal while simultaneously differentiating into various specific cell types [1]. This property makes them a crucial component in tissue physiology, maintaining homeostasis, and supporting regeneration after injury [2]. Based on their differentiation potential, stem cells are grouped into several categories. Pluripotent stem cells (PSCs), including embryonic stem cells (ESCs) and induced pluripotent stem cells (iPSCs), are capable of forming all somatic cell types [3]. Meanwhile, multipotent stem cells, such as mesenchymal stem cells (MSCs) and hematopoietic stem cells (HSCs), can only generate a few cell types within a specific tissue lineage [4]. Furthermore, adult stem cells reside in specific tissues and play a role in maintaining homeostasis and local regeneration [5]. In contrast, ESCs isolated from blastocysts have broader differentiation capabilities, but their use faces ethical issues and the risk of tumorigenesis [6].
The ability of stem cells to differentiate in a directed manner is a crucial foundation for cellular physiology and tissue regeneration [7]. This differentiation process is a complex mechanism regulated by molecular signaling pathways, transcription factors, microRNAs, and epigenetic mechanisms such as Deoxyribonucleic Acid (DNA) methylation and histone modifications [8]. Key pathways, including Wingless-related integration site (Wnt), Notch, Hedgehog, and Bone Morphogenetic Protein/Transforming Growth Factor-beta (BMP/TGF-β), influence cell fate determination, triggering the transition from self-renewal to specialization, and maintaining the balance between proliferation and apoptosis [9]. These mechanisms are not only crucial during embryonic development but also play a role in adult tissue regeneration and the maintenance of organ function [10]. Disruption of the regulation of stem cell differentiation can trigger various pathological conditions, including regeneration failure, organ degeneration, and tumorigenesis [11].
In the realm of regenerative therapy, understanding the physiology and mechanisms of stem cell differentiation has significant clinical implications [12]. Stem cells are being utilized to design therapeutic strategies for a variety of conditions, including cardiovascular disease, neurodegenerative disorders, organ injury, and hematopoietic disorders [13]. iPSCs and ESCs allow for the engineering of a patient’s own cells, minimizing the risk of immune reactions, while MSCs possess immunomodulatory capabilities and produce paracrine factors that support endogenous tissue regeneration [14]. Furthermore, technological advances such as 3D organoid culture, biomaterial scaffolds, and gene editing facilitate the regulation of differentiation, control of cell integration, and functional testing prior to in vivo transplantation, thereby improving the safety and effectiveness of clinical therapies [15].
Alongside these developments, research also highlights the importance of using in vitro and in vivo experimental models to understand stem cell physiology and differentiation mechanisms [16]. 2D monolayer culture systems, 3D organoids, co-culture systems, and xenotransplantation animal models enable the analysis of molecular pathways, niche interactions, and regenerative capacity of stem cells under conditions more closely resembling natural physiology [17]. This approach is crucial for assessing efficacy, tissue integration, and potential risks of tumorigenesis before clinical application [18].
This review aims to provide a comprehensive overview of stem cell physiology and cellular differentiation mechanisms, focusing on molecular pathways, epigenetic regulation, and interactions with the microenvironment. It also assesses their therapeutic potential and clinical challenges. This approach is expected to provide a comprehensive understanding of the role of stem cells in maintaining tissue homeostasis, organ regeneration, and the development of modern regenerative therapies. It also identifies opportunities for future research and clinical applications.
Data collection method
A narrative, theory-anchored review was conducted to synthesize advances in stem cell physiology, differentiation mechanisms, and therapeutic applications. Literature was identified via PubMed/MEDLINE, Scopus, Web of Science, and targeted Google Scholar searches for peer-reviewed articles published from January 2016 to November 2025. Boolean combinations of keywords/MeSH terms covered signaling pathways (Wnt/Notch/Hedgehog/BMP-TGF-β), core factors (OCT4/SOX2/NANOG), epigenetic regulation, experimental models (2D, organoids, animal, clinical), bioengineering (biomaterials, bioprinting, CRISPR), and target organs (heart, CNS, liver). Inclusion encompassed primary studies, reviews, meta-analyses, guidelines, and methods relevant to stem cell identity, signaling/epigenetics, niche-mechanics-metabolism, and translational safety; exclusions were non-peer-reviewed preprints (unless corroborated), editorial pieces without data, and off-topic studies. Two-stage screening with reference snowballing was applied; overlapping datasets prioritized the most comprehensive or recent source. Data were extracted on design, model/system, key pathways, epigenetic/metabolic readouts, and translational endpoints and synthesized across five axes (pluripotency control, signaling crosstalk, niche-metabolism coupling, experimental models, efficacy/safety). Bias was mitigated by emphasizing high-quality evidence and cross-validation, while topic breadth and model heterogeneity limited meta-analysis and leave rapidly evolving areas potentially undercovered.
Stem cell physiology
Stem cell physiology reflects the unique ability of cells to maintain their identity through self-renewal, while maintaining pluripotent or multipotent potential, which is governed by cell cycle regulation, transcriptional signaling pathways, and complex epigenetic mechanisms.
Basic characteristics of stem cells
Stem cells are characterized by two key properties that underlie their role in tissue physiology and regenerative therapies: Self-renewal and differentiation potential [19]. Self-renewal allows stem cells to divide symmetrically or asymmetrically, thus maintaining the stem cell population without losing its identity [20]. This process is regulated by a complex network of molecular signals, including the Wnt/β-catenin, Notch, and Phosphoinositide 3-kinase/Protein Kinase B (PI3K/AKT) pathways, which control cell proliferation while preventing premature differentiation [21]. Furthermore, epigenetic regulation plays a crucial role through histone modifications, DNA methylation, and microRNAs, ensuring that the expression of key genes that determine stem cell identity is maintained according to physiological needs [22].
Differentiation potential reflects the ability of stem cells to produce offspring with specific phenotypes [23]. Based on their level of potency, stem cells are classified as: Pluripotent, which can form almost all types of somatic cells; multipotent, which is limited to a few specific cell lineages; and unipotent, which produces only one specialized cell type [24]. For example, embryonic stem cells (ESCs) are pluripotent, while MSCs are multipotent and can differentiate into osteoblasts, chondrocytes, and adipocytes [25]. Furthermore, interactions with the microenvironmental niche and external signals also determine the direction of differentiation, so that phenotypic changes occur in a coordinated and adaptive manner according to tissue needs [26].
The balance between self-renewal and differentiation is crucial for maintaining tissue homeostasis [27]. Imbalances in this regulation can trigger excessive cell proliferation or reduced regenerative capacity, potentially leading to degenerative diseases and tumorigenesis [28].
Cell cycle regulation and homeostasis mechanisms
The cell cycle is a coordinated series of stages that allow stem cells to maintain self-renewal while preparing for differentiation when needed [29]. In stem cells, cell cycle regulation not only controls proliferation but also maintains genome integrity and the balance between symmetric and asymmetric division [30]. The major stages of the cell cycle—Gap 1 phase (G1), Synthesis phase (S), Gap 2 phase (G2), and Mitosis phase (M)—are regulated by complex interactions between cyclin-dependent kinases (CDKs), cyclins, and CDK inhibitors (CKIs), which act as checkpoints to prevent the proliferation of cells experiencing DNA damage or mitotic errors [31].
Stem cells have a relatively short G1 phase, allowing rapid proliferation while maintaining differentiation capacity [32]. Molecular signaling pathways, such as PI3K/AKT, Mitogen-Activated Protein Kinase/Extracellular Signal-Regulated Kinase (MAPK/ERK), and Notch, play a role in regulating CDK and cyclin activity, allowing stem cells to balance proliferation and maintain their identity [33]. DNA checkpoint mechanisms, including Tumor protein p53 (p53) and Ataxia Telangiectasia Mutated/ATM and Rad3-related (ATM/ATR) activation, ensure that cells with severe genetic damage undergo apoptosis or premature differentiation, thus minimizing the risk of mutation and tumorigenesis [34].
Stem cell homeostasis is also influenced by interactions with the microenvironmental niche, which provides autocrine and paracrine signals to maintain the stem cell population in a quiescent or active state [35]. In the quiescent state, stem cells are in the Gap 0 phase (G0), minimizing excessive division while maintaining regenerative capacity [36]. Activation of stem cells from the quiescent state occurs in response to physiological needs or tissue injury, controlled by a combination of growth factor signals, extracellular matrix, and mechanotransduction [37].
In addition to cell cycle regulation, stem cell homeostasis also relies on epigenetic and metabolic mechanisms [38]. Histone modifications and DNA methylation regulate the expression of genes associated with proliferation and differentiation, while metabolic pathways, such as glucose oxidation and oxidative phosphorylation, adjust energy availability to the cellʼs specific conditions [39]. The combination of cell cycle regulation, niche signals, and epigenetic control ensures that stem cells maintain their identity, genetic integrity, and regenerative capacity, which underpins stem cell physiology and stem cell-based therapeutic applications [40].
Transcription factors and signaling networks that maintain stem cell identity
Stem cell identity is controlled by tight transcriptional regulation, with nuclear transcription factors acting as master regulators in maintaining pluripotency and self-renewal [41]. In pluripotent stem cells, factors such as Octamer-binding transcription factor 4 (OCT4), SRY-related HMG-box gene 2 (SOX2), and NANOG form a transcriptional network that interacts both autoregulatory and cross-regulatory [42]. OCT4 maintains pluripotent status by activating genes that promote self-renewal and suppressing premature differentiation pathways [43]. SOX2 works synergistically with OCT4 to maintain the expression of key target genes, while NANOG strengthens the stability of the transcriptional network and increases resistance to external differentiation signals [44]. Table 1 summarizes the nuclear transcription factors and the molecular signaling networks that play a role in maintaining pluripotent stem cell identity.
In addition to nuclear transcription factors, molecular signaling networks from the external environment also play a crucial role in modulating transcriptional activity and maintaining stem cell homeostasis [45]. In mouse ESCs, the Leukemia Inhibitory Factor/Signal Transducer and Activator of Transcription 3 (LIF/STAT3) pathway, and in human ESCs, the Fibroblast Growth Factor/Extracellular Signal-Regulated Kinase (FGF/ERK) pathway, interact with nuclear transcription factors to balance self-renewal and differentiation readiness [46]. The Wnt/β-catenin pathway supports proliferation and maintains stem cell identity by enhancing the expression of pluripotency genes, while the Notch and Hedgehog pathways play a role in maintaining the stem cell niche and modulating responses to environmental signals [47].
The interactions between transcription factors and signaling pathways are not linear, but rather form a dynamic and redundant network [48]. For example, activation of β-catenin through the Wnt pathway can increase the expression of OCT4 and NANOG, while the Notch pathway helps stabilize SOX2 expression in the quiescent state [49]. Epigenetic mechanisms are also integrated into this network; histone modifications and DNA methylation facilitate the access of transcription factors to chromatin and maintain the gene expression patterns characteristic of stem cells [50].
Coordination between transcription factors and signaling networks is crucial for stem cells to remain pluripotent and respond to physiological needs [3]. Disruption of these networks, whether due to genetic mutations or dysregulated signals from the microenvironment, can result in loss of stem cell identity, premature differentiation, or neoplastic transformation [51].
Table 1 Transcription factors and signaling networks in the maintenance of stem cell identity.
Component |
Main function |
Related signal paths |
Interaction mechanism |
OCT4 |
Maintaining pluripotent status and self-renewal |
Wnt/β-catenin and LIF/STAT3 |
Activates self-renewal genes and suppresses premature differentiation pathways and expression can be enhanced by β-catenin |
SOX2 |
Supports pluripotency and expression of critical target genes |
Notch |
Works synergistically with OCT4 and stabilization of expression by Notch in quiescent conditions |
NANOG |
Strengthening the stability of the transcription network |
Wnt/β-catenin |
Supports resistance to external differentiation signals and expression can be enhanced by β-catenin |
LIF/STAT3 |
Maintain self-renewal on mouse ESC |
Core transcription factors |
Interacts with OCT4/SOX2/NANOG to balance proliferation and differentiation readiness |
FGF/ERK |
Maintaining self-renewal in human ESC |
Core transcription factors |
Modulating OCT4, SOX2, and NANOG activity for self-renewal-differentiation balance |
Wnt/β-catenin |
Supports stem cell proliferation and identity |
OCT4 and NANOG |
β-catenin activation increases OCT4 and NANOG expression |
Notch |
Maintaining niche and cell response to the environment |
SOX2 |
Stabilizing SOX2 expression under quiescent conditions |
Hedgehog |
Modulating cell response to niche |
Core transcription factors |
Supports homeostasis and readiness for stem cell differentiation |
Epigenetics (DNA methylation and histone modifications) |
Facilitates access of transcription factors to chromatin |
OCT4, SOX2, and NANOG |
Ensures stem cell-specific gene expression profiles; supports transcription and signaling networks |
Epigenetic mechanisms that influence stem cell stability
Stem cell stability and the ability to maintain their identity are largely determined by epigenetic regulation, which regulates gene expression without altering the DNA sequence [52]. Key epigenetic mechanisms include histone modification, DNA methylation, and regulation by non-coding ribonucleic acid (RNA), which work synergistically to maintain self-renewal and pluripotency [53].
Histone modifications play a crucial role in regulating chromatin accessibility and gene transcriptional status [54]. For example, H3 lysine 4 methylation (H3K4me3) is associated with active genes that support pluripotency, while H3 lysine 27 trimethylation (H3K27me3) marks repressed differentiation genes, forming bivalent domains [55]. This bivalent configuration allows stem cells to remain flexible, ready to differentiate rapidly upon receiving specific signals, while maintaining their basic cellular identity [56].
DNA methylation in gene promoter regions is also crucial for stabilizing stem cell phenotypes [57]. Genes that support pluripotency generally remain hypomethylated, maintaining their expression, while genes that promote differentiation remain hypermethylated to prevent premature activation [58]. These methylation dynamics are regulated by DNA methyltransferase (DNMT) enzymes and Ten-Eleven Translocation (TET) dioxygenases, which interact with nuclear transcription factors such as OCT4, SOX2, and NANOG, thus maintaining a balance between self-renewal and differentiation readiness [59].
In addition, microRNAs and long non-coding RNAs (lncRNAs) play a role in maintaining stem cell stability by targeting specific genes involved in the cell cycle, apoptosis, and differentiation pathways [60]. For example, microRNAs from the microRNA (miR)-290/295 family in mouse ESCs or miR-302 in human ESCs suppress the expression of pro-differentiation factors while supporting proliferation [61]. Some lncRNAs also function as scaffolds to recruit chromatin remodeling complexes, thereby enhancing the expression of pluripotency genes and suppressing differentiation genes [62].
The integration of these various epigenetic mechanisms creates a dynamic chromatin landscape, allowing stem cells to maintain their identity while remaining responsive to external signals [63]. Epigenetic disruption, whether due to mutations in chromatin-modifying enzymes or exposure to damaging environments, can lead to loss of pluripotency, premature differentiation, or neoplastic transformation [64]. Stem cell identity and physiological homeostasis are maintained through the coordinated regulation of self-renewal capacity, cell cycle control, pluripotency transcription factors, epigenetic mechanisms, and key signaling pathways, as illustrated in Figure 1.
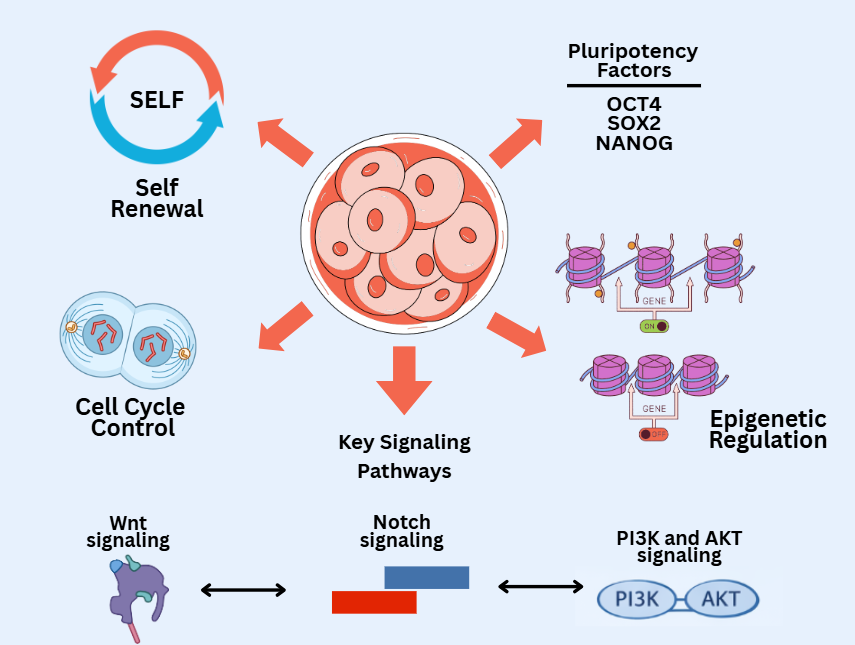
Figure 1 Regulation of stem cell identity and physiology.
Molecular mechanisms of cellular differentiation
Cell differentiation is a coordinated process regulated by key molecular signaling pathways, specific transcriptional regulation, microRNAs, and epigenetic dynamics, which together determine cell fate and its interaction with the microenvironmental niche.
Main signal path
Cell differentiation is controlled by evolutionarily conserved molecular signaling pathways that play a role in cell fate determination, proliferation, and interactions with the microenvironment [65]. Pathways such as Wnt, Notch, Hedgehog, and Bone Morphogenetic Protein/Transforming Growth Factor-beta (BMP/TGF-β) form complex regulatory networks, enabling the phenotypic transition of stem cells into specific cells in a coordinated and timely manner [9]. Table 2 summarizes the major molecular signaling pathways that regulate stem cell differentiation and maintenance.
The Wnt pathway acts as a key regulator of pluripotency and early differentiation [10]. Wnt activation results in the accumulation of β-catenin in the cytoplasm and its translocation to the nucleus, where it interacts with T-cell factor/Lymphoid enhancer-binding factor (TCF/LEF) transcription factors to activate the expression of genes supporting pluripotency [47]. This pathway also plays a role in determining the mesoderm and endoderm lineages in embryonic stem cells and balancing proliferation and differentiation in adult tissues [66]. Dysregulation of the Wnt pathway can lead to excessive proliferation or failure of differentiation, potentially leading to tumorigenesis [67].
Notch is an intercellular communication pathway that relies on ligand-membrane receptor interactions [68]. Notch activation leads to proteolytic cleavage of the receptor, generating intracellular fragments (NICDs) that enter the nucleus to modulate target gene expression [69]. This pathway plays a crucial role in maintaining stem cells in a quiescent state, determining the direction of progenitor cell differentiation, and regulating tissue proliferation [70]. Notch effects are contextual, strongly influenced by cell type, signal strength, and interactions with other pathways such as Wnt and Hedgehog [71].
The Hedgehog pathway controls cell proliferation, tissue patterning, and specialization [72]. Activation of this pathway through the ligand Sonic Hedgehog (Shh) inactivates the protein Patched, allowing Smoothened to activate the transcription factor Glioma-associated oncogene (Gli) [73]. Gli then regulates gene expression that supports stem cell proliferation and specific differentiation [74]. The Hedgehog pathway plays a crucial role in organ formation during embryogenesis and in maintaining homeostasis of adult tissues, including the nervous and hematopoietic systems [75].
Bone Morphogenetic Protein (BMP) and Transforming Growth Factor-β (TGF-β) are cytokine ligands that regulate differentiation through activation of serine/threonine kinase receptors and phosphorylation of Smads [76]. These pathways determine mesodermal and endodermal cell lineage specialization and interact with the Wnt and Notch pathways to direct differentiation spatiotemporally [77]. BMPs tend to promote mesenchymal cell differentiation, while TGF-β can induce epithelial differentiation while suppressing excessive proliferation, thus playing a crucial role in maintaining tissue homeostasis [78].
Coordination between the Wnt, Notch, Hedgehog, and BMP/TGF-β pathways forms a dynamic signaling network, enabling stem cells to respond to both internal and external stimuli [9]. Cross-reactions between these pathways ensure directed, stable, and physiologically appropriate differentiation, while maintaining a stem cell population ready for tissue regeneration [79].
Table 2 Key signaling pathways in stem cell differentiation and maintenance.
Signal path |
Activation mechanism |
Main function |
Interaction/Coordination |
Wnt/β-catenin |
Wnt activation → accumulation of β-catenin → translocation to the nucleus → interaction with TCF/LEF |
Maintaining pluripotency, specifying mesoderm and endoderm lineages, and regulating proliferation vs differentiation |
Interacting with Notch, BMP/TGF-β and dysregulation can lead to excessive proliferation or failure of differentiation |
Notch |
Ligand-membrane receptor → receptor cleavage → NICD enters nucleus → target gene modulation |
Maintains stem cell quiescence, determines the direction of progenitor cell differentiation, and regulates tissue proliferation |
Contextual effects and interactions with Wnt and Hedgehog |
Hedgehog (Hh) |
Sonic Hedgehog (Shh) ligand inactivates Patched → Smoothened activates → Gli regulates gene expression |
Regulates proliferation, tissue patterning, and cell specialization and is important during embryogenesis and adult tissue homeostasis |
Coordination with Notch and Wnt for organ formation and homeostasis |
BMP/TGF-β |
Ligand activates serine/threonine kinase receptors → Smad phosphorylation → modulation of gene expression |
Determines mesoderm and endoderm specialization, promotes mesenchymal (BMP) or epithelial (TGF-β) differentiation, and regulates tissue homeostasis |
Interacts with Wnt and Notch to direct spatiotemporal differentiation |
Specific transcriptional regulation and microRNAs
Stem cell differentiation relies heavily on specific transcriptional regulation, where transcription factors regulate the expression of genes that determine cell fate [80]. Factors such as GATA-binding transcription factor (GATA), PU box-binding protein 1 (PU.1), Runt-related transcription factor (RUNX), and Paired box (PAX) direct differentiation pathways toward specific lineages, such as hematopoietic, mesenchymal, or neural [81]. The interactions between these transcription factors form a coordinated regulatory network, enabling directed cellular transitions while maintaining phenotypic integrity and stability [82]. For example, the combination of GATA1 and PU.1 determines the specialization of hematopoietic progenitor cells toward erythroid or myeloid, while RUNX2 promotes osteogenic differentiation in mesenchymal stem cells [83].
In addition to transcriptional regulation, miRNAs act as post-transcriptional regulators, stabilizing differentiation pathways and preventing the expression of genes inconsistent with the desired cell fate [84]. miRNAs target specific mRNAs for degradation or translational inhibition, thereby reinforcing differentiation decisions driven by transcription factors [85]. For example, miR-290/295 in mouse ESCs and miR-302/367 in human ESCs repress pro-differentiation genes while promoting self-renewal, while miR-124 and miR-9 direct neural progenitors toward a mature neuronal phenotype [61]. miRNAs also serve as buffers against fluctuations in extrinsic signals, maintaining consistent gene expression during differentiation and enabling adaptive responses to microenvironmental conditions [86].
The interaction between specific transcription factors and microRNAs forms a multilevel regulatory network that ensures proper cellular differentiation [85]. Transcription factors can regulate miRNA expression, while miRNAs can modulate the levels of specific transcription factors, creating a feedback loop that reinforces the direction of differentiation [87]. This dynamic allows stem cells to balance self-renewal and specialization, ensuring that differentiation occurs only when both internal and external signals are adequate [88].
Dysregulation of transcription factors or microRNAs can trigger premature differentiation, loss of stem cell identity, or neoplastic transformation, emphasizing the importance of maintaining the integrity of these regulatory networks [89].
Epigenetic dynamics during differentiation
The process of cell differentiation in stem cells involves complex and coordinated epigenetic changes, enabling cells to adopt specific phenotypes while maintaining genetic integrity [90]. These epigenetic dynamics include histone modifications, DNA methylation and demethylation, and regulation by non-coding RNA, which collectively orchestrate specific gene expression programs for specific cell lineages [91].
During differentiation, histone modifications play a role in opening or closing chromatin to allow transcription factors to access it [92]. For example, trimethylation of H3 lysin 27 (H3K27me3) marks repressed differentiation genes in stem cells, while acetylation of H3 lysin 27 (H3K27ac) activates specific target genes as cells begin to differentiate [93]. Bivalent domains, which are combinations of H3K4me3 and H3K27me3 at specific genes, allow cells to remain flexible, allowing differentiation genes to be activated immediately upon receiving specific signals [56].
DNA methylation also exhibits significant dynamics [94]. Genes that support pluripotency tend to be hypomethylated to maintain active expression, while differentiation genes remain hypermethylated until the differentiation pathway is activated [95]. DNMT enzymes and TET dioxygenases play a role in regulating this methylation, enabling the proper transition between self-renewal and specialization states [59].
Additionally, non-coding RNAs, including microRNAs and lncRNAs, play a crucial role in epigenetic modulation during differentiation [96]. MicroRNAs target pro-differentiation factor mRNAs for degradation or translational inhibition, thereby facilitating the linear transition from stem cell phenotype to programmed cell phenotype [97]. LncRNAs can serve as scaffolds to recruit chromatin remodeling complexes or histone modification enzymes, which modulate chromatin status at key differentiation genes [98].
These epigenetic dynamics are flexible yet controlled, allowing stem cells to respond adaptively to both internal and external signals [99]. Dysregulation of these mechanisms, whether due to mutations in epigenetic enzymes or exposure to environmental stress, can lead to premature differentiation, loss of cell identity, or neoplastic transformation [100].
Interaction with micro-environmental niches
Stem cells do not function autonomously but are constantly influenced by their microenvironmental niche—a cellular and molecular network that regulates stem cell behavior, proliferation, and differentiation [101]. This niche consists of stromal cells, extracellular matrix (ECM), growth factors, cytokines, and mechanical and metabolic components that form a dynamic and adaptive microecosystem [102]. These interactions ensure a balance between self-renewal and differentiation in the stem cell pool according to the physiological needs of the tissue [103].
Communication between stem cells and their niche occurs through several mechanisms. First, paracrine and autocrine signals, such as Leukemia Inhibitory Factor (LIF), Wnt, Notch ligands, and BMP/TGF-β, convey molecular instructions that determine cell fate [104]. These pathways not only influence proliferation and differentiation but also maintain the quiescent state of stem cells during homeostasis [105]. Second, physical interactions with the ECM and cell adhesion via integrins or cadherins transmit mechanotransduction signals, which modulate gene expression, transcription factor activity, and cytoskeleton reorganization [106].
Furthermore, the microenvironmental niche also regulates the availability of metabolites and oxygen, which influence the metabolic profile of stem cells [107]. Hypoxic conditions in certain niches, such as the bone marrow, increase the expression of Hypoxia Inducible Factor 1 Alpha (HIF-1α), which supports self-renewal while preventing premature differentiation [108]. Niches also provide protective mechanisms, including anti-apoptotic factors and antioxidant enzymes, to maintain genome integrity and prevent the accumulation of mutations [109].
These interactions are dynamic and context-dependent; changes in the niche due to injury, inflammation, or aging can trigger stem cell activation, modulate differentiation pathways, or even dysregulate proliferation [110]. Disruption of the relationship between stem cells and their niche can result in loss of tissue homeostasis, decreased regenerative capacity, or neoplastic transformation [111]. The molecular mechanisms of stem cell differentiation are illustrated in Figure 2, showing how major signaling pathways converge on transcriptional regulation and lineage commitment.
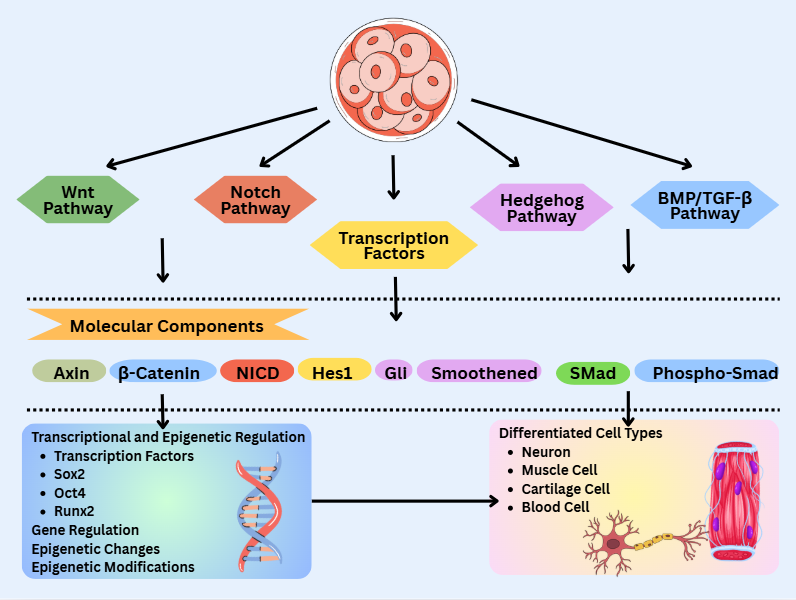
Figure 2 Stem cell differentiation pathways.
Interaction between stem cell physiology and differentiation
The interplay between stem cell physiology and cellular differentiation exhibits a dynamic balance between self-renewal, proliferation, and specialization, which is regulated by molecular signaling pathways, transcriptional regulation, and microenvironmental factors, and can be studied using both in vitro and in vivo experimental models.
How stem cell physiology determines differentiation capacity
Stem cell physiology directly influences differentiation capacity through a complex integration of cell cycle regulation, molecular signaling pathways, transcription factors, and epigenetic mechanisms [112]. Effective self-renewal, optimal metabolic conditions, and interactions with the microenvironmental niche form the basis of stem cellsʼ ability to differentiate appropriately [113].
First, cell cycle status influences differentiation readiness. Stem cells with a longer G1 phase are more ready to respond to differentiation signals, while cells undergoing rapid proliferation tend to maintain self-renewal [32]. Checkpoint regulation, CDK/cyclin activity, and pathways such as PI3K/AKT and MAPK/ERK balance proliferation and differentiation readiness, allowing cellular transitions to occur in a controlled manner [33].
Second, the metabolic profile of stem cells plays a crucial role. Pluripotent stem cells generally rely on glycolytic metabolism, supporting rapid proliferation while maintaining pluripotency [114]. As differentiation begins, a shift toward oxidative metabolism occurs, providing energy and metabolites essential for the development of specific cell phenotypes [115]. Thus, the physiological status of stem cells determines their ability to effectively respond to differentiation signals [38].
Third, nuclear transcription factors such as OCT4, SOX2, and NANOG, along with epigenetic regulation (histone modification, DNA methylation, and microRNA), form the genetic and chromatin basis for differentiation [116]. Transcription factors maintain the expression of pluripotency genes while suppressing premature differentiation pathways [117]. When cells receive specific signals, epigenetic changes unlock differentiation genes, allowing stem cells to execute a linear program toward their targeted phenotype [118].
Fourth, interactions with the microenvironmental niche provide an external context that directs differentiation pathways [119]. Paracrine factors, ECM adhesion, mechanotransduction, and hypoxic conditions can modulate the activity of the Wnt, Notch, Hedgehog, or BMP/TGF-β pathways, allowing stem cells to adjust proliferation and differentiation according to the physiological needs of the tissue [51].
The relationship between self-renewal, proliferation, and specialization
Stem cells maintain tissue homeostasis through a complex coordination of self-renewal, proliferation, and differentiation [120]. Self-renewal allows stem cells to maintain a population with pluripotent or multipotent identities, while proliferation ensures sufficient cell numbers for tissue regeneration [121]. Differentiation allows stem cells to produce phenotypically programmed progeny, forming functional tissues [122]. These three aspects are dynamically interconnected, allowing cellular differentiation to proceed without depleting the stem cell reservoir [123].
Self-renewal and proliferation are controlled by intrinsic and extrinsic signaling pathways, including Wnt/β-catenin, Notch, Hedgehog, and PI3K/AKT [124]. These pathways balance symmetric division, which increases the number of stem cells, with asymmetric division, which produces one stem cell and one progenitor cell for differentiation [125]. Symmetric division supports the expansion of the stem cell population during growth or regeneration, while asymmetric division allows integration between self-renewal and specialization [126]. Cell cycle checkpoint mechanisms and epigenetic control ensure proliferation occurs without loss of cell identity or the introduction of potentially pathological genetic mutations [127].
Cellular specialization relies on the integration of signals from specific transcription pathways and epigenetic mechanisms [128]. Core transcription factors such as OCT4, SOX2, and NANOG maintain the expression of pluripotency genes, while activation of lineage-specific factors such as GATA, RUNX, or PAX drives stem cells to transition to a specific phenotype [129]. Epigenetic dynamics, including histone modifications and DNA methylation, stabilize differentiation decisions, so that progenitor cell proliferation remains aligned with the desired specialization [91].
The interplay between self-renewal, proliferation, and specialization allows stem cells to adaptively respond to the physiological needs of tissues [130]. For example, tissue injury triggers stem cell proliferation through the activation of signaling pathways, followed by specific differentiation to replace lost cells while maintaining the self-renewing reservoir [131]. Conversely, dysregulation of this relationship can lead to loss of the stem cell reservoir, excessive proliferation, or premature differentiation, potentially leading to regeneration failure or tumorigenesis [132].
Examples of in vitro and in vivo experimental models
A deep understanding of stem cell physiology and cellular differentiation mechanisms has largely been achieved through in vitro and in vivo experimental models, which allow for controlled observation of molecular and phenotypic processes [133]. These models have become essential tools for elucidating signaling pathways, transcriptional regulation, epigenetic interactions, and the therapeutic potential of stem cells [1]. Table 3 summarizes the in vitro and in vivo experimental models used to study stem cell physiology and differentiation mechanisms.
In vitro models allow precise manipulation of the cell environment and direct observation of proliferation, self-renewal, and differentiation [134]. ESC culture is a classic model, in which mouse or human ESCs are maintained in media containing growth factors such as LIF or basic Fibroblast Growth Factor (bFGF) to maintain pluripotency [135]. Differentiation is directed using a combination of extrinsic factors, such as BMP4 for the mesodermal pathway or retinoic acid for the neuroectodermal pathway [136].
iPSCs are also an important model for studying linear differentiation in human cells, allowing the evaluation of tissue-specific phenotypes without the need for embryonic material [137]. 2D monolayer culture protocols or 3D organoids are used to mimic the native tissue environment, including brain, liver, pancreas, and heart organoids, which replicate intercellular interactions, polarity, and three-dimensional differentiation [138].
Co-culture systems and biomaterial scaffolds are also used to mimic microenvironmental niches [139]. For example, co-culture of MSCs with endothelial cells or osteoblasts allows studies of paracrine signaling and intercellular interactions in the regulation of self-renewal and differentiation [140]. Synthetic polymer matrices or hydrogels support proliferation and linear differentiation by providing mechanical and adhesive signals that mimic native tissue [141].
Reporter genes and live-cell imaging are often used to monitor the expression of transcription factors or signaling pathways in real time [142]. For example, using Green Fluorescent Protein (GFP) reporters for OCT4 or SOX2 in ESCs allows observation of pluripotency status when the Wnt or Notch pathways are manipulated [143].
Meanwhile, in vivo models provide a complete physiological context for evaluating cell integration, specialization, and tissue function [144]. Mouse chimeras and transplantation are classic models, in which ESCs or iPSCs are introduced into embryonic or adult tissues to assess the contribution of stem cells to organ formation and tissue homeostasis [145]. Another example is the xenotransplantation of human stem cells into immunodeficiency models (e.g., NOD/SCID mice) to evaluate linear differentiation and therapeutic potential [146].
In vivo lineage tracing using Cyclization recombinase-Locus of crossover in P1 (Cre-LoxP) or fluorescent reporter systems allows tracking the origin and fate of progenitor cells in complex tissues, as well as identifying cell populations involved in regeneration or pathology [147]. Furthermore, injury-induced regeneration models, such as those involving the heart, brain, or bone marrow, allow evaluation of how stem cells proliferate, differentiate, and interact with their microenvironmental niches in both physiological and pathological contexts [148].
Combined in vitro and in vivo approaches are also being applied, for example by transplanting organoids into experimental animals to assess their integration and functional capabilities, or by pre-conditioning stem cells in vitro before use in in vivo regenerative therapies [149]. As illustrated in Figure 3, stem cell fate is dynamically regulated by reciprocal interactions with the surrounding microenvironmental niche, including extracellular matrix components, stromal cells, vascular-derived signals, oxygen gradients, and biochemical as well as mechanical cues.
Table 3 Examples of in vitro and in vivo experimental models for stem cell studies.
Experimental model |
Stem cell types |
Purpose/Application |
Key methods/Techniques |
Reference |
ESC culture |
Embryonic Stem Cell (mouse/human) |
Maintaining pluripotency and studying linear differentiation |
Media with growth factors (LIF, bFGF) and directed differentiation with BMP4 and retinoic acid |
[134-136] |
iPSC culture |
Induced pluripotent stem cell |
Assessing specific differentiation of human tissues without the use of embryos |
2D monolayer or 3D organoid culture (brain, liver, pancreas, and heart) |
[137,138] |
Co-culture System |
MSC, ESC, and iPSC |
Understanding paracrine interactions, self-renewal, and differentiation |
Co-culture with endothelial cells or osteoblasts and scaffold biomaterials and hydrogels |
[139-141] |
Reporter gene and live-cell imaging |
ESC and iPSC |
Tracking the expression of transcription factors or signaling pathways in real-time |
GFP reporter for OCT4/SOX2 and monitoring of the Wnt or Notch pathway |
[142,143] |
Chimera / Transplant |
ESC and iPSC |
Evaluation of cell integration and contribution to organ formation and homeostasis |
Transplantation into embryos or adult tissues and xenotransplantation into NOD/SCID mice |
[144-146] |
Lineage tracing in vivo |
ESC, iPSC, and progenitor cells |
Tracing the origin and fate of cells and identifying regenerative populations |
Cre-LoxP or fluorescent reporter system |
[147] |
Injury-induced regeneration model |
ESC, iPSC, and MSC |
Assessing proliferation, differentiation, and interaction with the post-injury niche |
Heart, brain, and bone marrow injuries and evaluation of stem cell responses |
[148] |
Organoid transplantation |
iPSC-derived organoid |
Assess network integration and functionality |
In vitro pre-conditioning followed by in vivo transplantation |
[149] |
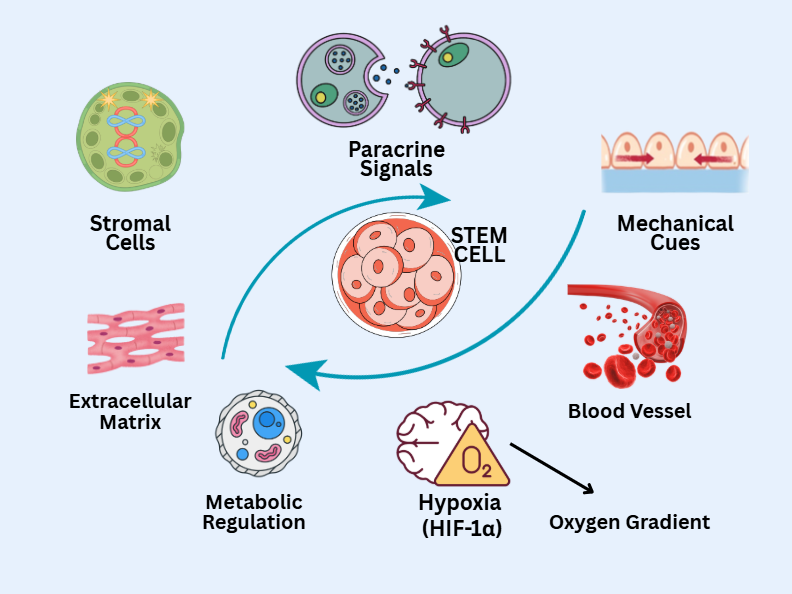
Figure 3 Stem cell–niche interactions.
Therapeutic potential and clinical applications
Stem cells have great therapeutic potential in tissue regeneration and the treatment of degenerative diseases and cancer, with strategies that include regenerative therapy of vital organs, microenvironmental modulation, and the use of programmed cells such as iPSCs, ESCs, and MSCs.
Regenerative therapy
Stem cell-based regenerative therapies aim to restore damaged tissue function through cell integration, linear differentiation, and stimulation of endogenous tissue repair [131]. Both pluripotent and multipotent stem cells provide promising platforms for therapies in the heart, nervous system, and liver, each with specific challenges and approaches [150]. Table 4 summarizes the applications of regenerative therapies using stem cells in various organs and body systems.
Myocardial injury from cardiac infarction causes massive loss of cardiomyocytes, which are difficult to regenerate effectively [151]. Stem cells, including human iPSC-derived cardiomyocytes and MSCs, are used to replace lost cells and restore cardiac contractile function [152]. Preclinical studies in mouse and pig models have shown that stem cell transplantation can increase ventricular ejection fraction and reduce fibrogenesis through paracrine mechanisms, including the secretion of angiogenic factors such as VEGF and FGF2, as well as anti-inflammatory effects [153]. Furthermore, in vitro cardiac organoids are used to evaluate electrical integration and cell contractility prior to in vivo transplantation [154].
Central nervous system injuries, such as spinal cord injury or neurodegenerative diseases, present significant challenges due to the limited regeneration capacity of adult neurons [155]. Neural progenitor cells (NPCs) and iPSC-derived neurons are used to replace lost neurons while supporting remyelination by oligodendrocytes [156]. In mouse models of spinal cord injury, NPC transplantation has been shown to enhance axon regeneration, reduce gliosis, and improve motor function [157]. Therapeutic mechanisms include direct differentiation, modulation of the inflammatory microenvironment, and secretion of neurotrophic factors such as Brain-Derived Neurotrophic Factor (BDNF) and Glial Cell Line-Derived Neurotrophic Factor (GDNF) [158]. Furthermore, 3D brain organoids are used to model neurodegenerative diseases and evaluate the potential for neuronal differentiation and tissue integration [159].
The liver has natural regenerative abilities, but extensive damage from cirrhosis or chronic hepatitis often requires additional intervention [160]. Stem cells, including hepatic progenitor cells, iPSC-derived hepatocyte-like cells, and MSCs, are used to improve hepatocyte function while modulating the inflammatory microenvironment [161]. In a mouse model of chemical liver injury, stem cell transplantation demonstrated restoration of liver function, increased expression of metabolic enzymes, and reduced fibrosis through directed differentiation and secretion of anti-fibrotic factors [162]. Furthermore, liver organoids are used to assess metabolic function and toxicology prior to clinical application [163].
The success of regenerative therapy is determined not only by cell transplantation alone, but also by the stem cellsʼ ability to respond to local signaling pathways such as Wnt, Notch, and BMP/TGF-β, as well as interact with the microenvironmental niche [164]. Factors such as oxygen levels, cell growth, and the extracellular matrix influence cell proliferation, differentiation, and integration, resulting in coordinated tissue repair [131]. In vitro preconditioning of stem cells using specific factors or biomaterial scaffolds can improve cell viability and integration after in vivo transplantation [165].
Table 4 Examples of regenerative therapy using stem cells in specific organs and systems.
Organ/System |
Stem cell types |
Goals of therapy |
Mechanism/Strategy |
Experimental model |
Reference |
Heart |
Human iPSC-derived cardiomyocytes and MSCs |
Replacing lost cardiomyocyte cells and improving contractile function |
Cell integration, linear differentiation, secretion of paracrine factors (VEGF and FGF2), and modulation of fibrogenesis |
Mouse and pig models and in vitro cardiac organoids |
[151-154] |
Central nervous system |
Neural progenitor cells (NPC) and iPSC-derived neurons |
Replacing lost neurons, supporting oligodendrocyte remyelination, and improving motor function |
Direct differentiation, modulation of the inflammatory microenvironment, and secretion of neurotrophic factors (BDNF and GDNF) |
Spinal cord injury mouse model and 3D brain organoid |
[155-159] |
Liver |
Hepatic progenitor cells, iPSC-derived hepatocyte-like cells, and MSC |
Improve hepatocyte function, reduce fibrosis, and modulate inflammation |
Direct differentiation, secretion of anti-fibrotic factors, and increased expression of metabolic enzymes |
Chemical liver injury mouse model and in vitro liver organoids |
[160-163] |
Implications in the treatment of degenerative diseases and cancer
Stem cells have significant therapeutic potential in the treatment of degenerative diseases and cancer, thanks to their unique ability to self-renewal, linear differentiation, and modulate the tissue microenvironment [166]. Stem cell-based therapies aim to repair damaged tissues, regenerate specific cells, and normalize disrupted molecular pathways, opening up opportunities for precision and regenerative medical approaches [167].
In degenerative diseases, including neurodegenerative, cardiovascular, and musculoskeletal disorders, tissue damage is often irreversible or has limited natural regenerative capacity [168]. NPCs and iPSC-derived neurons are used in Parkinsonʼs and Alzheimerʼs models to replace lost neurons, promote remyelination by oligodendrocytes, and increase the secretion of neurotrophic factors such as BDNF and GDNF [169]. Preclinical studies in mice have shown improvements in motor and cognitive function, confirming that the therapeutic mechanism involves direct differentiation and modulation of the inflammatory microenvironment [170].
In cardiovascular therapy, iPSC-derived cardiomyocytes and MSCs are used to replace cardiomyocytes lost due to myocardial infarction [171]. In addition to differentiating into contractile cells, stem cells also stimulate angiogenesis through the secretion of VEGF and FGF2, and suppress the local inflammatory response, thus improving cardiac function recovery [172]. In vitro approaches, such as cardiac organoids or 3D cardiac tissue culture, allow for evaluation of electrophysiological integration before transplantation, improving the safety and effectiveness of therapy [173].
In musculoskeletal tissue regeneration, MSCs and bone progenitors play a role in osteogenic differentiation and the secretion of anti-inflammatory factors, which help reduce bone and cartilage degradation and accelerate tissue recovery [174]. The use of 3D scaffolds and co-culture with stromal cells allows cellular interactions that mimic the native niche, thereby increasing differentiation efficiency [175].
In cancer therapy, stem cells play a dual role, serving both as therapeutic targets and as research models [176]. The study of cancer stem cells (CSCs) helps explain tumor heterogeneity, resistance to chemotherapy, and metastatic potential [177]. CSCs maintain an aggressive tumor cell population through self-renewal and limited differentiation, making conventional therapies often incapable of eradicating all tumor cells [178].
Stem cell-based therapies offer strategies for targeting CSCs specifically or as therapeutic vectors [179]. For example, MSCs modified to express antitumor factors or prodrug systems can influence the tumor microenvironment, inhibit CSC proliferation, and minimize damage to normal tissue [180]. Furthermore, cancer organoid models and iPSC-derived tumor models enable mapping of resistance pathways, drug screening, and the development of precision therapies tailored to a patientʼs molecular profile [181].
The use of stem cells in degenerative diseases and cancer emphasizes the need for controlled differentiation, cell integration, and clinical safety [182]. Precisely modified cells are capable of balancing self-renewal, specialization, and niche interaction, thus supporting tissue regeneration or effective tumor cell elimination [51]. This approach opens up opportunities for the development of regenerative therapies and precision oncology, which combine molecular strategies, bioengineering, and cellular transplantation [183].
Cell integration, safety, risk of tumorigenesis
Stem cell-based therapies have significant regenerative potential, but their implementation faces significant challenges related to cell integration, regulation of differentiation, and the risk of tumorigenesis [184]. The success of these approaches depends heavily on the cellsʼ ability to interact functionally with target tissues, ensuring patient safety, and preventing uncontrolled cell proliferation [13].
Effective cell integration is essential for tissue function recovery [185]. Transplanted stem cells must be able to migrate to the injured area, interact with their microenvironmental niche, and establish physiological connections with surrounding cells [131]. For example, in cardiovascular therapy, transplanted cardiomyocytes must integrate electrophysiologically with native cardiomyocytes to ensure synchronous heart contractions [186]. Meanwhile, in the nervous system, transplanted neurons must form stable, functional synapses to maintain normal impulse transmission [187]. Disturbed microenvironmental conditions, such as those caused by inflammation or fibrosis, can inhibit cell migration and differentiation, therefore the use of biomaterial scaffolds and in vitro preconditioning are applied to enhance cell integration [139].
Pluripotent stem cells have a vast differentiation capacity, placing them at risk of remaining undifferentiated or undergoing inappropriate differentiation after transplantation [188]. Cells that retain pluripotency or differentiate uncontrollably can disrupt tissue homeostasis, reduce therapeutic effectiveness, and lead to clinical complications [189]. To minimize this risk, strategies such as in vitro pre-differentiation, the use of growth factors or specialized matrices, and the selection of differentiated cell populations prior to transplantation are employed [190]. Furthermore, monitoring differentiation markers and assessing genomic integrity are essential to ensure therapeutic safety [191].
The risk of tumor formation is a major challenge, especially when using pluripotent stem cells [192]. Cells that are not fully differentiated or have genetic mutations have the potential to form teratomas or neoplasms after transplantation [193]. This risk is further increased by long-term in vitro expansion and dysregulation of proliferation pathways such as Wnt, PI3K/AKT, and Myelocytomatosis oncogene (MYC) [194]. To minimize this risk, strategies such as strict selection of differentiated cells, the implementation of suicide gene systems, and long-term monitoring with molecular imaging and tumor biomarkers are employed [195]. This approach is crucial for detecting abnormal proliferation or tumorigenesis at an early stage [196].
Addressing these challenges requires a multidimensional approach that integrates an understanding of stem cell physiology, differentiation pathways, niche interactions, and epigenetic regulation [197]. Strategies such as in vitro preconditioning, the use of biomaterial scaffolds, regulation of signaling pathways, and the selection of pre-differentiated cell populations can support optimal cell integration, reduce uncontrolled proliferation, and enhance clinical safety [198]. The combined application of these strategies ensures that stem cell therapy effectively and safely balances self-renewal, differentiation, and restoration of tissue function [199].
Programmed cell differentiation-based therapy
Programmed stem cell-based therapies, such as iPSCs, ESCs, and MSCs, have emerged as innovative strategies for tissue regeneration and the treatment of degenerative diseases [200]. These cellsʼ abilities for self-renewal, directed differentiation, and microenvironmental modulation make them promising candidates for precision therapy and individualized treatments [1].
ESCs possess high pluripotency, allowing them to differentiate into all somatic cell types [137]. In therapeutic applications, ESCs are used to replace cells lost due to injury or degenerative diseases, such as cardiomyocytes in myocardial infarction, neurons in neurodegenerative disorders, and hepatocytes in chronic liver damage [201]. The ESC differentiation process is regulated by a combination of growth factors and molecular signals, including Bone Morphogenetic Protein (BMP), Wnt, and Fibroblast Growth Factor (FGF), which determine the cell’s lineage-specific pathways [202]. Major challenges in ESC therapy include the risk of tumorigenesis and immunological rejection, making in vitro pre-differentiation strategies and the use of biomaterial scaffolds crucial to ensure safe integration in vivo [203].
iPSCs are obtained by reprogramming somatic cells through the expression of nuclear transcription factors such as OCT4, SOX2, Kruppel-Like Factor 4 (KLF4), and cellular Myelocytomatosis oncogene (c-MYC), thus achieving pluripotency similar to ESCs [204]. The advantage of iPSCs lies in their ability to generate autologous cells, which reduces the risk of immune reactions. iPSCs are used in regenerative therapies for the heart, nervous system, and liver, as well as as disease models for drug screening and molecular mechanism research [131]. In vitro models, including 2D cultures, 3D organoids, and co-culture systems, allow for controlled differentiation, monitoring of lineage markers, and optimization of interactions with the microenvironmental niche before in vivo transplantation [205].
MSCs are multipotent cells capable of differentiating into osteoblasts, chondrocytes, and adipocytes, and possess significant immunomodulatory properties [206]. MSCs are widely used in regenerative therapies for bone, cartilage, heart, and inflamed tissues [207]. In addition to their linear differentiation capabilities, MSCs act through paracrine mechanisms by secreting growth factors, anti-inflammatory cytokines, and angiogenic molecules that support endogenous tissue regeneration [208]. The advantages of MSCs include ease of harvest from bone marrow, adipose tissue, or the placenta, and a relatively low risk of tumorigenesis compared to pluripotent cells [209].
Programmed cell therapy faces challenges related to differentiation control, integration into target tissues, and the potential for abnormal proliferation [12]. Mitigation strategies include in vitro pre-differentiation, the use of biomaterial scaffolds, regulation of specific signaling pathways, and monitoring marker expression and cell viability before transplantation [198]. The success of therapy depends heavily on optimal cell integration with host tissue and modulation of the microenvironment, allowing precise and safe restoration of tissue function [207].
Challenges and future perspectives
Although stem cell-based therapies hold significant potential for tissue regeneration and the treatment of degenerative diseases, their implementation still faces numerous scientific, technical, and clinical challenges [164]. Limited understanding of stem cell physiology and the limitations of current experimental models are major obstacles to realizing safe and effective therapies [210].
Key limitations include heterogeneity within stem cell populations, the inability to fully control differentiation consistently, and a limited understanding of stem cell interactions with the in vivo microenvironmental niche [211]. In vitro models often fail to fully mimic the physiological conditions of intact tissues, while in vivo models face challenges such as biological variability and limitations in scale and long-term monitoring [212]. Furthermore, the risk of tumorigenesis in pluripotent stem cells and the potential for immune reactions to transplanted cells remain critical issues requiring more effective mitigation strategies [213].
Technological advances present promising opportunities to overcome the limitations of stem cell therapy [19]. Gene editing techniques, such as Clustered Regularly Interspaced Short Palindromic Repeats-CRISPR-associated protein 9 (CRISPR-Cas9), enable targeted gene modification to correct mutations, enhance linear differentiation, or reduce the risk of tumorigenesis [214]. 3D organoid cultures enable the creation of more accurate miniature human tissues to assess cell function, differentiation, and interactions [138]. Furthermore, bioprinting and advanced biomaterial scaffolds provide platforms for constructing three-dimensional tissues that support more physiological integration of transplanted cells [215]. The combined application of these technologies enhances control over cell proliferation, differentiation, and integration with high precision [216].
The safety and effectiveness of stem cell therapy can be enhanced through a multidimensional approach [196]. This approach includes in vitro pre-differentiation, selection of differentiated cell populations, monitoring of genomic integrity and lineage marker expression, and the use of biomaterial scaffolds to support homing and integration with target tissues [12]. Long-term monitoring using molecular imaging and stem cell biomarkers allows early identification of abnormal proliferation or tumorigenic risk [217]. Furthermore, combination with growth factors, signaling pathway modulation, or cell-microenvironment therapy strategies enhances the ability of stem cells to adapt to the physiological needs of the tissue [218].
Kesimpulan
Stem cell physiology and cellular differentiation processes are controlled by complex interactions between molecular signaling pathways, transcription factors, epigenetic mechanisms, and the tissue microenvironment. A deeper understanding of these mechanisms opens up opportunities for designing safer and more effective regenerative therapy strategies, including for degenerative diseases, tissue injury, and cancer. Future research should focus on optimizing differentiation control, cell integration, reducing the risk of tumorigenesis, and utilizing technologies such as organoids, bioprinting, and gene editing to improve the precision and efficacy of clinical therapies.
Acknowledgements
The authors would like to express their sincere gratitude to the National Research and Innovation Agency (BRIN), Indonesia for providing research facilities and support throughout this study. We also extend our appreciation to Universitas Airlangga for academic guidance, collaborative opportunities, and institutional support that greatly contributed to the completion of this work.
Funding Statement
This work was supported by the National Research and Innovation Agency (BRIN) and the Indonesia Endowment Fund for Education (LPDP) through the RIIM Program – Competitive Wave 7 (Decree No. 61/II.7/HK/2024). The funding body had no role in the design of the study, the collection, analysis, or interpretation of data, or in writing the manuscript.
Declaration of Generative AI in Scientific Writing
The authors declare that no generative AI tools were used in the writing or preparation of this manuscript.
CRediT Author Statement
Aswin Rafif Khairullah: Conceptualization, Literature review, and Writing – original draft. Saifur Rehman: Literature review, Data curation, and Writing – original draft. Imam Mustofa: Literature review, Data curation, and Writing – original draft. Bima Putra Pratama: Reference management, Data curation, and Validation. Eny Martindah: Conceptualization, Supervision, Funding acquisition, and Writing – review and editing. Arif Nur Muhammad Ansori: Writing – review and editing, Validation, and Supervision. Mohammad Sukmanadi: Manuscript preparation, Critical review, and Validation. Ilma Fauziah Ma’ruf: Manuscript preparation, Critical review, and Validation. Analinasari Analinasari: Manuscript preparation, Critical review, and Validation. Riza Zainuddin Ahmad: Methodological review, Critical revision, and Validation. Sri Suryatmiati Prihandani: Manuscript preparation, Critical review, and Validation. Bantari Wisynu Kusuma Wardhani: Writing – review and editing, Supervision, and Validation. Siti Hamidatul Aliyah: Manuscript preparation, Critical review, and Validation. Wita Yulianti: Reference management, Resources, and Validation. Dea Anita Ariani Kurniasih: Writing – review and editing, Visualization, and Validation Fadhila Utari: Project administration, Supervision, Writing – review and editing, and Validation.
Conflict of Interest Statement
The authors declare that there are no commercial or financial relationships that could be construed as a potential conflict of interest in the conduct and publication of this work.
Data Availability Statement
All data generated or analyzed in this study are included in this published article. No additional datasets were generated or analyzed during the current study.
References
[1] IZ Sadiq, FS Abubakar, BS Katsayal, B Ibrahim, A Adamu, MA Usman, M Aliyu, MA Suleiman and A Muhammad. Stem cells in regenerative medicine: Unlocking therapeutic potential through stem cell therapy, 3D bioprinting, gene editing, and drug discovery. Biomedical Engineering Advances 2025; 9, 100172.
[2] SM Alnasser, AS Alrobian, MS Alfayez, OT Almutairi, SS Almutairi and TS Alkeraidees. Pharmacological modulation of stem cells signaling pathway for therapeutic applications. Stem Cell Research & Therapy 2025; 16, 327.
[3] G Liu, BT David, M Trawczynski and RG Fessler. Advances in pluripotent stem cells: History, mechanisms, technologies, and applications. Stem Cell Reviews and Reports 2020; 16(1), 3-32.
[4] JY Lee and SH Hong. Hematopoietic stem cells and their roles in tissue regeneration. International Journal of Stem Cells 2020; 13(1), 1-12.
[5] G Mannino, C Russo, G Maugeri, G Musumeci, N Vicario, D Tibullo, R Giuffrida, R Parenti and DL Furno. Adult stem cell niches for tissue homeostasis. Journal of Cellular Physiology 2022; 237(1), 239-257.
[6] O Hamadelseed and T Skutella. Stem cell-based therapeutic strategies for down syndrome and Alzheimerʼs disease. Stem Cell Research and Therapy 2025; 16, 420.
[7] Y Jin, S Li, Q Yu, T Chen and D Liu. Application of stem cells in regeneration medicine. MedComm 2023; 4(4), 291.
[8] L Tan, X Liu, H Dou and Y Hou. Characteristics and regulation of mesenchymal stem cell plasticity by the microenvironment — specific factors involved in the regulation of MSC plasticity. Genes & Diseases 2022; 9(2), 296-309.
[9] M Pelullo, S Zema, F Nardozza, S Checquolo, I Screpanti and D Bellavia. Wnt, Notch, and TGF-β pathways impinge on hedgehog signaling complexity: An open window on cancer. Frontiers in Genetics 2019; 10, 711.
[10] C Xue, Q Chu, Q Shi, Y Zeng, J Lu and L Li. Wnt signaling pathways in biology and disease: Mechanisms and therapeutic advances. Signal Transduction and Targeted Therapy 2025; 10, 106.
[11] C Mani, PH Reddy and K Palle. DNA repair fidelity in stem cell maintenance, health, and disease. Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease 2020; 1866(4), 165444.
[12] BM Hussen, M Taheri, RK Yashooa, GH Abdullah, SR Abdullah, RK Kheder and SA Mustafa. Revolutionizing medicine: Recent developments and future prospects in stem-cell therapy. International Journal of Surgery 2024; 110(12), 8002-8024.
[13] HE Marei. Stem cell therapy: A revolutionary cure or a pandoraʼs box. Stem Cell Research & Therapy 2025; 16, 255.
[14] CW Zeng. Stem cell-based approaches for spinal cord injury: The promise of iPSCs. Biology 2025; 14(3), 314.
[15] B Yu, D Zhou, F Wang, X Chen, M Li and J Su. Organoids for tissue repair and regeneration. Materials Today Bio 2025; 33, 102013.
[16] A del Sol and S Jung. The importance of computational modeling in stem cell research. Trends in Biotechnology 2021; 39(2), 126-136.
[17] M Hofer and MP Lutolf. Engineering organoids. Nature Reviews Materials 2021; 6(5), 402-420.
[18] X Liu, Z Zhou, Y Zhang, H Zhong, X Cai and R Guan. Recent progress on the organoids: Techniques, advantages and applications. Biomedicine and Pharmacotherapy 2025; 185, 117942.
[19] W Zakrzewski, M Dobrzyński, M Szymonowicz and Z Rybak. Stem cells: Past, present, and future. Stem Cell Research & Therapy 2019; 10(1), 68.
[20] S Bolkent. Cellular and molecular mechanisms of asymmetric stem cell division in tissue homeostasis. Genes to Cells 2024; 29(12), 1099-1110.
[21] F Ju, MM Atyah, N Horstmann, S Gul, R Vago, CJ Bruns, Y Zhao, QZ Dong and N Ren. Characteristics of the cancer stem cell niche and therapeutic strategies. Stem Cell Research and Therapy 2022; 13(1), 233.
[22] A Zare, A Salehpour, A Khoradmehr, S Bakhshalizadeh, V Najafzadeh, S Almasi-Turk, M Mahdipour, R Shirazi and A Tamadon. Epigenetic modification factors and microRNAs network associated with differentiation of embryonic stem cells and induced pluripotent stem cells toward cardiomyocytes: A review. Life 2023; 13(2), 569.
[23] RB Narasimha, S Shreya, VA Jayabal, V Yadav, PK Rath, BP Mishra, S Kancharla, P Kolli, G Mandadapu, S Kumar, AK Mohanty and MK Jena. Stem cell therapy for diseases of livestock animals: An in-depth review. Veterinary Sciences 2025; 12(1), 67.
[24] R Cancedda and M Mastrogiacomo. The phoenix of stem cells: Pluripotent cells in adult tissues and peripheral blood. Frontiers in Bioengineering and Biotechnology 2024; 12, 1414156.
[25] K Thanaskody, AS Jusop, GJ Tye, WSWK Zaman, SA Dass and F Nordin. MSCs vs. iPSCs: Potential in therapeutic applications. Frontiers in Cell and Developmental Biology 2022; 10, 1005926.
[26] R Farahzadi, B Valipour, S Montazersaheb and E Fathi. Targeting the stem cell niche micro-environment as therapeutic strategies in aging. Frontiers in Cell and Developmental Biology 2023; 11, 1162136.
[27] TM Nguyen and M Aragona. Regulation of tissue architecture and stem cell dynamics to sustain homeostasis and repair in the skin epidermis. Seminars in Cell & Developmental Biology 2022; 130, 79-89.
[28] R He, Y Liu, W Fu, X He, S Liu, D Xiao and Y Tao. Mechanisms and cross-talk of regulated cell death and their epigenetic modifications in tumor progression. Molecular Cancer 2024; 23(1), 267.
[29] S Treichel and MD Filippi. Linking cell cycle to hematopoietic stem cell fate decisions. Frontiers in Cell and Developmental Biology 2023; 11, 1231735.
[30] B Evano, S Khalilian, GL Carrou, G Almouzni and S Tajbakhsh. Dynamics of asymmetric and symmetric divisions of muscle stem cells in vivo and on artificial niches. Cell Reports 2020; 30(10), 3195-3206.
[31] L Ding, J Cao, W Lin, H Chen, X Xiong, H Ao, M Yu, J Lin and Q Cui. The roles of cyclin-dependent kinases in cell-cycle progression and therapeutic strategies in human breast cancer. International Journal of Molecular Sciences 2020; 21(6), 1960.
[32] L Liu, W Michowski, A Kolodziejczyk and P Sicinski. The cell cycle in stem cell proliferation, pluripotency and differentiation. Nature Cell Biology 2019; 21(9), 1060-1067.
[33] I Pellarin, A DallʼAcqua, A Favero, I Segatto, V Rossi, N Crestan, J Karimbayli, B Belletti and G Baldassarre. Cyclin-dependent protein kinases and cell cycle regulation in biology and disease. Signal Transduction and Targeted Therapy 2025; 10, 11.
[34] J Shen, Q Wang, Y Mao, W Gao and S Duan. Targeting the p53 signaling pathway in cancers: Molecular mechanisms and clinical studies. MedComm 2023; 4(3), 288.
[35] E Fuchs and HM Blau. Tissue stem cells: Architects of their niches. Cell Stem Cell 2020; 27(4), 532-556.
[36] O Marescal and IM Cheeseman. Cellular mechanisms and regulation of quiescence. Developmental Cell 2020; 55(3), 259-271.
[37] CTJ van Velthoven and TA Rando. Stem cell quiescence: Dynamism, restraint, and cellular idling. Cell Stem Cell 2019; 24(2), 213-225.
[38] X Li, O Jiang and S Wang. Molecular mechanisms of cellular metabolic homeostasis in stem cells. International Journal of Oral Science 2023; 15, 52.
[39] P Saggese, A Sellitto, CA Martinez, G Giurato, G Nassa, F Rizzo, R Tarallo and C Scafoglio. Metabolic regulation of epigenetic modifications and cell differentiation in cancer. Cancers 2020; 12(12), 3788.
[40] X Huang, Y Wang, Q Li, X Li and C Wang. Coupling of stemness maintenance with cell cycle control in stem cells. Frontiers in Cell and Developmental Biology 2025; 13, 1693489.
[41] G Divisato, F Passaro, T Russo and S Parisi. The key role of microRNAs in self-renewal and differentiation of embryonic stem cells. International Journal of Molecular Sciences 2020; 21(17), 6285.
[42] CM MacCarthy, G Wu, V Malik, Y Menuchin-Lasowski, T Velychko, G Keshet, R Fan, I Bedzhov, GM Church, R Jauch, V Cojocaru, HR Schöler and S Velychko. Highly cooperative chimeric super-SOX induces naive pluripotency across species. Cell Stem Cell 2024; 31(1), 127-147.
[43] EJ Lee, R Sun and J Kim. The self-renewal function of Oct-4 can be replaced by the EWS-Oct-4 fusion protein in embryonic stem cells. Cellular and Molecular Life Sciences 2025; 82(1), 166.
[44] S Zhang, X Xiong and Y Sun. Functional characterization of SOX2 as an anticancer target. Signal Transduction and Targeted Therapy 2020; 5, 135.
[45] H He, M Yang, S Li, G Zhang, Z Ding, L Zhang, G Shi and Y Li. Mechanisms and biotechnological applications of transcription factors. Synthetic and Systems Biotechnology 2023; 8(4), 565-577.
[46] M Mossahebi-Mohammadi, M Quan, JS Zhang and X Li. FGF signaling pathway: A key regulator of stem cell pluripotency. Frontiers in Cell and Developmental Biology 2020; 8, 79.
[47] J Liu, Q Xiao, J Xiao, C Niu, Y Li, X Zhang, Z Zhou, G Shu and G Yin. Wnt/β-catenin signalling: Function, biological mechanisms, and therapeutic opportunities. Signal Transduction and Targeted Therapy 2022; 7(1), 3.
[48] L Martini, SH Baek, I Lo, BA Raby, EK Silverman, ST Weiss, K Glass and A Halu. Detecting and dissecting signaling crosstalk via the multilayer network integration of signaling and regulatory interactions. Nucleic Acids Research 2024; 52(1), 5.
[49] H Aria, M Azizi, S Nazem, B Mansoori, F Darbeheshti, A Niazmand, A Daraei and Y Mansoori. Competing endogenous RNAs regulatory crosstalk networks: The messages from the RNA world to signaling pathways directing cancer stem cell development. Heliyon 2024; 10(15), 35208.
[50] AJ Fritz, M El Dika, RH Toor, PD Rodriguez, SJ Foley, R Ullah, D Nie, B Banerjee, D Lohese, KM Tracy, KC Glass, S Frietze, PN Ghule, JL Heath, AN Imbalzano, A van Wijnen, J Gordon, JB Lian, JL Stein and GS Stein. Epigenetic-mediated regulation of gene expression for biological control and cancer: Cell and tissue structure, function, and phenotype. Results and Problems in Cell Differentiation 2022; 70, 339-373.
[51] H Lee, B Kim, J Park, S Park, G Yoo, S Yum, W Kang, JM Lee, H Youn and B Youn. Cancer stem cells: Landscape, challenges and emerging therapeutic innovations. Signal Transduction and Targeted Therapy 2025; 10, 248.
[52] A Harvey, G Caretti, V Moresi, A Renzini and S Adamo. Interplay between metabolites and the epigenome in regulating embryonic and adult stem cell potency and maintenance. Stem Cell Reports 2019; 13(4), 573-589.
[53] S Bakrim, NE Omari, OME Yaagoubi, A Khalid, AN Abdalla, SMA Hamza, SE Ibrahim, F Atifi, Y Zaid, A Bouyahya, ME Fessikh, LC Ming and T Aanniz. Epi-nutrients for cancer prevention: Molecular mechanisms and emerging insights. Cell Biology and Toxicology 2025; 41(1), 116.
[54] HY Ryu. Histone modification pathways suppressing cryptic transcription. Epigenomes 2024; 8(4), 42.
[55] Y Yu, X Li, R Jiao, Y Lu, X Jiang and X Li. H3K27me3-H3K4me1 transition at bivalent promoters instructs lineage specification in development. Cell & Bioscience 2023; 13(1), 66.
[56] E Bryan, D Valsakumar, NJ Idigo, M Warburton, KM Webb, KA McLaughlin, C Spanos, S Lenci, V Major, C Ambrosi, S Andrews, T Baubec, J Rappsilber and P Voigt. Nucleosomal asymmetry shapes histone mark binding and promotes poising at bivalent domains. Molecular Cell 2025; 85(3), 471-489.
[57] A Walewska, A Janucik, M Tynecka, M Moniuszko and A Eljaszewicz. Mesenchymal stem cells under epigenetic control - the role of epigenetic machinery in fate decision and functional properties. Cell Death & Disease 2023; 14, 720.
[58] Q Wang, F Xiong, G Wu, W Liu, J Chen, B Wang and Y Chen. Gene body methylation in cancer: Molecular mechanisms and clinical applications. Clinical Epigenetics 2022; 14, 154.
[59] AT Davletgildeeva and NA Kuznetsov. The role of DNMT methyltransferases and TET dioxygenases in the maintenance of the DNA methylation level. Biomolecules 2024; 14(9), 1117.
[60] S Chen, J Zhu, F Wang, Z Guan, Y Ge, X Yang and J Cai. LncRNAs and their role in cancer stem cells. Oncotarget 2017; 8(66), 110685-110692.
[61] G Divisato, S Piscitelli, M Elia, E Cascone and S Parisi. MicroRNAs and stem-like properties: The complex regulation underlying stemness maintenance and cancer development. Biomolecules 2021; 11(8), 1074.
[62] J Chen, Y Wang, C Wang, JF Hu and W Li. LncRNA functions as a new emerging epigenetic factor in determining the fate of stem cells. Frontiers in Genetics 2020; 11, 277.
[63] K Park, MC Jeon, D Lee, JI Kim and SW Im. Genetic and epigenetic alterations in aging and rejuvenation of human. Molecules and Cells 2024; 47(12), 100137.
[64] Y An, Q Wang, K Gao, C Zhang, Y Ouyang, R Li, Z Ma, T Wu, L Zhou, Z Xie, R Zhang and G Wu. Epigenetic regulation of aging and its rejuvenation. MedComm 2025; 6(9), 70369.
[65] WH Matsui. Cancer stem cell signaling pathways. Medicine (Baltimore) 2016; 95(1), 8-19.
[66] P Song, Z Gao, Y Bao, L Chen, Y Huang, Y Liu, Q Dong and X Wei. Wnt/β-catenin signaling pathway in carcinogenesis and cancer therapy. Journal of Hematology & Oncology 2024; 17, 46.
[67] M Tufail, CH Jiang and N Li. Wnt signaling in cancer: From biomarkers to targeted therapies and clinical translation. Molecular Cancer 2025; 24(1), 107.
[68] B Zhou, W Lin, Y Long, Y Yang, H Zhang, K Wu and Q Chu. Notch signaling pathway: Architecture, disease, and therapeutics. Signal Transduction and Targeted Therapy 2022; 7, 95.
[69] SJ Bray and M Gomez-Lamarca. Notch after cleavage. Current Opinion in Cell Biology 2018; 51, 103-109.
[70] S Parambath, NR Selvraj, P Venugopal and R Aradhya. Notch signaling: An emerging paradigm in the pathogenesis of reproductive disorders and diverse pathological conditions. International Journal of Molecular Sciences 2024; 25(10), 5423.
[71] V Kumar, M Vashishta, L Kong, X Wu, JJ Lu, C Guha and BS Dwarakanath. The role of Notch, hedgehog, and wnt signaling pathways in the resistance of tumors to anticancer therapies. Frontiers in Cell and Developmental Biology 2021; 9, 650772.
[72] GB Carballo, JR Honorato, GPF de Lopes and TCLSE Spohr. A highlight on Sonic hedgehog pathway. Cell Communication and Signaling 2018; 16, 11.
[73] AM Skoda, D Simovic, V Karin, V Kardum, S Vranic and L Serman. The role of the Hedgehog signaling pathway in cancer: A comprehensive review. Bosnian Journal of Basic Medical Sciences 2018; 18(1), 8-20.
[74] L Zhang, J Wang, N Xu, J Guo, Y Lin, X Zhang, R Ji, Y Ji, H Li, X Han, W Li, X Cheng, J Qin, M Tian, M Xu and X Zhang. Correction: POU3F4 up-regulates Gli1 expression and promotes neuronal differentiation and synaptic development of hippocampal neural stem cells. Stem Cell Research and Therapy 2024; 15(1), 501.
[75] J Jing, Z Wu, J Wang, G Luo, H Lin, Y Fan and C Zhou. Hedgehog signaling in tissue homeostasis, cancers, and targeted therapies. Signal Transduction and Targeted Therapy 2023; 8, 315.
[76] T Katagiri and T Watabe. Bone morphogenetic proteins. Cold Spring Harbor Perspectives in Biology 2016; 8(6), 021899.
[77] J Nickel and TD Mueller. Specification of BMP signaling. Cells 2019; 8(12), 1579.
[78] F Dituri, C Cossu, S Mancarella and G Giannelli. The interactivity between TGFβ and BMP signaling in organogenesis, fibrosis, and cancer. Cells 2019; 8(10), 1130.
[79] Q Li, S Geng, H Luo, W Wang, YQ Mo, Q Luo, L Wang, GB Song, JP Sheng and B Xu. Signaling pathways involved in colorectal cancer: Pathogenesis and targeted therapy. Signal Transduction and Targeted Therapy 2024; 9, 266.
[80] MM Mehlferber, M Kuyumcu-Martinez, CL Miller and GM Sheynkman. Transcription factors and splice factors - interconnected regulators of stem cell differentiation. Current Stem Cell Reports 2023; 9(2), 31-41.
[81] MH Lentjes, HE Niessen, Y Akiyama, AP de Bruïne, V Melotte and M van Engeland. The emerging role of GATA transcription factors in development and disease. Expert Reviews in Molecular Medicine 2016; 18, 3.
[82] PS Hoppe, M Schwarzfischer, D Loeffler, KD Kokkaliaris, O Hilsenbeck, N Moritz, M Endele, A Filipczyk, A Gambardella, N Ahmed, M Etzrodt, DL Coutu, MA Rieger, C Marr, MK Strasser, B Schauberger, I Burtscher, O Ermakova, A Bürger, H Lickert, …, T Schroeder. Early myeloid lineage choice is not initiated by random PU.1 to GATA1 protein ratios. Nature 2016; 535(7611), 299-302.
[83] Y Chen, R Liang, Y Li, L Jiang, D Ma, Q Luo and G Song. Chromatin accessibility: Biological functions, molecular mechanisms and therapeutic application. Signal Transduction and Targeted Therapy 2024; 9, 340.
[84] M Arnott, NF Sampilo and JL Song. Transcription of microRNAs is regulated by developmental signaling pathways and transcription factors. Frontiers in Cell and Developmental Biology 2024; 12, 1356589.
[85] P Perri, M Ponzoni, MV Corrias, I Ceccherini, S Candiani and T Bachetti. A focus on regulatory networks linking microRNAs, transcription factors and target genes in neuroblastoma. Cancers 2021; 13(21), 5528.
[86] J OʼBrien, H Hayder, Y Zayed and C Peng. Overview of microRNA biogenesis, mechanisms of actions, and circulation. Frontiers in Endocrinology (Lausanne) 2018; 9, 402.
[87] FH Nazki and CP Bracken. Regulation and dysregulation of microRNA - transcription factor axes in differentiation and neuroblastoma. Cellular and Molecular Life Sciences 2025; 82, 304.
[88] N Li, B Long, W Han, S Yuan and K Wang. microRNAs: Important regulators of stem cells. Stem Cell Research and Therapy 2017; 8, 110.
[89] A Drysch, A Ahuja, D Prasad, R Jain, S Romanos, A Alwakeal and C Ahuja. Transcriptomic comparisons of somatic and cancer stem cells. Biomedicines 2025; 13(8), 2039.
[90] B Srinageshwar, P Maiti, GL Dunbar and J Rossignol. Role of epigenetics in stem cell proliferation and differentiation: Implications for treating neurodegenerative diseases. International Journal of Molecular Sciences 2016; 17(2), 199.
[91] Y Cui, J Deng, Y Zhang, L Du, F Jiang, C Li, W Chen, H Zhang and Z He. Epigenetic regulation by DNA methylation, histone modifications and chromatin remodeling complexes in controlling spermatogenesis and their dysfunction with male infertility. Cellular and Molecular Life Sciences 2025; 82, 343.
[92] P Gujral, V Mahajan, AC Lissaman and AP Ponnampalam. Histone acetylation and the role of histone deacetylases in normal cyclic endometrium. Reproductive Biology and Endocrinology 2020; 18, 84.
[93] J Yuan, Q Jiang, T Gong, D Fan, J Zhang, F Chen, X Zhu, X Wang, Y Qiao, H Chen, Z Liu and J Su. Loss of grand histone H3 lysine 27 trimethylation domains mediated transcriptional activation in esophageal squamous cell carcinoma. npj Genomic Medicine 2021; 6, 65.
[94] Y Liu, Y Wang, D Bao, H Chen, M Gong, S Sun and G Zou. Cross-kingdom DNA methylation dynamics: Comparative mechanisms of 5mC/6mA regulation and their implications in epigenetic disorders. Biology 2025; 14(5), 461.
[95] M Yagi, S Yamanaka and Y Yamada. Epigenetic foundations of pluripotent stem cells that recapitulate in vivo pluripotency. Laboratory Investigation 2017; 97(10), 1133-1141.
[96] S Kumar, EA Gonzalez, P Rameshwar and JP Etchegaray. Non-coding RNAs as mediators of epigenetic changes in Malignancies. Cancers 2020; 12(12), 3657.
[97] MJ Pajares, E Alemany-Cosme, S Goñi, E Bandres, C Palanca-Ballester and J Sandoval. Epigenetic regulation of microRNAs in cancer: Shortening the distance from bench to bedside. International Journal of Molecular Sciences 2021; 22(14), 7350.
[98] B Xie and A Dean. Chromatin-associated RNAs regulate gene expression and chromatin structure. Noncoding RNA 2025; 11(5), 68.
[99] S Ji, M Xiong, H Chen, Y Liu, L Zhou, Y Hong, M Wang, C Wang, X Fu and X Sun. Cellular rejuvenation: Molecular mechanisms and potential therapeutic interventions for diseases. Signal Transduction and Targeted Therapy 2023; 8, 116.
[100] K Szabó, F Balogh, D Romhányi, L Erdei, B Toldi, R Gyulai, L Kemény and G Groma. Epigenetic regulatory processes involved in the establishment and maintenance of skin homeostasis-the role of microbiota. International Journal of Molecular Sciences 2025; 26(2), 438.
[101] B Yushkov, V Chereshnev, E Korneva, V Yushkova and A Sarapultsev. Stem-cell niches in health and disease: Microenvironmental determinants of regeneration and pathology. Cells 2025; 14(13), 981.
[102] A Marangio, A Biccari, E DʼAngelo, F Sensi, G Spolverato, S Pucciarelli and M Agostini. The study of the extracellular matrix in chronic inflammation: A way to prevent cancer initiation? Cancers 2022; 14(23), 5903.
[103] E Novoseletskaya, O Grigorieva, P Nimiritsky, N Basalova, R Eremichev, I Milovskaya, K Kulebyakin, M Kulebyakina, S Rodionov, N Omelyanenko and A Efimenko. Mesenchymal stromal cell-produced components of extracellular matrix potentiate multipotent stem cell response to differentiation stimuli. Frontiers in Cell and Developmental Biology 2020; 8, 555378.
[104] M Inaba, YM Yamashita and M Buszczak. Keeping stem cells under control: New insights into the mechanisms that limit niche-stem cell signaling within the reproductive system. Molecular Reproduction and Development 2016; 83(8), 675-683.
[105] Z Deng, T Fan, C Xiao, H Tian, Y Zheng, C Li and J He. TGF-β signaling in health, disease, and therapeutics. Signal Transduction and Targeted Therapy 2024; 9, 61.
[106] K Wagh, M Ishikawa, DA Garcia, DA Stavreva, A Upadhyaya and GL Hager. Mechanical regulation of transcription: Recent advances. Trends in Cell Biology 2021; 31(6), 457-72.
[107] BT Jackson and LWS Finley. Metabolic regulation of the hallmarks of stem cell biology. Cell Stem Cell 2024; 31(2), 161-180.
[108] W Chen, P Wu, F Yu, G Luo, L Qing and J Tang. HIF-1α regulates bone homeostasis and angiogenesis, participating in the occurrence of bone metabolic diseases. Cells 2022; 11(22), 3552.
[109] H Moukham, A Lambiase, GD Barone, F Tripodi and P Coccetti. Exploiting natural niches with neuroprotective properties: A comprehensive review. Nutrients 2024; 16(9), 1298.
[110] A Brunet, MA Goodell and TA Rando. Ageing and rejuvenation of tissue stem cells and their niches. Nature Reviews Molecular Cell Biology 2023; 24, 45-62.
[111] B Liu, J Qu, W Zhang, JCI Belmonte and GH Liu. A stem cell aging framework, from mechanisms to interventions. Cell Reports 2022; 41(3), 111451.
[112] J Muhr and DW Hagey. The cell cycle and differentiation as integrated processes: Cyclins and CDKs reciprocally regulate Sox and Notch to balance stem cell maintenance. Bioessays 2021; 43(7), 2000285.
[113] D Bačenková, M Trebuňová, E Dosedla, J Čajková and J Živčák. Interactions of hematopoietic and associated mesenchymal stem cell populations in the bone marrow microenvironment, in vivo and in vitro model. International Journal of Molecular Sciences 2025; 26(18), 9036.
[114] JG Lees, DK Gardner and AJ Harvey. Pluripotent stem cell metabolism and mitochondria: Beyond ATP. Stem Cells International 2017; 2017(1), 2874283.
[115] L Racine, R Parmentier, S Niphadkar, J Chhun, JA Martignoles, F Delhommeau, S Laxman and A Paldi. Metabolic adaptation pilots the differentiation of human hematopoietic cells. Life Science Alliance 2024; 7(8), 202402747.
[116] S Thiagalingam. Epigenetic memory in development and disease: Unraveling the mechanism. Biochimica et Biophysica Acta: Reviews on Cancer 2020; 1873(2), 188349.
[117] D Gökbuget and R Blelloch. Epigenetic control of transcriptional regulation in pluripotency and early differentiation. Development 2019; 146(19), 164772.
[118] A Basu and VK Tiwari. Epigenetic reprogramming of cell identity: Lessons from development for regenerative medicine. Clinical Epigenetics 2021; 13, 144.
[119] M Abdul-Al, GK Kyeremeh, M Saeinasab, SH Keshel and F Sefat. Stem cell niche microenvironment: Review. Bioengineering 2021; 8(8), 108.
[120] N Khorasani, M Sadeghi and A Nowzari-Dalini. A computational model of stem cell molecular mechanism to maintain tissue homeostasis. PLoS One 2020; 15(7), 0236519.
[121] M Chehelgerdi, FB Dehkordi, M Chehelgerdi, H Kabiri, H Salehian-Dehkordi, M Abdolvand, S Salmanizadeh, M Rashidi, A Niazmand, S Ahmadi, S Feizbakhshan, S Kabiri, N Vatandoost and T Ranjbarnejad. Exploring the promising potential of induced pluripotent stem cells in cancer research and therapy. Molecular Cancer 2023; 22, 189.
[122] MF Pittenger, DE Discher, BM Péault, DG Phinney, JM Hare and AI Caplan. Mesenchymal stem cell perspective: Cell biology to clinical progress. npj Regenerative Medicine 2019; 4, 22.
[123] I Rosa, M Marini and M Manetti. Telocytes: An emerging component of stem cell niche microenvironment. Journal of Histochemistry & Cytochemistry 2021; 69(12), 795-818.
[124] W Manni and W Min. Signaling pathways in the regulation of cancer stem cells and associated targeted therapy. MedComm 2022; 3(4), 176.
[125] F Lasorsa, M Rutigliano, M Milella, M Ferro, SD Pandolfo, F Crocetto, R Autorino, M Battaglia, P Ditonno and G Lucarelli. Cancer stem cells in renal cell carcinoma: Origins and biomarkers. International Journal of Molecular Sciences 2023; 24(17), 13179.
[126] S Chao, H Yan and P Bu. Asymmetric division of stem cells and its cancer relevance. Cell Regeneration 2024; 13, 5.
[127] JJ Bower, LD Vance, M Psioda, SL Smith-Roe, DA Simpson, JG Ibrahim, KA Hoadley, CM Perou and WK Kaufmann. Patterns of cell cycle checkpoint deregulation associated with intrinsic molecular subtypes of human breast cancer cells. npj Breast Cancer 2017; 3, 9.
[128] P Geara and FJ Dilworth. Epigenetic integration of signaling from the regenerative environment. Current Topics in Developmental Biology 2024; 158, 341-374.
[129] N Swain, M Thakur, J Pathak and B Swain. SOX2, OCT4 and NANOG: The core embryonic stem cell pluripotency regulators in oral carcinogenesis. Journal of Oral and Maxillofacial Pathology 2020; 24(2), 368-373.
[130] VF Fiore, J Almagro and E Fuchs. Shaping epithelial tissues by stem cell mechanics in development and cancer. Nature Reviews Molecular Cell Biology 2025; 26, 442-455.
[131] MA Sharifi, NF Yasin, T Kamarul and AM Sharifi. The regenerative journey: Exploring stem cell roles from injury detection to tissue repair. Stem Cell Research and Therapy 2025; 16, 541.
[132] X Zhang, D Zhang, Y Zhang, J Wang and J Lu. Restoration of skeletal muscle function via mesenchymal stem cells: Mechanistic insights and therapeutic advances in myasthenia gravis. Frontiers in Cell and Developmental Biology 2025; 13, 1658062.
[133] M Engel, D Do-Ha, SS Muñoz and L Ooi. Common pitfalls of stem cell differentiation: A guide to improving protocols for neurodegenerative disease models and research. Cellular and Molecular Life Sciences 2016; 73, 3693-3709.
[134] I Pereira, MJ Lopez-Martinez and J Samitier. Advances in current in vitro models on neurodegenerative diseases. Frontiers in Bioengineering and Biotechnology 2023; 11, 1260397.
[135] L Sutherland, M Ruhe, D Gattegno-Ho, K Mann, J Greaves, M Koscielniak, S Meek, Z Lu, M Waterfall, R Taylor, A Tsakiridis, H Brown, SK Maciver, A Joshi, M Clinton, LH Chamberlain, A Smith and T Burdon. LIF-dependent survival of embryonic stem cells is regulated by a novel palmitoylated Gab1 signalling protein. Journal of Cell Science 2018; 131(18), 222257.
[136] AM Setiawan, TA Kamarudin and NA Ghafar. The role of BMP4 in adipose-derived stem cell differentiation: A minireview. Frontiers in Cell and Developmental Biology 2022; 10, 1045103.
[137] J Cerneckis, H Cai and Y Shi. Induced pluripotent stem cells (iPSCs): Molecular mechanisms of induction and applications. Signal Transduction and Targeted Therapy 2024; 9, 112.
[138] E Suarez-Martinez, I Suazo-Sanchez, M Celis-Romero and A Carnero. 3D and organoid culture in research: Physiology, hereditary genetic diseases and cancer. Cell & Bioscience 2022; 12, 39.
[139] X Zhao, Q Li, Z Guo and Z Li. Constructing a cell microenvironment with biomaterial scaffolds for stem cell therapy. Stem Cell Research and Therapy 2021; 12, 583.
[140] D Perepletchikova, P Kuchur, L Basovich, I Khvorova, A Lobov, K Azarkina, N Aksenov, S Bozhkova, V Karelkin and A Malashicheva. Endothelial-mesenchymal crosstalk drives osteogenic differentiation of human osteoblasts through Notch signaling. Cell Communication and Signaling 2025; 23, 100.
[141] R Cruz-Acuña and AJ García. Synthetic hydrogels mimicking basement membrane matrices to promote cell-matrix interactions. Matrix Biology 2017; 57-58, 324-333.
[142] I Serganova and RG Blasberg. Molecular imaging with reporter genes: Has its promise been delivered? The Journal of Nuclear Medicine 2019; 60(12), 1665-1681.
[143] J Bayerl, M Ayyash, T Shani, YS Manor, O Gafni, R Massarwa, Y Kalma, A Aguilera-Castrejon, M Zerbib, H Amir, D Sheban, S Geula, N Mor, L Weinberger, SN Tassa, V Krupalnik, B Oldak, N Livnat, S Tarazi, S Tawil, …, JH Hanna. Principles of signaling pathway modulation for enhancing human naive pluripotency induction. Cell Stem Cell 2021; 28(9), 1549-1565.
[144] Z Lin, S Aryal, YH Cheng and AJ Gesquiere. Integration of in vitro and in vivo models to predict cellular and tissue dosimetry of nanomaterials using physiologically based pharmacokinetic modeling. ACS Nano 2022; 16(12), 19722-19754.
[145] B Wen, G Wang, E Li, OA Kolesnichenko, Z Tu, S Divanovic, TV Kalin and VV Kalinichenko. In vivo generation of bone marrow from embryonic stem cells in interspecies chimeras. Elife 2022; 11, 74018.
[146] S Park, MK Ha, Y Lee, J Song and TH Yoon. Effects of immune cell heterogeneity and protein corona on the cellular association and cytotoxicity of gold nanoparticles: A single-cell-based, high-dimensional mass cytometry study. ACS Nanoscience Au 2023; 3(4), 323-334.
[147] T Wang, X Chen, K Wang, J Ju, X Yu, S Wang, C Liu and K Wang. Cre-loxP-mediated genetic lineage tracing: Unraveling cell fate and origin in the developing heart. Frontiers in Cardiovascular Medicine 2023; 10, 1085629.
[148] P Saeedi, LS Nilchiani, B Zand, M Hajimirghasemi and R Halabian. An overview of stem cells and cell products involved in trauma injury. Regenerative Therapy 2025; 29, 60-76.
[149] HI Chen, JA Wolf, R Blue, MM Song, JD Moreno, GL Ming and H Song. Transplantation of human brain organoids: Revisiting the science and ethics of brain chimeras. Cell Stem Cell 2019; 25(4), 462-472.
[150] ZS Razavi, S Farokhi, G Mahmoudvand, A Karimi-Rouzbahani, B Farasati-Far, S Tahmasebi-Ghorabi, H Pazoki-Toroudi, M Saadat-Fakhr and H Afkhami. Stem cells and bio scaffolds for the treatment of cardiovascular diseases: New insights. Frontiers in Cell and Developmental Biology 2024; 12, 1472103.
[151] QY Guo, JQ Yang, XX Feng and YJ Zhou. Regeneration of the heart: From molecular mechanisms to clinical therapeutics. Military Medical Research 2023; 10, 18.
[152] L Bagno, KE Hatzistergos, W Balkan and JM Hare. Mesenchymal stem cell-based therapy for cardiovascular disease: Progress and challenges. Molecular Therapy 2018; 26(7), 1610-1623.
[153] FM Moraddahande, SME Meybodi, M Matin, N Soleimani, N Ghasemzadeh and AD Firoozabadi. Current status and new horizons in stem cell therapy in cardiovascular regenerative medicine (CaVaReM): An update. European Journal of Medical Research 2025; 30, 837.
[154] Z Huang, K Jia, Y Tan, Y Yu, W Xiao, X Zhou, J Yi and C Zhang. Advances in cardiac organoid research: Implications for cardiovascular disease treatment. Cardiovascular Diabetology 2025; 24, 25.
[155] Y Fu, X Rui, S Zhu, C Guo, H Li, Z Pan, X Wu and W He. Research status of regenerative difficulties after central nervous system injury. Regenerative Therapy 2025; 29, 493-498.
[156] N Nagoshi, M Khazaei, JE Ahlfors, CS Ahuja, S Nori, J Wang, S Shibata and MG Fehlings. Human spinal oligodendrogenic neural progenitor cells promote functional recovery after spinal cord injury by axonal remyelination and tissue sparing. Stem Cells Translational Medicine 2018; 7(11), 806-818.
[157] V Jagrit, J Koffler and JN Dulin. Combinatorial strategies for cell transplantation in traumatic spinal cord injury. Frontiers in Neuroscience 2024; 18, 1349446.
[158] S Ahmadi, M Nabiuni, M Tahmaseb and E Amini. Enhanced neural differentiation of epidermal neural crest stem cell by synergistic effect of lithium carbonate and crocin on BDNF and GDNF expression as neurotrophic factors. Iranian Journal of Pharmaceutical Research 2021; 20(2), 95-106.
[159] S Shaikh, L Siddique, HT Khalifey, R Mahereen, T Raziq, RM Firdous, A Siddique, IM Shakir, Z Ahmed, A Akbar, EA Alshehri, R Chinappan, A Alzhrani, TA Mir and A Yaqinuddin. Brain organoid model systems of neurodegenerative diseases: Recent progress and future prospects. Frontiers in Neuroscience 2025; 19, 1604435.
[160] S Hora and T Wuestefeld. Liver injury and regeneration: Current understanding, new approaches, and future perspectives. Cells 2023; 12(17), 2129.
[161] A Messina, E Luce, M Hussein and A Dubart-Kupperschmitt. Pluripotent-stem-cell-derived hepatic cells: Hepatocytes and organoids for liver therapy and regeneration. Cells 2020; 9(2), 420.
[162] HR Moon, SJ Mun, TH Kim, H Kim, D Kang, S Kim, JH Shin, D Choi, SJ Ahn and MJ Son. Guidelines for manufacturing and application of organoids: Liver. International Journal of Stem Cells 2024; 17(2), 120-129.
[163] J Sun, W Chang, B Li, X Zhang, Y Deng, Y Cai, Z Gu and Z Xie. Organoids pharmacology and toxicology. EngMedicine 2025; 2(3), 100089.
[164] L Wei, W Yan, W Shah, Z Zhang, M Wang, B Liu, Z Xue, Y Cao, X Hou, K Zhang, B Yan and X Wang. Advancements and challenges in stem cell transplantation for regenerative medicine. Heliyon 2024; 10(16), 35836.
[165] M Shafiq, Y Jung and SH Kim. Insight on stem cell preconditioning and instructive biomaterials to enhance cell adhesion, retention, and engraftment for tissue repair. Biomaterials 2016; 90, 85-115.
[166] S Zhidu, T Ying, J Rui and Z Chao. Translational potential of mesenchymal stem cells in regenerative therapies for human diseases: Challenges and opportunities. Stem Cell Research and Therapy 2024; 15, 266.
[167] T Tominari, C Sathyaprakash and Y Aoki. Stem/progenitor cell-based therapy for Duchenne muscular dystrophy. Frontiers in Cell and Developmental Biology 2025; 13, 1640275.
[168] J Chen, R Zhou, Y Feng and L Cheng. Molecular mechanisms of exercise contributing to tissue regeneration. Signal Transduction and Targeted Therapy 2022; 7, 383.
[169] A Shao, S Tu, J Lu and J Zhang. Crosstalk between stem cell and spinal cord injury: Pathophysiology and treatment strategies. Stem Cell Research and Therapy 2019; 10, 238.
[170] J Han, X Zhang, L Kang and J Guan. Extracellular vesicles as therapeutic modulators of neuroinflammation in Alzheimerʼs disease: A focus on signaling mechanisms. Journal of Neuroinflammation 2025; 22, 120.
[171] QD Vo, Y Saito, K Nakamura, T Iida and S Yuasa. Induced pluripotent stem cell-derived cardiomyocytes therapy for ischemic heart disease in animal model: A meta-analysis. International Journal of Molecular Sciences 2024; 25(2), 987.
[172] SZ Moradi, F Jalili, Z Hoseinkhani and K Mansouri. Regenerative medicine and angiogenesis; focused on cardiovascular disease. Advanced Pharmaceutical Bulletin 2022; 12(4), 686-699.
[173] X Du, H Jia, Y Chang, Y Zhao and J Song. Progress of organoid platform in cardiovascular research. Bioactive Materials 2024; 40, 88-103.
[174] D Han, W Liu, J Gong, Y Ma and Z Sun. Challenges and future perspectives in using mesenchymal stem cells for efficient bone fracture healing. Frontiers in Bioengineering and Biotechnology 2025; 13, 1568914.
[175] WH Abuwatfa, WG Pitt and GA Husseini. Scaffold-based 3D cell culture models in cancer research. Journal of Biomedical Science 2024; 31, 7.
[176] DT Chu, TT Nguyen, NLB Tien, DK Tran, JH Jeong, PG Anh, VV Thanh, DT Truong and TC Dinh. Recent progress of stem cell therapy in cancer treatment: Molecular mechanisms and potential applications. Cells 2020; 9(3), 563.
[177] LTH Phi, IN Sari, YG Yang, SH Lee, N Jun, KS Kim, YK Lee and HY Kwon. Cancer stem cells (CSCs) in drug resistance and their therapeutic implications in cancer treatment. Stem Cells International 2018; 2018, 5416923.
[178] X Chu, W Tian, J Ning, G Xiao, Y Zhou, Z Wang, Z Zhai, G Tanzhu, J Yang and R Zhou. Cancer stem cells: Advances in knowledge and implications for cancer therapy. Signal Transduction and Targeted Therapy 2024; 9, 170.
[179] M Mazloomi, A Doustmihan, S Alimohammadvand, H Hamishehkar, MR Hamblin and RJ Esfahlan. Advanced drug delivery platforms target cancer stem cells. Asian Journal of Pharmaceutical Sciences 2025; 20(3), 101036.
[180] X Xuan, C Tian, M Zhao, Y Sun and C Huang. Mesenchymal stem cells in cancer progression and anticancer therapeutic resistance. Cancer Cell International 2021; 21, 595.
[181] J Bae, YS Choi, G Cho and SJ Jang. The patient-derived cancer organoids: Promises and challenges as platforms for cancer discovery. Cancers 2022; 14(9), 2144.
[182] S Isik, S Osman, B Yeman-Kiyak, SRM Shamshir and NME Sanchez. Advances in neurodegenerative disease therapy: Stem cell clinical trials and promise of engineered exosomes. CNS Neuroscience and Therapeutics 2025; 31(9), 70577.
[183] SV Mathan and RP Singh. Cancer stem cells connecting to immunotherapy: Key insights, challenges, and potential treatment opportunities. Cancers 2025; 17(13), 2100.
[184] BI Mendi, R Hirani, A Sayegh, M Hassan, L Fleshner, B Farabi, MF Atak and B Safai. Utilization of stem cells in medicine: A narrative review. International Journal of Molecular Sciences 2025; 26(19), 9659.
[185] P Nitti, A Narayanan, R Pellegrino, S Villani, M Madaghiele and C Demitri. Cell-tissue interaction: The biomimetic approach to design tissue engineered biomaterials. Bioengineering 2023; 10(10), 1122.
[186] AM Sacco, C Castaldo, FD Meglio, D Nurzynska, S Palermi, R Spera, R Gnasso, G Zinno, V Romano and I Belviso. The long and winding road to cardiac regeneration. Applied Sciences 2023; 13(16), 9432.
[187] F Wang, X Xie, X Xing and X Sun. Excitatory synaptic transmission in ischemic stroke: A new outlet for classical neuroprotective strategies. International Journal of Molecular Sciences 2022; 23(16), 9381.
[188] MS Lach, MA Rosochowicz, M Richter, I Jagiełło, WM Suchorska and T Trzeciak. The induced pluripotent stem cells in articular cartilage regeneration and disease modelling: Are we ready for their clinical use? Cells 2022; 11(3), 529.
[189] S Gao, X Hou, Y Jiang, Z Xu, T Cai, J Chen and G Chang. Integrated analysis of hematopoietic differentiation outcomes and molecular characterization reveals unbiased differentiation capacity and minor transcriptional memory in HPC/HSC-iPSCs. Stem Cell Research and Therapy 2017; 8, 13.
[190] AY Movahed, R Bagheri, P Savatier, T Šarić and S Moradi. Elimination of tumorigenic pluripotent stem cells from their differentiated cell therapy products: An important step toward ensuring safe cell therapy. Stem Cell Reports 2025; 20(7), 102543.
[191] J Han, D Lim and K Yang. Gene editing strategies to address current challenges in stem cell-derived β cell therapy for type 1 Diabetes. Journal of Tissue Engineering 2025; 16, 20417314251373039.
[192] K Wuputra, CC Ku, DC Wu, YC Lin, S Saito and KK Yokoyama. Prevention of tumor risk associated with the reprogramming of human pluripotent stem cells. Journal of Experimental and Clinical Cancer Research 2020; 39, 100.
[193] O Gordeeva and S Khaydukov. Tumorigenic and differentiation potentials of embryonic stem cells depend on TGFβ family signaling: Lessons from teratocarcinoma cells stimulated to differentiate with retinoic acid. stem cells international 2017; 2017, 7284872.
[194] B Liu, Y Lu, A Taledaohan, S Qiao, Q Li and Y Wang. The promoting role of HK II in tumor development and the research progress of its inhibitors. Molecules 2023; 29(1), 75.
[195] A Ahmad, M Imran and H Ahsan. Biomarkers as biomedical bioindicators: Approaches and techniques for the detection, analysis, and validation of novel biomarkers of diseases. Pharmaceutics 2023; 15(6), 1630.
[196] GE Leonov, LR Grinchevskaya, OV Makhnach, MV Samburova, DV Goldshtein and DI Salikhova. Safety assessment of stem cell-based therapies: Current standards and advancing frameworks. Cells 2025; 14(21), 1660.
[197] W Zhou, X Liu, Z Li, B Jia, X Lei, K Sun, P Yang, S He, D Wang, H Zhang and S Wang. Epigenetic crosstalk between stem cells and tumors: Mechanisms and emerging perspectives. American Journal of Stem Cells 2025; 14(3), 98-133.
[198] BN Kharbikar, P Mohindra and TA Desai. Biomaterials to enhance stem cell transplantation. Cell Stem Cell 2022; 29(5), 692-721.
[199] EH Moretti, ALY Lin, L Peruzzotti-Jametti, S Pluchino and S Mozafari. Neural stem cell-derived extracellular vesicles for advanced neural repair. Journal of Neurochemistry 2025; 169(8), 70170.
[200] RS Mahla. Stem cells applications in regenerative medicine and disease therapeutics. International Journal of Cell Biology 2016; 2016, 6940283.
[201] YD Rocca, A Mazzone, GD Marconi, O Trubiani, J Pizzicannella and F Diomede. Stem cells in regenerative medicine: A journey from adult stem cells to induced pluripotent cells. International Journal of Molecular Sciences 2025; 26(17), 8255.
[202] G Chen, S Yin, H Zeng, H Li and X Wan. Regulation of embryonic stem cell self-renewal. Life 2022; 12(8), 1151.
[203] SH Sharami, E Vojoudi, Z Zarei-Behjani, A Hasanzadi, SS Shoarishoar, A Vaez and Z Barabadi. From challenges to solutions: The promise of regenerative medicine and tissue engineering in restoring female reproductive health. Heliyon 2025; 11(12), 43484.
[204] Y Chen, M Li and Y Wu. The occurrence and development of induced pluripotent stem cells. Frontiers in Genetics 2024; 15, 1389558.
[205] K Moskal, N Khurana, L Siegert, YS Lee, H Clevers, E Elinav and J Puschhof. Modeling cancer-microbiome interactions in vitro: A guide to co-culture platforms. International Journal of Cancer 2025; 156(11), 2053-2067.
[206] L Hu, C Yin, F Zhao, A Ali, J Ma and A Qian. Mesenchymal stem cells: Cell fate decision to osteoblast or adipocyte and application in osteoporosis treatment. International Journal of Molecular Sciences 2018; 19(2), 360.
[207] X Han, R Liao, X Li, C Zhang, S Huo, L Qin, Y Xiong, T He, G Xiao and T Zhang. Mesenchymal stem cells in treating human diseases: Molecular mechanisms and clinical studies. Signal Transduction and Targeted Therapy 2025; 10, 262.
[208] D Bian, Y Wu, G Song, R Azizi and A Zamani. The application of mesenchymal stromal cells (MSCs) and their derivative exosome in skin wound healing: A comprehensive review. Stem Cell Research and Therapy 2022; 13, 24.
[209] A Musiał-Wysocka, M Kot and M Majka. The pros and cons of mesenchymal stem cell-based therapies. Cell Transplantation 2019; 28(7), 801-812.
[210] AP Reyes and SM Calado. human stem cells in disease modelling and treatment: Bridging the gap between bench and bedside. Biomedicines 2025; 13(9), 2313.
[211] I Mastrolia, EM Foppiani, A Murgia, O Candini, AV Samarelli, G Grisendi, E Veronesi, EM Horwitz and M Dominici. Challenges in clinical development of mesenchymal stromal/stem cells: Concise review. Stem Cells Translational Medicine 2019; 8(11), 1135-1148.
[212] D Shi, G Mi, M Wang and TJ Webster. In vitro and ex vivo systems at the forefront of infection modeling and drug discovery. Biomaterials 2019; 198, 228-249.
[213] YC Lin, CC Ku, K Wuputra, CJ Liu, DC Wu, M Satou, Y Mitsui, S Saito and KK Yokoyama. Possible strategies to reduce the tumorigenic risk of reprogrammed normal and cancer cells. International Journal of Molecular Sciences 2024; 25(10), 5177.
[214] JM Alamillo, CM López, FJM Rivas, F Torralbo, M Bulut and S Alseekh. Clustered regularly interspaced short palindromic repeats/CRISPR-associated protein and hairy roots: A perfect match for gene functional analysis and crop improvement. Current Opinion in Biotechnology 2023; 79, 102876.
[215] CK Bektas, J Luo, B Conley, KN Le and KB Lee. 3D bioprinting approaches for enhancing stem cell-based neural tissue regeneration. Acta Biomaterialia 2025; 193, 20-48.
[216] K Loukelis, N Koutsomarkos, AG Mikos and M Chatzinikolaidou. Advances in 3D bioprinting for regenerative medicine applications. Regenerative Biomaterials 2024; 11, 033.
[217] Y Zhou, L Tao, J Qiu, J Xu, X Yang, Y Zhang, X Tian, X Guan, X Cen and Y Zhao. Tumor biomarkers for diagnosis, prognosis and targeted therapy. Signal Transduction and Targeted Therapy 2024; 9, 132.
[218] S Shan, Q Li, T Criswell, A Atala and Y Zhang. Stem cell therapy combined with controlled release of growth factors for the treatment of sphincter dysfunction. Cell & Bioscience 2023; 13(1), 56.