
Trends
Sci.
2026;
23(6):
12369
Blumea balsamifera Extract Induces Apoptosis and Inhibit Cell Migration on HepG2 Hepatocellular Carcinoma: An Integrated Experimental and Computational Study
Muhammad Hermawan Widyananda1, Adinda Romdiatus Saadah2, Yuslinda Annisa2, Nuraini Rosyadah2, Dinia Rizqi Dwijayanti2, Feri Eko Hermanto3 and Nashi Widodo1,2,*
1Biosystem Study Center, Universitas Brawijaya, Malang, Indonesia
2Department of Biology, Faculty of Mathematics and Natural Sciences, Universitas Brawijaya, Malang, Indonesia
3Department of Intelligent Livestock Industry, Faculty of Animal Science, Universitas Brawijaya, Malang, Indonesia
(*Corresponding author’s e-mail: [email protected])
Received: 21 October 2025, Revised: 28 November 2025, Accepted: 15 December 2025, Published: 15 February 2026
Abstract
Hepatocellular carcinoma (HCC) is one of the most aggressive types of cancer and requires alternative therapeutic strategies beyond current treatments. Blumea balsamifera is a medicinal plant with potential anticancer properties that may offer a novel approach for HCC treatment. This study aimed to evaluate the anti-HCC activity of B. balsamifera extract through an integrated experimental and computational approach. Experimental assays were performed to assess cytotoxicity, apoptosis induction, mitochondrial membrane potential, and cell migration, while computational analyses were applied to predict the molecular mechanisms involved using network pharmacology, molecular docking, and molecular dynamic simulation. The extract demonstrated cytotoxic activity against HepG2 cells (IC₅₀ = 199.09 ± 11.86 µg/mL) and significantly induced apoptosis in a dose-dependent manner, with the IC₅₀ concentration triggering apoptosis in more than 80% of cells. In addition, B. balsamifera extract markedly inhibited HepG2 cell migration, with 74% of the wound gap remaining open at the IC₅₀ concentration. Network pharmacology analysis further predicted that the bioactive compounds of B. balsamifera may act by targeting key HCC-related proteins involved in cell death and migration pathways, including TOP1, AKT1, PPM1D, SRC, AKR1C2, DNMT1, and PTH. Collectively, these findings provide both experimental evidence and computational insights supporting the potential of B. balsamifera as a promising natural therapeutic candidate for hepatocellular carcinoma.
Keywords: Apoptosis, Blumea balsamifera, Cell migration, Hepatocellular carcinoma
Introduction
Hepatocellular
carcinoma (HCC) is the most common liver cancer and remains a major
global health burden [1].
It ranks as the sixth most frequently diagnosed cancer and the third
leading cause of
cancer-related mortality worldwide, with
increasing incidence particularly in regions with high prevalence of
hepatitis B and C virus infection, alcohol abuse, and metabolic
liver diseases [2].
Previous epidemiological studies reported that HBV infection had
strong correlation with hepatocarcinogenicity [3].
Despite advances in diagnosis and treatment, the prognosis of HCC
remains poor due to its late detection, aggressive progression, and
high recurrence rate [4].
Hepatocellular carcinoma (HCC) is characterized by the dysregulation
of multiple signaling pathways, which makes its treatment
particularly challenging.
Hepatocellular carcinoma (HCC) is characterized by the dysregulation of multiple signaling pathways that drive tumor initiation and progression. For example, the PI3K–AKT signaling pathway is frequently activated through the upregulation of EGFR, PI3K, AKT, and mTORC1, along with the downregulation or mutation of negative regulators such as TSC1/2 and PTEN [5]. Similarly, the JAK–STAT signaling pathway is often altered; JAK1 missense mutations that promote cytokine-independent growth have been identified in approximately 9% of HBV-related HCC cases [6]. The MAPK signaling pathway also undergoes frequent dysregulation, particularly due to RAS mutations that enhance cell proliferation and survival [7]. Collectively, these molecular alterations complicate therapeutic management, as conventional chemotherapeutic agents often lead to drug resistance and undesirable side effects [8]. Therefore, multi-target therapeutic strategies are required, and herbal medicines have emerged as promising candidates. Natural compounds are especially appealing because of their structural diversity, broad biological activities, and ability to modulate multiple molecular targets consistent with the complex pathophysiology of HCC.
Blumea balsamifera (family Asteraceae) is a medicinal plant widely used in traditional medicine throughout Southeast Asia [9]. It has been reported to exhibit diverse pharmacological properties, including antioxidant, anti-inflammatory, and anticancer activities [10,11]. Extracts of B. balsamifera have demonstrated significant cytotoxic effects against various cancer cell types, including cervical and leukemia cells [12,13]. These anticancer effects are primarily attributed to its rich content of bioactive compounds such as blumeatin, quercetin, luteolin, and several other phenolic constituents [14]. Previous research has shown that B. balsamifera extract exerts antiproliferative effects on HepG2 cells by downregulating the expression of retinoblastoma (Rb), cyclin, and a proliferation-inducing ligand (APRIL) [15]. However, further studies exploring the anti-hepatocellular carcinoma mechanisms of B. balsamifera remain limited. Therefore, the present study aims to provide new insights into the anti-HCC potential of B. balsamifera extract, particularly through mechanisms involving apoptosis induction and inhibition of cell migration.
In this study, we investigated the anticancer activity of B. balsamifera ethanolic extract against HepG2 hepatocellular carcinoma cells using an integrated experimental and computational approach. Specifically, we evaluated its cytotoxic, pro-apoptotic, and anti-migratory effects through in vitro assays, while complementing these findings with network pharmacology analyses to predict molecular targets and pathways involved. The objective of this study was to provide both experimental evidence and computational insights into the potential of B. balsamifera as a candidate natural product–based therapeutic for HCC.
Materials and methods
Extraction of Blumea balsamifera
Blumea balsamifera leaves (Batch No. 220530SBL.F.MMB) were obtained from the UPT Herbal Materia Medika Laboratory in Batu, East Java, Indonesia. Six grams of B. balsamifera powder was added to 60 mL of 96% ethanol (Merck, Germany) (1:10 ratio). Extraction was performed using the Microwave-Assisted Extraction (MAE) method (Anton-Paar, Austria) controlled according to the specific protocol, a holding temperature of 50 °C for 20 min (5 min warming up, 10 min holding time, and 5 min cooling down) at a power level of 1,500 Watts. The extract was filtered, and then evaporated using a Buchi R-210 rotary evaporator (50 rpm, 37 °C) (ThermoScientific, Singapore) [16].
HepG2 Cell preparation
The liver cancer cell line HepG2 were obtained from the Animal Physiology Anatomy Laboratory, Brawijaya University, Malang. The cells were grown in DMEM (Dulbecco’s Modified Eagle Medium) high glucose basal medium (Gibco, USA) supplemented with 10% Fetal Bovine Serum (Gibco, USA) and 1% penicillin-streptomycin (Gibco, USA). The cells were incubated at 37 °C and 5% CO2.
Cytotoxicity assay
The HepG2 cells were seeded in a 96-well plate (NEST Scientific, USA) at a density of 7,500 cells/well and incubated at 37 °C 5% CO2 for 24 h. The B. balsamifera extract was first dissolved in dimethyl sulfoxide (DMSO) to prepare a 100,000 µg/mL stock solution, then diluted in complete medium to obtain final treatment concentrations of 0 (Control), 20, 40, 80, 160, and 320 µg/mL. The cells were treated to these concentrations and incubated for 24 h. The treatment medium was replaced by 5% WST-1 (Sigma-Aldrich, USA) and incubated for 30 min in CO2 incubator. Absorbance measurements were performed in 450 nm using Multiskan skyhigh Microplate Spectrophotometer (Thermo Scientific, USA). The absorbance values were used to calculate cell viability, and the IC₅₀ value was determined by plotting the percentage of viable cells against the logarithmic concentrations of the extract and fitting the data to a linear regression dose–response curve. The assay was performed in three independent replications [17].
Apoptosis assay
The cells were seeded in 24-well plates at a density of 75,000 cells per well and incubated at 37°C with 5% CO₂ for 24 h. The cells were treated with B. balsamifera extract at concentrations of 0 µg/mL (untreated/negative control), 99.55, 199.09, and 398.18 µg/mL, as well as 5 µg/mL cisplatin (positive control), and further incubated for an additional 24 h. The centrifugation pellet was added with 50 µL annexin-V and PI (1:2 ratio, desolved in binding buffer) (BioLegend, USA), then incubated in the dark for 20 min. Apoptosis analysis was performed using a flow cytometer (BD FACSCalibur™, San Jose, CA), and the data were analyzed using FlowJO v10.8.1. (BD, USA). Flow cytometry analysis began with FSC–SSC gating to exclude debris and isolate intact cell populations. Annexin V–FITC/PI staining was then analyzed using quadrant gating to differentiate viable, early apoptotic, late apoptotic, and necrotic cells. The assay was conducted in triplicate [18].
Rhodamine 123 mitochondrial membrane potential assay
Cells were seeded in 24-well plates at a density of 75,000 cells per well and incubated overnight at 37 °C with 5% CO₂. The cells were treated with B. balsamifera extract at concentrations of 0 µg/mL (untreated), 99.55, 199.09, and 398.18 µg/mL, as well as 5 µg/mL cisplatin, and incubated for 24 h. After treatment, cells were exposed to 2 µM rhodamine 123 and incubated for 1 h in a CO₂ incubator. The cells were then centrifuged, resuspended in DMEM-HG basal medium, and incubated for 20 min at 4 °C. Following PBS washing, samples were analyzed by flow cytometry, and data were processed using CellQuest software (BD Biosciences, USA). Debris was excluded using FSC-SSC gating, and mitochondrial membrane potential loss was quantified using histogram-based gating of rhodamine 123 fluorescence. The assays were performed in triplicate [18].
Cell migration assay
HepG2 cells were seeded in a 24-well plate at a density of 3×105 cells/well and incubated for 24 h. The cells were scratched vertically using a sterile yellow tip. The detached cells were washed with PBS. The cells were treated with extracts at doses of 0, 99.55, 199.09, 398.18 µg/mL, and 5 µg/mL Cisplatin. Cell migration was observed at 0, 12, and 24 h after treatment using an inverted phase-contrast microscope (10×magnification) (Olumpus IX71, Japan). Wound closure was quantified using ImageJ with the MRI_Wound_Healing_Tool.ijm macro (U.S. National Institutes of Health, USA), which automatically thresholds the images and identifies the wound region. The macro calculates the gap area by measuring the pixel area of the cell-free zone in each image. The migration assay was conducted in triplicate [17].
Statistical analysis
Data analysis was conducted using IBM SPSS Statistics version 23 (IBM Corp., USA). The results were expressed as mean ± standard deviation (SD). Group differences were evaluated using one-way analysis of variance (ANOVA) followed by Tukey’s HSD post hoc test. A p-value of ≤ 0.01 was considered statistically significant.
Target protein prediction
The active compounds of B. balsamifera extract were obtained from our previous study using liquid chromatography high resolution mass spectrometry [19]. The target protein of each active compound were predicted using STITCH server (http://stitch.embl.de/) [20]. STITCH settings were configured using Homo sapiens as the species, the full STRING network as the network type, a 0.4 confidence cut-off, and a maximum of 50 interactions. All targets protein of each active compound were merged with proteins related to hepatocellular carcinoma from GeneCards (https://www.genecards.org/) using Venny diagram 2.1 (https://bioinfogp.cnb.csic.es/tools/venny/). The complex interaction between active compound and the target proteins were visualized using Cytoscape 3.10.3.
Hub protein prediction and functional annotation
The 190 target protein related to HCC were analyzed for finding the hub protein in the network. The most essential protein in the network calculated by degree of centrality were analyzed using CytoHubba plugin on Cytoscape. In cytoHubba, hub gene identification was performed using the MCC (Maximal Clique Centrality) scoring method, with the options ‘Check the first-stage nodes’, ‘Display the shortest path’, and ‘Display the expanded subnetwork’ enabled during analysis. The ranking of hub proteins was visualized through node size and color, where larger nodes and brighter colors indicate higher hub scores. The role of proteins in the network were analyzed using Functional Annotation in DAVID webserver (https://davidbioinformatics.nih.gov/). The libraries used in this analysis included Gene Ontology, KEGG Pathway, and WikiPathways. Only terms related to diabetes and those with a false discovery rate (FDR) of less than 0.05 were selected.
Molecular docking
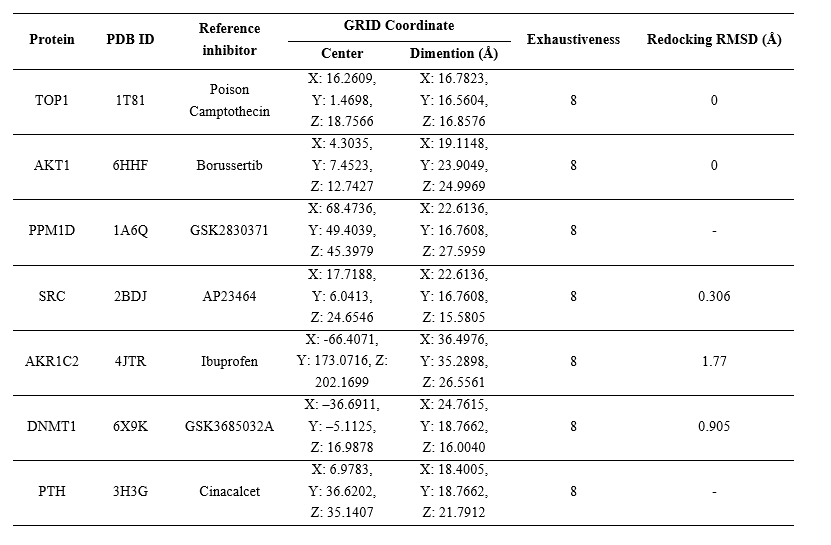
Molecular docking was conducted to assess the interactions between B. balsamifera active compounds and their target proteins, focusing on HCC-related and hub proteins identified from the network analysis. The three-dimensional (3D) structures of the active compounds were obtained from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). Target proteins, including TOP1, AKT1, PPM1D, SRC, AKR1C2, DNMT1, and PTH, were retrieved from the RCSB Protein Data Bank (https://www.rcsb.org/). The reference inhibitors were extracted from the respective protein crystallographic structures, except for PPM1D and PTH, for which the inhibitors GSK2830371 (CID: 70983932) and Cinacalcet (CID: 156419) were obtained from PubChem. The 3D structures of the active compounds were geometry-optimized using Open Babel [21], while all non-essential molecules such as water and co-crystallized ligands were removed from the protein structures using Biovia Discovery Studio 2021. Molecular docking simulations were carried out using AutoDock Vina integrated within the PyRx 0.9.5 interface to predict binding affinities and interaction poses between the ligands and target proteins [22]. The pose with the lowest (most negative) binding affinity value was selected as the optimal docking result. Validation was carried out using the re-docking method and the RMSD value was obtained using PyMOL 2.3.3 software. The molecular docking configuration were showed in Table 1.
Table 1 The details of the molecular docking method.
Molecular dynamic simulation
Molecular dynamics (MD) simulations were conducted for both protein–active compound and protein–inhibitor complexes using Yet Another Scientific Artificial Reality Application (YASARA) [23]. Each system was placed in a cubic simulation cell and solvated with explicit water molecules. The simulation environment was set to mimic human physiological conditions (pH 7.4, temperature 310 K, and salt concentration 0.9%). The AMBER14 force field was applied throughout the simulation. MD simulations were run for 20 nanoseconds, with trajectory data saved every 25 picoseconds. The simulation was executed using the md_run.mcr script. Structural stability was assessed by calculating the root mean square deviation (RMSD), radius of gyration (Rg), and number of hydrogen bond using md_analyze.mcr.
Results and discussion
The B. balsamifera extract toxic to HepG2
The cytotoxic effect of B. balsamifera extract on HepG2 cells is presented in Figure 1(A). Cell viability decreased significantly with increasing extract concentrations, demonstrating a clear dose-dependent pattern. A marked reduction in viability was observed at concentrations of 160 µg/mL and 320 µg/mL, indicating strong cytotoxic activity at higher doses. The calculated IC₅₀ value of the extract was 199.09 ± 11.86 µg/mL, suggesting that B. balsamifera extract exhibits moderately cytotoxic (IC50: 21 - 200 µg/mL) against HepG2 cells according to the classification by U.S. National Cancer Institute (NCI) [24].
This finding remains biologically meaningful because mild or moderate cytotoxicity in plant extracts often reflects selective rather than nonspecific toxicity [25]. Such profiles are advantageous in early-stage anticancer screening, as they suggest potential for targeting malignant cells while sparing normal tissues [26]. The decrease in HepG2 viability with increasing extract concentration may result from the combined action of flavonoids and phenolic compounds such as quercetin, which have been shown to induce oxidative stress, cell-cycle arrest, and apoptosis in hepatocellular carcinoma models [27,28]. Therefore, the cytotoxic response observed in this study provides preliminary evidence that B. balsamifera harbors bioactive constituents capable of inducing cell death and reducing viability in HCC cells, possibly through multiple molecular mechanisms.
The B. balsamifera extract induce apoptosis of HepG2 cell
Annexin V-FITC/PI flow cytometry showed a significant increase in early and late apoptotic HepG2 cells after treatment with B. balsamifera extract compared with the control and cisplatin groups (Figure 1(B)). The flow cytometry gating profiles (Figure 1(D)) further illustrated this effect, showing a pronounced increase in cell populations within the Annexin V–positive (early apoptotic) and Annexin V/PI–double positive (late apoptotic) quadrants, indicating that the extract effectively induces apoptosis in a dose-dependent manner. At the IC₅₀ concentration (199 µg/mL), a higher proportion of cells were in late apoptosis (56%), suggesting that most apoptotic cells had progressed beyond the early stage by the 24-hour time point. In contrast, at the higher concentration of 398 µg/mL, early apoptosis increased (51%) while late apoptosis decreased (34%), a pattern consistent with dose-dependent acceleration of apoptosis in which stronger cytotoxic stress triggers rapid initiation of apoptosis, resulting in more cells being captured in the early apoptotic phase. This apoptotic response was further supported by the rhodamine 123 mitochondrial assay, which showed a gradual decline in mitochondrial membrane potential as the extract concentration increased, a well-known indicator of apoptosis (Figures 1(C) - 1(E)).
The induction of apoptosis in HepG2 cells following B. balsamifera extract treatment indicates that the cytotoxic effect observed earlier is primarily mediated through programmed cell death rather than necrosis. The predominance of apoptosis over necrosis indicates that the cytotoxicity of B. balsamifera occurs primarily through programmed cell death mechanisms. Consistent with established pathways of phytochemical-induced apoptosis, this process is likely mediated by mitochondrial membrane potential disruption, cytochrome c release, and caspase activation [29]. These findings collectively suggest that B. balsamifera exerts its anticancer activity via mitochondria-mediated apoptosis, supporting its candidacy for further development as a natural therapeutic agent against hepatocellular carcinoma.
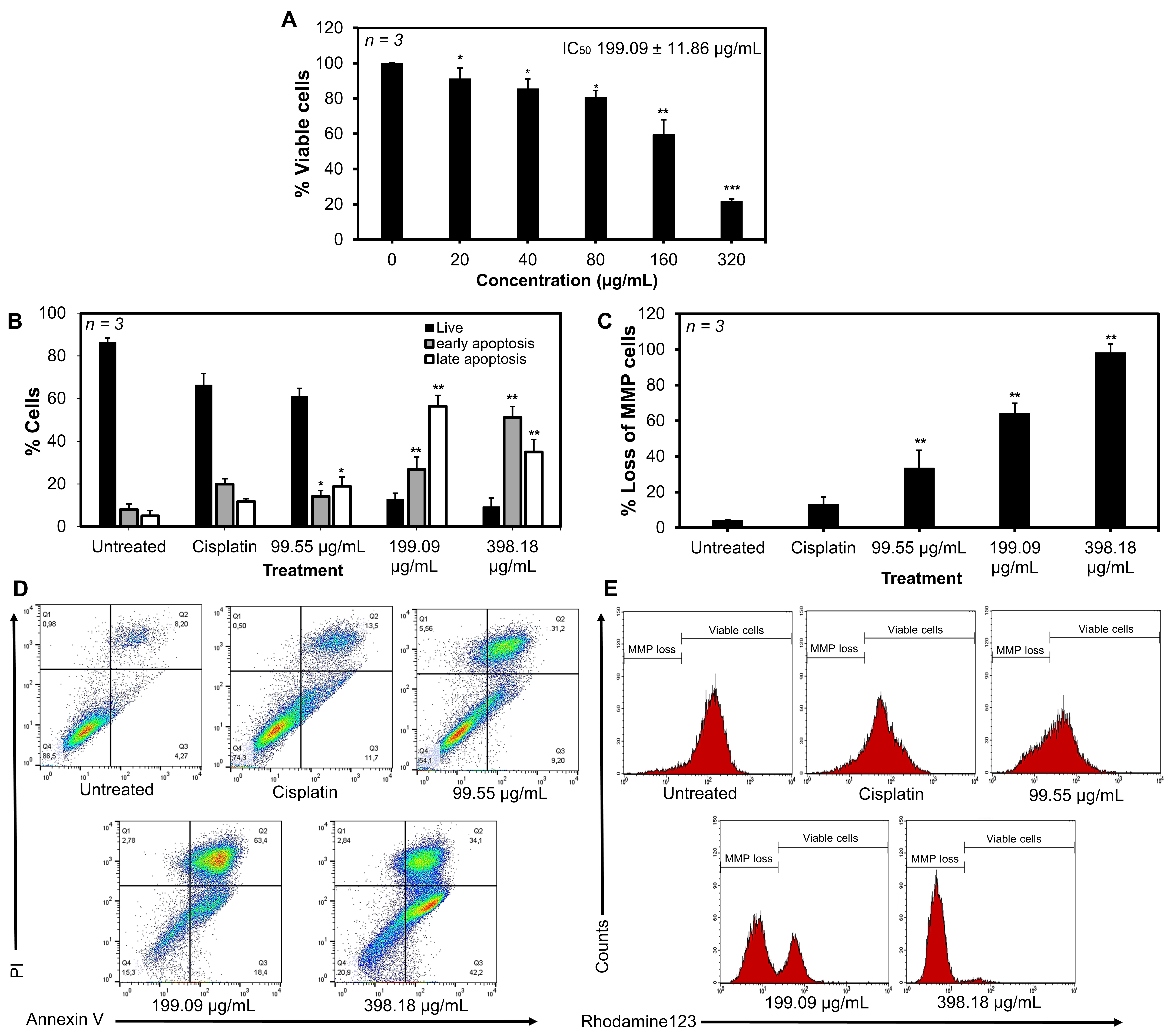
Figure 1 Cytotoxicity and apoptosis-inducing activity of the ethanolic extract of B. balsamifera in HepG2 cells. (A) B. balsamifera extract exhibited cytotoxic effects on HepG2 cells with an IC50 value of 199.09 ± 11.86 µg/mL. (B & D) The extract significantly induced apoptosis at concentrations of 199.09 and 398.18 µg/mL. (C & E) The extract induced a concentration-dependent loss of mitochondrial membrane potential (MMP) in HepG2 cells. For cytotoxicity assays, (*) indicates a significant difference compared with 0 µg/mL, (**) indicates a significant difference compared with 0 - 80 µg/mL, and (***) indicates a significant difference compared with all tested concentrations. For apoptosis and rhoamine123 assays, (*) indicates a significant difference compared with the untreated group, and (**) indicates a significant difference compared with both the untreated group and cisplatin. Data are presented as the mean of three replicates ± SD, with p-value < 0.01.
The B. balsamifera extract inhibit cell migration of HepG2 cell
The scratch assay revealed that treatment with B. balsamifera extract suppressed the migratory ability of HepG2 cells. In the control group, the scratch gap area progressively narrowed over time, reflecting the strong migratory and proliferative capacity of hepatocellular carcinoma cells. By contrast, in the extract-treated group, particularly at a concentration of 199 µg/mL, the gap area remained wide, indicating a significant reduction in cell migration rate (Figure 2). At the highest concentration, the wound area could not be clearly detected because most cells had undergone apoptosis, resulting in extensive cell loss within the culture.
The inhibition of HepG2 cell migration by B. balsamifera extract suggests an anti-metastatic potential, as cell migration is a critical step in tumor invasion and metastasis. This effect is consistent with previous findings that flavonoids such as quercetin and naringenin could suppress migration through downregulation of matrix metalloproteinase (MMP-2 and MMP-9) and inhibition of PI3K/AKT and MAPK signaling pathways [30]. The persistence of a wide gap area at higher concentrations highlights the ability of the extract to interfere with cytoskeletal remodeling and cellular motility. Taken together, these findings provide evidence that B. balsamifera may contribute not only to cytotoxic and pro-apoptotic effects but also to the suppression of metastatic progression in hepatocellular carcinoma.
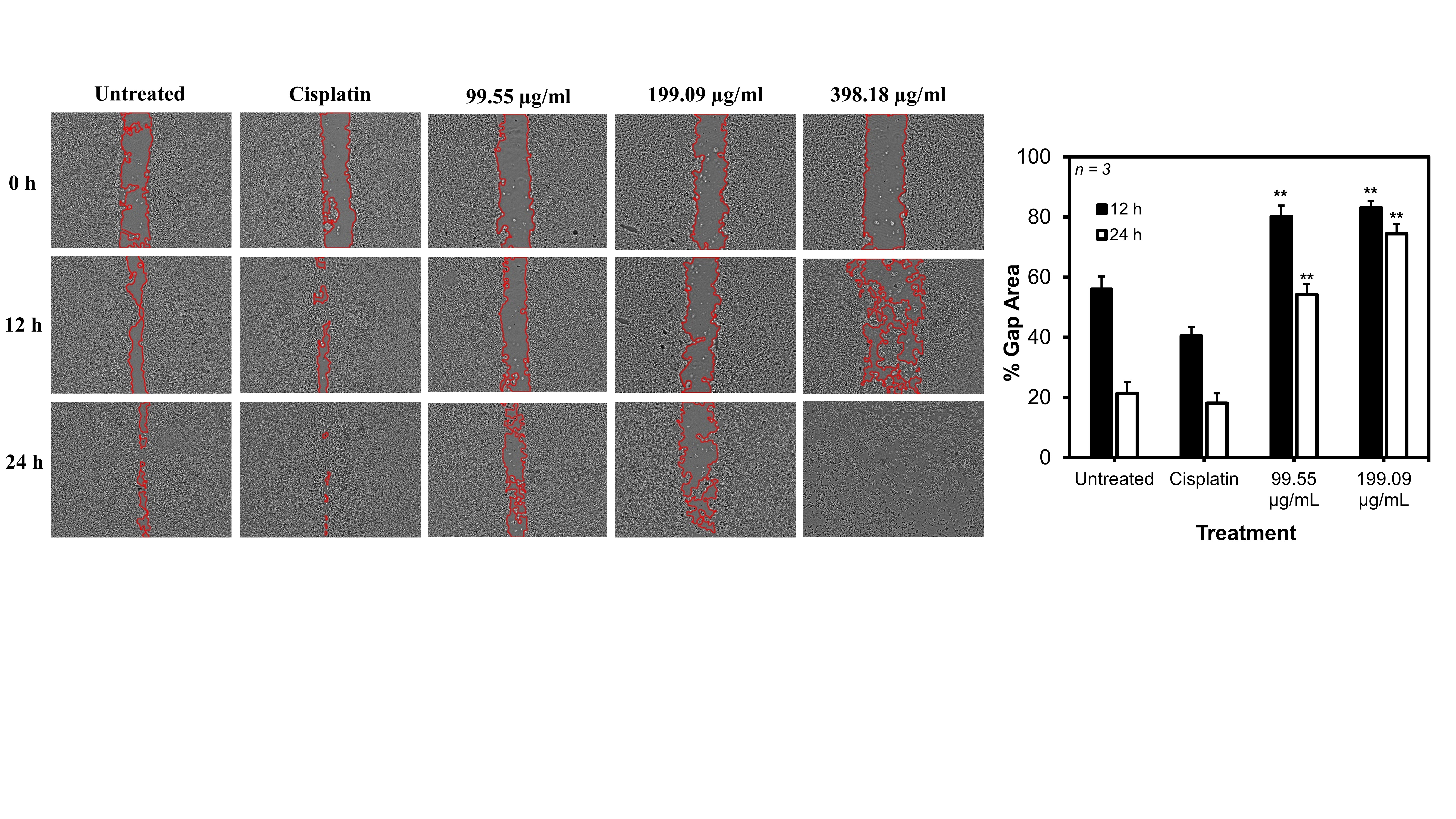
Figure 2 B. balsamifera extract inhibits HepG2 cell migration, as indicated by the wider gap area in the scratch assay. Data are presented as the mean of three replicates ± SD, with p-value < 0.01. (**) indicates a significant difference compared with both the untreated group and cisplatin.
Target proteins of active compounds in B. balsamifera extract
The STITCH analysis identified nine active compounds from B. balsamifera (quercetin, tuberonic acid, salicylate, isorhamnetin, 3,4-dicaffeoyl, lecanoric acid, chlorogenic acid, naringenin, and aurantio-obtus) that were predicted to interact with multiple protein targets (Figure 3(A)). Each compound exhibited multi-target potential, reflecting the complex pharmacology. Integration of these predicted targets with hepatocellular carcinoma (HCC)-related proteins obtained from the GeneCards database identified 190 overlapping proteins associated with both B. balsamifera compounds and HCC (Figure 3(B)). A compound–target network was subsequently constructed to visualize these interactions, revealing a highly interconnected topology in which several compounds were linked to multiple targets, while certain proteins functioned as hub nodes connecting multiple bioactive constituents (Figure 3(C)). These results suggest that B. balsamifera may exert its anticancer effects through multi-target modulation of key proteins involved in HCC-related signaling pathways.
The identification of 190 overlapping targets between B. balsamifera compounds and HCC-related proteins underscores the therapeutic potential of this plant in liver cancer. The multi-target nature of the interactions is consistent with previous reports that flavonoids such as quercetin and isorhamnetin can simultaneously regulate diverse signaling pathways implicated in tumorigenesis [10,11]. This property is advantageous in cancer therapy, as HCC involves a highly complex molecular landscape that often requires modulation of multiple pathways rather than single-target interventions.
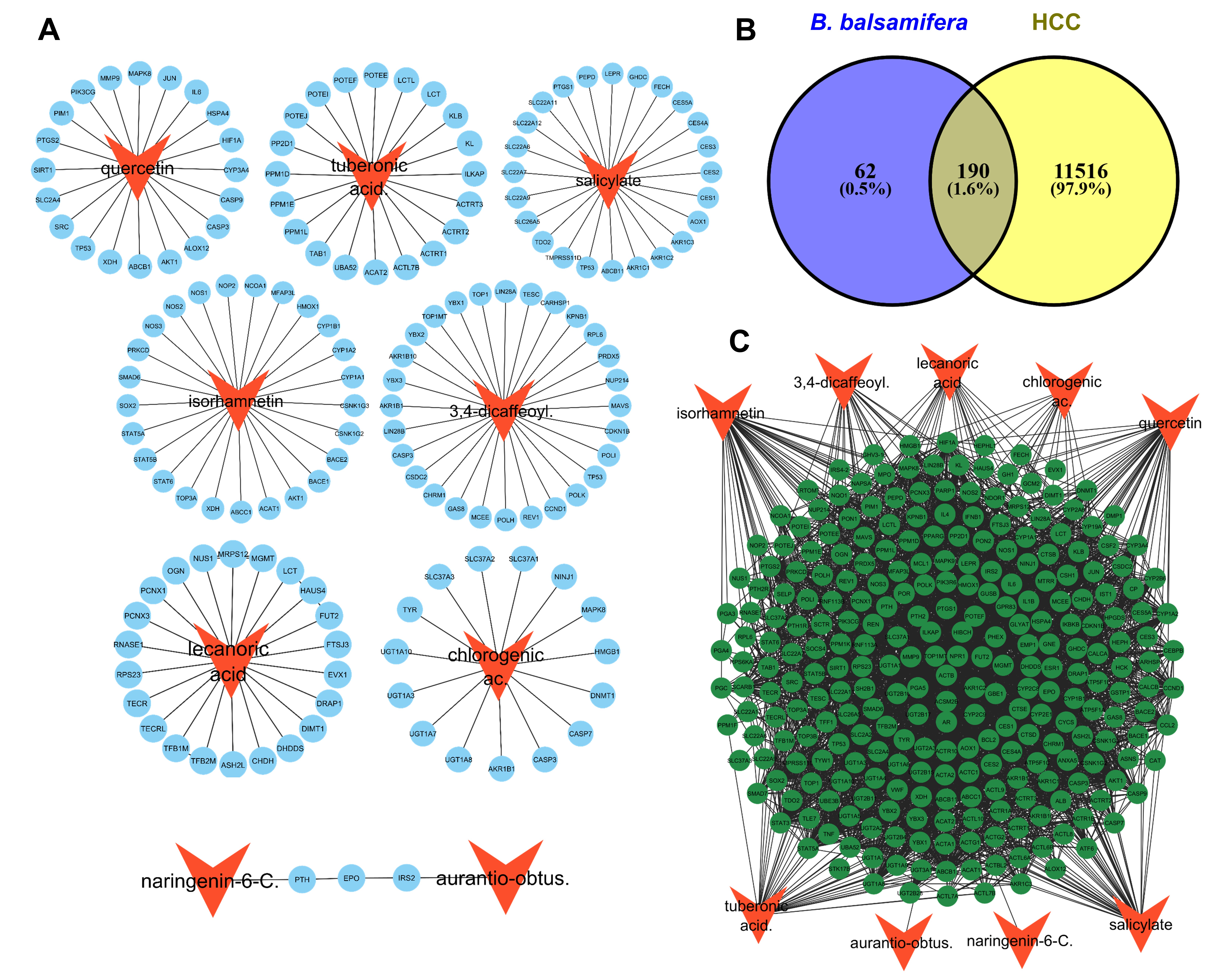
Figure 3 Network pharmacology representing hepatocellular carcinoma (HCC)-related targets of compounds contained in B. balsamifera extract. (A) Predicted protein targets of B. balsamifera compounds. (B) Overlap analysis identified 190 targets associated with HCC. (C) Complex interactions between active compounds and HCC-related protein targets.
Hub proteins and functional annotation indicated a strong correlation to apoptosis and cell migration
Network analysis identified several hub proteins, including members of the UDP-glucuronosyltransferase (UGT1) and cytochrome P450 (CYP) families, as well as key cancer-related regulators such as MAPK, TP53, and AKT1. The coexistence of metabolic enzymes and signaling molecules among these hub nodes suggests that B. balsamifera compounds may simultaneously modulate hepatic metabolism and cancer-associated signaling pathways (Figure 4(A)).
Functional annotation of the 190 overlapping target proteins using Gene Ontology (GO), KEGG, and WikiPathways databases revealed significant enrichment in processes related to apoptosis and cell migration. GO analysis highlighted biological processes such as apoptosis, cell migration, and cell motility (Figure 4(B)). KEGG pathway enrichment indicated strong associations with apoptosis-related pathways, including PI3K–AKT and p53 signaling (Figure 4(C)), while WikiPathways analysis similarly revealed enrichment in PI3K–AKT, apoptosis, and ErbB signaling pathways (Figure 4(D)).
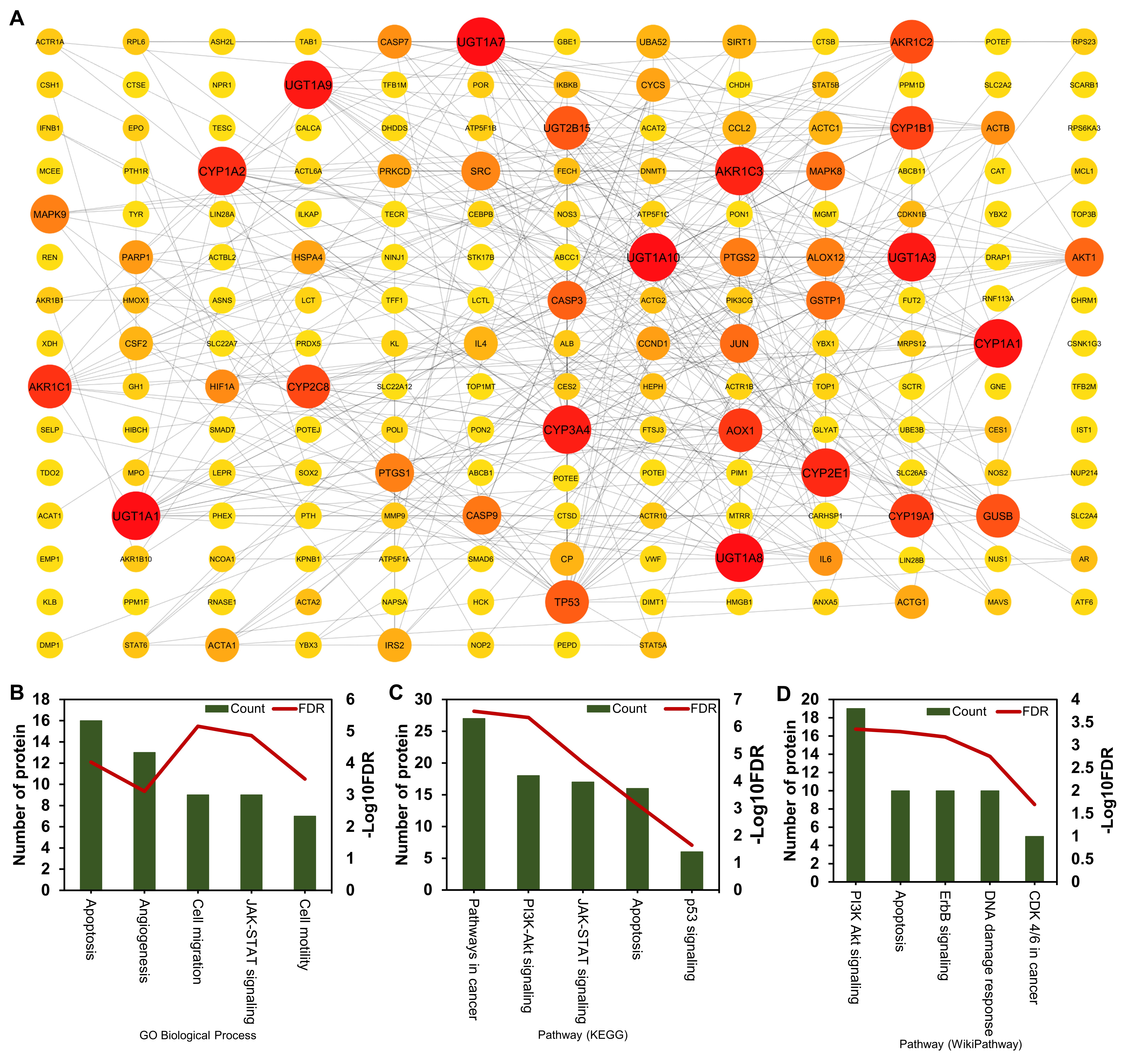
Figure 4 Hub proteins among the HCC-related targets identified using CytoHubba and functional annotation analysis. (A) Protein–protein interaction (PPI) network showing the hub proteins; larger node size and deeper red color indicate higher degree centrality. (B - D) Functional enrichment of the identified targets based on Gene Ontology (GO), KEGG, and WikiPathways analyses, highlighting their involvement in apoptosis and cell migration–related processes.
The compound–target network further highlights the importance of hub proteins with high connectivity, which likely play pivotal roles in mediating the biological effects of B. balsamifera. Previous studies have shown that such hub nodes in protein–protein interaction networks often represent essential disease-associated or druggable targets [31]. A similar approach has been applied in other network pharmacology studies, such as phytochemical investigations of Euphorbia hirta in breast cancer, where overlapping target prediction enhanced the understanding of therapeutic mechanisms [17]. Taken together, these findings provide a systems-level rationale that B. balsamifera may act through multi-target and network-based mechanisms to inhibit HCC progression. The identification of UGT1s and CYPs as hub proteins suggests that B. balsamifera may influence xenobiotic and drug metabolism, as well as the bio activation and detoxification of hepatic carcinogens [32]. Such interactions may alter hepatic metabolic capacity and potentially modulate cancer cell sensitivity to therapeutic agents.
Moreover, the emergence of MAPK, TP53, and AKT1 as hub proteins reinforces the link between B. balsamifera activity and classical mechanisms of hepatocarcinogenesis. Dysregulation of the MAPK pathway has been widely implicated in liver tumor progression through its role in proliferation, differentiation, and survival [33,34]. Similarly, TP53 is one of the most frequently mutated tumor suppressors in HCC, with loss of function driving uncontrolled proliferation and resistance to apoptosis [35]. Meanwhile, AKT1, as a central effector of the PI3K/AKT signaling pathway, is known to promote growth, angiogenesis, and apoptosis resistance in HCC [33]. Collectively, these findings suggest that B. balsamifera exerts its potential anticancer effects by targeting both metabolic enzymes and key oncogenic signaling nodes.
Interaction between active compounds and the most potential target
Target prediction using STITCH identified several potential protein targets for the active compounds of B. balsamifera. Subsequent screening using cBioPortal revealed one most promising target protein for each compound. The interactions between these active compounds and their respective targets were then evaluated through molecular docking. Docking simulations for TFB2M–lecanoric acid and IRS–aurantio-obtusin were excluded due to the absence of suitable reference inhibitors, which would compromise result validity.
The docking simulations showed that all other active compounds bound to their target proteins at the same binding site as the reference inhibitors. The binding affinity value was also not much different from the inhibitor (Table 2). Moreover, several active compounds interacted with the same key amino acid residues as the inhibitors, suggesting comparable binding modes. These findings indicate that the active compounds possess strong potential as inhibitors of TOP1, AKT1, PPM1D, SRC, AKR1C2, DNMT1, and PTH proteins (Figure 5).
Table 2 Binding affinity value of protein ligand interaction from molecular docking.
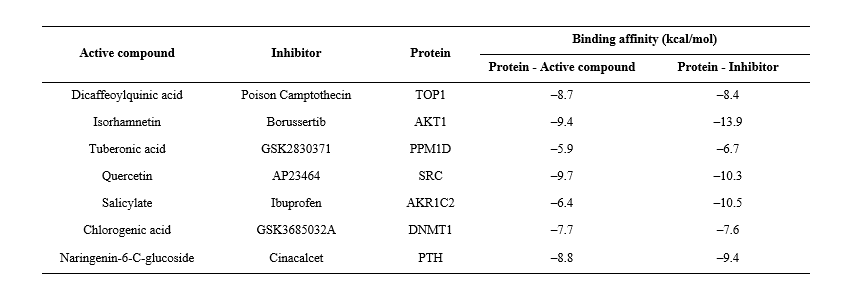
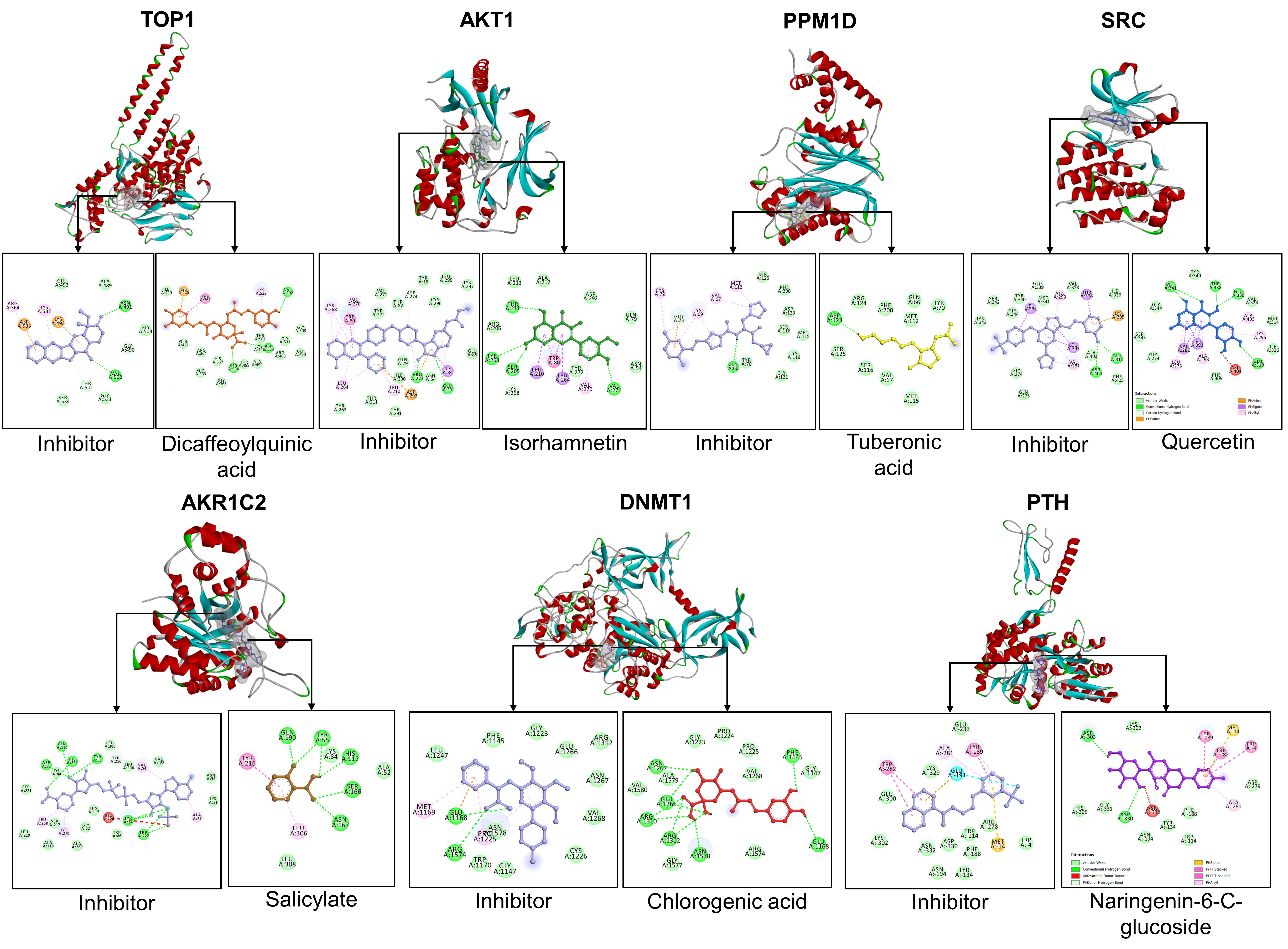
Figure 5 The molecular docking represents the interaction between active compound and its target protein.
Molecular Dynamic (MD) simulation of protein-active compound interaction
In this study, molecular dynamics (MD) simulations were performed to analyze the stability of protein structures and the stability of protein–active compound interactions over a 20 ns simulation period. The RMSD of the protein backbone represents structural stability based on atomic fluctuations within the backbone. The results showed that the backbone RMSD remained steady throughout the simulation, indicating that all proteins maintained stable structures (Figure 6(A)). This was supported by the radius of gyration (Rg), which did not increase during the simulation, confirming that the overall protein structures remained compact and stable (Figure 6(B)). The number of hydrogen bonds within the proteins also did not decrease, further demonstrating structural stability, as hydrogen bonds play an important role in maintaining secondary structure (Figure 6(C)).
The stability of protein–active compound interactions was represented by the RMSD of ligand movement and the number of hydrogen bonds formed between the protein and ligand. The RMSD values of ligand movement showed that all active compounds exhibited consistent trajectories, reflecting stable interactions with their target proteins (Figure 7(A)). The RMSD of ligand conformation also indicated that the structures of the active compounds remained stable throughout the simulation (Figure 7(B)). This interaction stability was further supported by the continuous formation of more than one hydrogen bond between the active compounds and their target proteins during the simulation (Figure 7(C)).
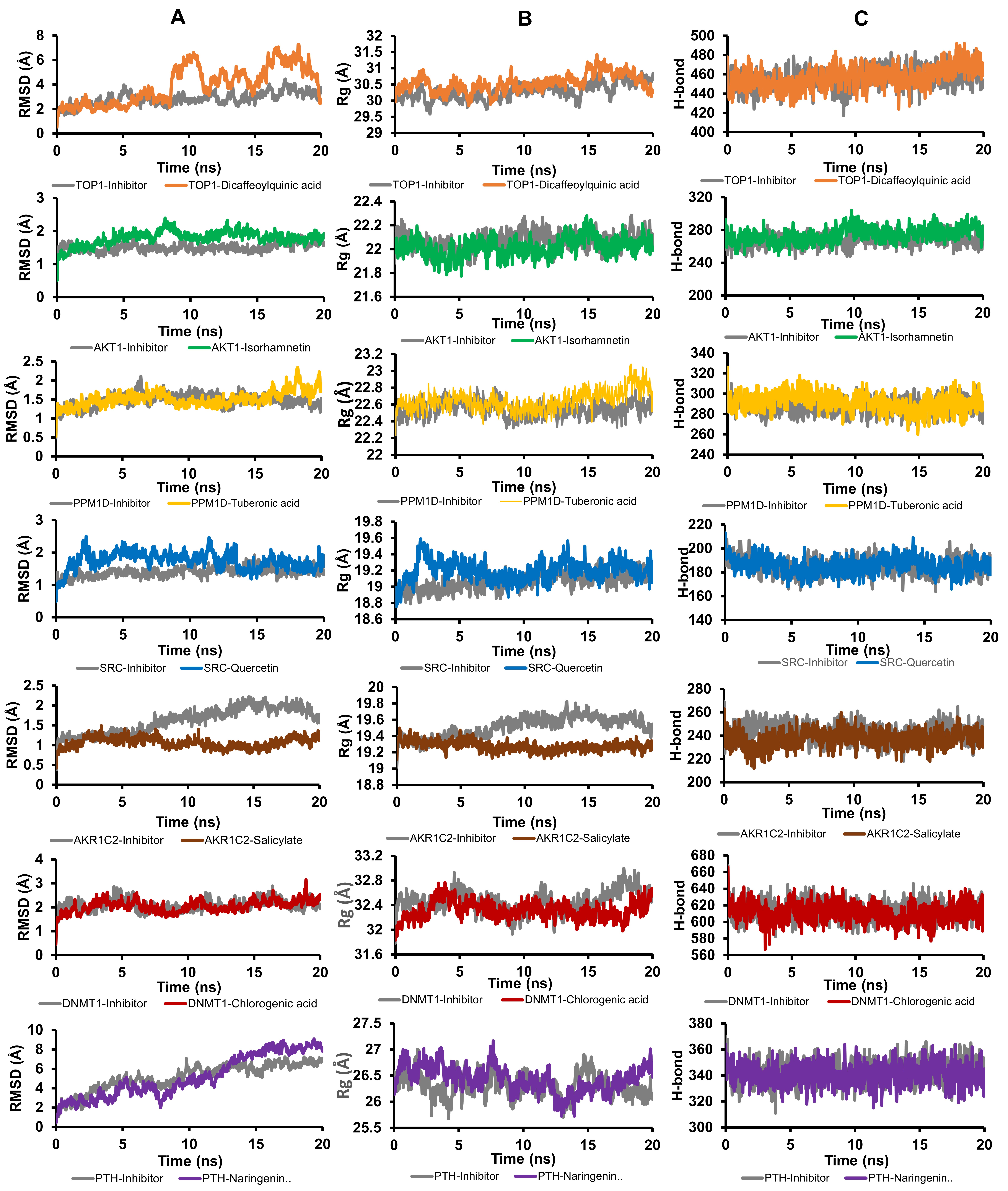
Figure 6 Molecular dynamic simulation represents the stability of protein structure during 20 ns simulation. A) Backbone protein RMSD, B) Radius of gyration (Rg). C) Number of hydrogen bond of protein.
The molecular dynamics simulation results provide strong evidence that the active compounds from B. balsamifera form stable and persistent interactions with their target proteins. The consistent RMSD values and sustained hydrogen bonding throughout the simulation indicate conformational stability of both the protein backbones and ligand-binding orientations [36,37]. These findings suggest that the docked complexes remain energetically favorable under physiological conditions, supporting the reliability of the docking predictions. Notably, the greater stability observed in the TOP1 and DNMT1 complexes compared to their reference inhibitors implies that the active compounds may exhibit comparable or even superior binding affinity and inhibitory potential. Similar patterns of ligand stability and hydrogen bond persistence have been correlated with enhanced binding strength and biological activity in previous simulation studies [38]. Collectively, these results reinforce the hypothesis that B. balsamifera constituents can act as multi-target inhibitors capable of modulating key oncogenic proteins in hepatocellular carcinoma.
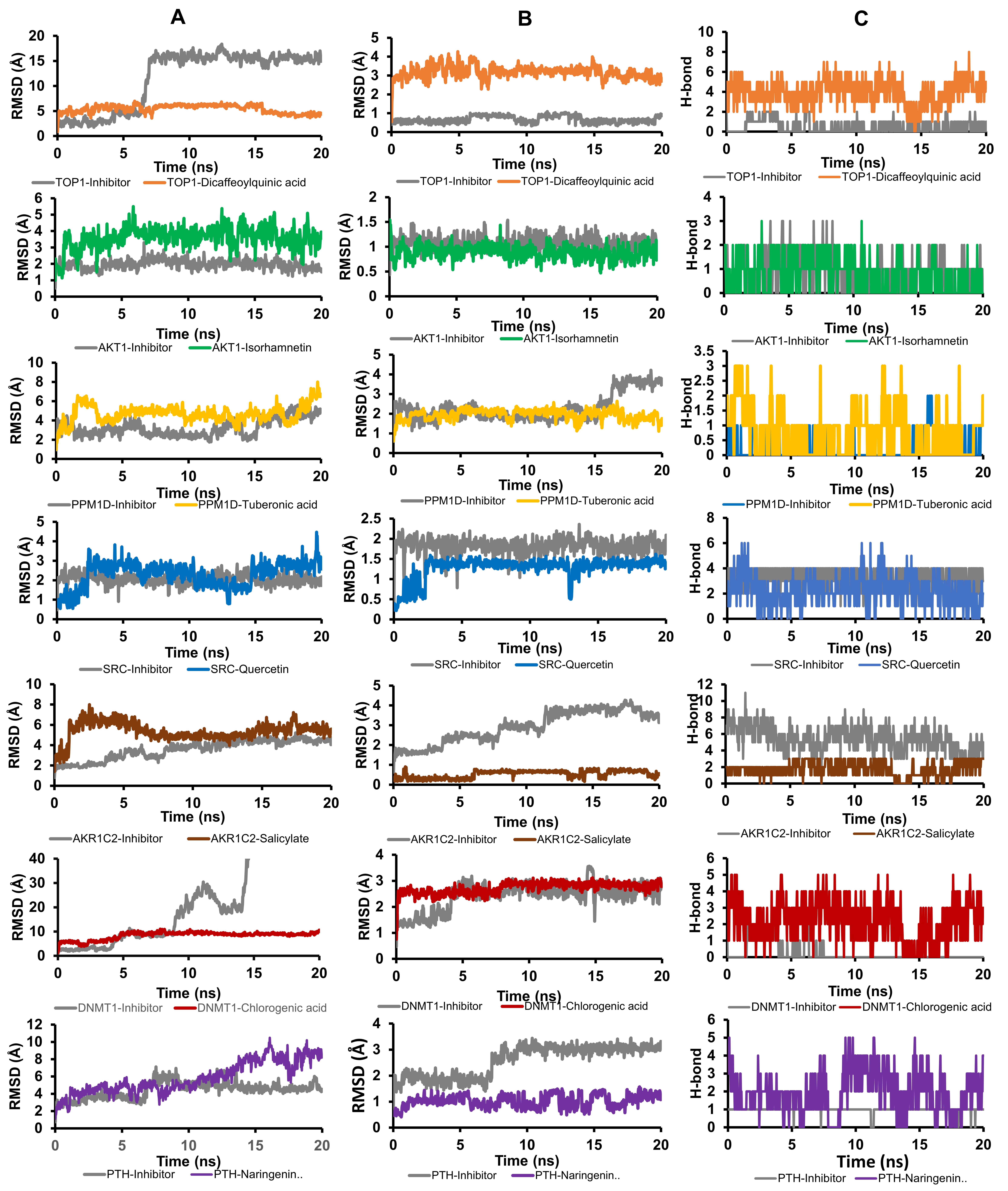
Figure 7 Molecular dynamic simulation represents the stability of protein-ligand interaction during 20 ns simulation. A) Ligand movement RMSD, B) RMSD of ligand conformation. C) Number of hydrogen bond in protein-ligand interaction.
Correlation of experimental and computational study
The molecular docking and dynamics analyses revealed that the active compounds of B. balsamifera could effectively interact with several key proteins involved in hepatocellular carcinoma progression. Dicaffeoylquinic acid exhibited stable binding to TOP1, a topoisomerase that regulates DNA supercoiling during replication and transcription. Inhibition of TOP1 disrupts DNA replication and induces DNA strand breaks, triggering apoptosis in rapidly dividing cancer cells [39]. Similarly, chlorogenic acid showed strong affinity toward DNMT1, a DNA methyltransferase responsible for maintaining aberrant DNA methylation patterns in tumors. Suppression of DNMT1 can lead to reactivation of silenced tumor suppressor genes and initiation of apoptotic pathways [40]. These interactions suggest that B. balsamifera constituents may exert their cytotoxic and pro-apoptotic effects by targeting nuclear enzymes essential for DNA integrity and epigenetic regulation.
Another crucial target identified was AKT1, which plays a central role in the PI3K–AKT signaling pathway that promotes cell survival and migration. The strong binding of isorhamnetin to AKT1 indicates potential inhibition of downstream phosphorylation cascades that suppress apoptosis and stimulate motility [33]. Likewise, tuberonic acid was predicted to inhibit PPM1D, a phosphatase that negatively regulates p53-mediated apoptosis. Inhibition of PPM1D can stabilize p53, enhancing apoptotic signaling and reducing the proliferation of malignant hepatocytes [41]. The combined inhibition of AKT1 and PPM1D by B. balsamifera compounds may therefore contribute to both apoptosis induction and inhibition of cell migration, consistent with the experimental findings from flow cytometry and scratch assays.
Additionally, quercetin and salicylate exhibited high affinity for SRC and AKR1C2, respectively, while naringenin-6-C-glucoside targeted PTH. SRC is a well-known proto-oncogene that drives cytoskeletal rearrangements and epithelial–mesenchymal transition (EMT), processes that enhance cell migration and invasion [42]. Previous study reported that Quercetin inhibit metastasis and invasion of HCC in vitro [43]. Similarly, inhibition of AKR1C2, an aldo–keto reductase involved in detoxification and hormone metabolism, can attenuate sensitize cancer cells to apoptosis [44]. Meanwhile, modulation of PTH signaling by naringenin derivatives may contribute to calcium-dependent apoptotic regulation in hepatocytes [45]. Collectively, these results suggest that B. balsamifera exerts multi-target inhibitory effects on proteins associated with cell survival, DNA repair, and migration, thereby promoting apoptosis and suppressing metastatic behavior in hepatocellular carcinoma cells.
Although this study demonstrates the therapeutic potential of B. balsamifera against hepatocellular carcinoma, further investigations are necessary. The next study using a normal hepatic cell line is needed to examine the selectivity of B. balsamifera extract. Migration assays under proliferation-blocked conditions need to be performed to allow a clearer interpretation of whether changes in wound closure are due to reduced motility or decreased proliferative activity. Future work should include in vivo validation to confirm the extract’s efficacy, pharmacokinetics, and systemic safety. Advanced transcriptomic and proteomic analyses would also help elucidate the precise molecular pathways modulated by B. balsamifera in hepatic cancer cells.
Conclusions
This study provides both experimental and computational evidence supporting the potential of B. balsamifera as a promising natural product–based therapeutic candidate for hepatocellular carcinoma (HCC). The extract exhibited dose-dependent cytotoxic and pro-apoptotic effects on HepG2 cells, accompanied by significant inhibition of cell migration. Network pharmacology, molecular docking, and molecular dynamics simulations collectively revealed that the active compounds of B. balsamifera interact stably with multiple HCC-related targets—including TOP1, AKT1, PPM1D, SRC, AKR1C2, DNMT1, and PTH—which are known to regulate apoptosis, cell survival, and migration pathways. These multi-target interactions underscore the polypharmacological nature of B. balsamifera and its potential to modulate key molecular mechanisms underlying cancer progression. Overall, the findings highlight B. balsamifera as a valuable source of bioactive compounds for further development in anti-HCC therapy.
Acknowledgement
This research was funded by the Regular Fundamental Research Program of the Ministry of Education, Culture, Research, and Technology (Kemendikbudristek) of the Republic of Indonesia (grant: 628/UN10.A0501/B/PT.01.03.2/2025). We gratefully acknowledge the AI-Center, Universitas Brawijaya, for providing computational resources for molecular dynamics analyses. We also thank the Integrated Research Laboratory (LRT), Universitas Brawijaya, for providing facilities for the extraction of B. balsamifera.
Declaration of generative AI in scientific writing
The authors declare that generative artificial intelligence (AI) tools were used to assist in the preparation of this manuscript. Specifically, ChatGPT (OpenAI) and Grammarly were employed for grammar correction and readability enhancement. No AI tools were used for data analysis, interpretation of results, or drawing scientific conclusions. The final manuscript reflects the authors’ own intellectual contributions and scientific judgment.
CRediT author statement
Muhammad Hermawan Widyananda: Conceptualization, Methodology, Data Curation, Validation, Writing - Original Draft. Adinda Romdiatus Sa’adah: Methodology, Formal Analysis, Data Curation, Investigation, Writing - Original Manuscript. Yuslinda Annisa: Validation, Investigation, Supervision. Nuraini Rosyadah: Supervision, Project Administration. Dinia Rizqi Dwijayanti: Methodology, Supervision. Feri Eko Hermanto: Writing - Review & Editing, Supervision. Nashi Widodo: Conceptualization, Methodology, Software, Validation, Resource, Supervision, Funding Acquisition.
References
[1] F Bray, M Laversanne, H Sung, J Ferlay, RL Siegel, I Soerjomataram and A Jemal. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians 2024; 74(3), 229-263.
[2] YC Shen, HC Hsu, TM Lin, YS Chang, LF Hu, LF Chen, SH Lin, PI Kuo, WS Chen, YC Lin, JH. Chen, YC Liang and CC Chang. H1-Antihistamines reduce the risk of hepatocellular carcinoma in patients with hepatitis B virus, hepatitis C virus, or dual hepatitis B virus-hepatitis C virus infection. Journal of Clinical Oncology 2022; 40(11), 1206-1219.
[3] J Balogh, D Victor, EH Asham, SG Burroughs, M Boktour, A Saharia, X Li, RM Ghobrial and HP Monsour. Hepatocellular carcinoma: A review. Journal of Hepatocellular Carcinoma 2016; 3, 41-53.
[4] D Papaconstantinou, DI Tsilimigras and TM Pawlik. Recurrent hepatocellular carcinoma: Patterns, detection, staging and treatment. Journal of Hepatocellular Carcinoma 2022; 9, 947-957.
[5] J Zheng, S Wang, L Xia, Z Sun, KM Chan, R Bernards, W Qin, J Chen, Q Xia and H Jin. Hepatocellular carcinoma: Signaling pathways and therapeutic advances. Signal Transduction and Targeted Therapy 2025; 10(1), 35.
[6] Z Kan, H Zheng, X Liu, S Li, TD Barber, Z Gong, H Gao, K Hao, MD Willard, J Xu, R Hauptschein, PA Rejto, J Fernandez, G Wang, Q Zhang, B Wang, R Chen, J Wang, NP Lee, W Zhou, Z Lin, Z Peng, K Yi, S Chen, L Li, X Fan, J Yang, R Ye, J Ju, K Wang, H Estrella, S Deng, P Wei, M Qiu, IH Wulur, J Liu, ME Ehsani, C Zhang, A Loboda, WK Sung, A Aggarwal, RT Poon, ST Fan, J Wang, J Hardwick, C Reinhard, H Dai, Y Li, JM Luk and M Mao. Whole-genome sequencing identifies recurrent mutations in hepatocellular carcinoma. Genome Research 2013; 23(9), 1422-1433.
[7] A Fernandez-Medarde and E Santos. Ras in Cancer and Developmental Diseases. Genes & Cancer 2011; 2(3), 344-358.
[8] AJ Gosalia, P Martin and PD Jones. Advances and future directions in the treatment of hepatocellular carcinoma, Gastroenterology & Hepatology 2017; 13(7), 398-410.
[9] IG Widhiantara and IM Jawi. Phytochemical composition and health properties of Sembung plant (Blumea balsamifera): A review. Veterinary World 2021; 14(5), 1185-1196.
[10] A Schieber, P Keller, P Streker, I.Klaiber and R Carle. Detection of isorhamnetin glycosides in extracts of apples (Malus domestica cv. “Brettacher”) by HPLC‐PDA and HPLC‐APCI‐MS/MS. Phytochemical Analysis 2002; 13(2), 87-94.
[11] W Wang, X Yuan, J Mu, Y Zou, L Xu, J Chen, X Zhu, B Li, Z Zeng, X Wu, Z Yin and Q Wang. Quercetin induces MGMT+ glioblastoma cells apoptosis via dual inhibition of Wnt3a/β-Catenin and Akt/NF-κB signaling pathways. Phytomedicine 2023; 118, 154933.
[12] NLPDPP Sari, NLPEK Sari, PNC Santoso, ES Dewi, DPO Lestari, A Lestarini and NW Armerinayanti. Anticancer Activity Test of 70% Alcohol Extract of Sembung Leaves (Blumea balsamifera) Against Cervical Cancer Cells In-Vitro. BIO Web Conference 2025; 184, 01011.
[13] NA Ismail, A Matawali, PC Lee and J Gansau. Blumea balsamifera (L.) DC. Elicit Anti-Kinase, Anti-Phosphatase and Cytotoxic Activities against Acute Promyelocytic Leukemia Cells (HL-60). Tropical Journal of Natural Product Research 2021; 5(4), 656-660.
[14] Y Pang, D Wang, Z Fan, X Chen, F Yu, X Hu, K Wang and L Yuan. Blumea balsamifera—A Phytochemical and Pharmacological Review. Molecules 2014; 19(7), 9453-9477.
[15] T Norikura, A Kojima-Yuasa, M Shimizu, X Huang, S Xu, S Kametani, SN Rho, DO Kennedy and I Matsui-Yuasa. Mechanism of growth inhibitory effect of Blumea balsamiferaExtract in hepatocellular carcinoma. Bioscience, Biotechnology, and Biochemistry 2008; 72(5), 1183-1189.
[16] MH Widyananda, ST Wicaksono, K Rahmawati, S Puspitarini, SM Ulfa, YD Jatmiko, M Masruri and N Widodo. A Potential Anticancer Mechanism of Finger Root (Boesenbergia rotunda) Extracts against a Breast Cancer Cell Line. Scientifica 2022; 2022, 9130252.
[17] MH Widyananda, L Muflikhah, SM Ulfa and N Widodo. Unveiling the antibreast cancer mechanism of Euphorbia hirta ethanol extract: Computational and experimental study. Journal of Biologically Active Products from Nature 2024; 14(3), 359-382.
[18] N Widodo, S Puspitarini, MH Widyananda, A Alamsyah, ST Wicaksono, M Masruri and YD Jatmiko. Anticancer activity of Caesalpinia sappan by downregulating mitochondrial genes in A549 lung cancer cell line [version 2; peer review: 2 approved]. F1000Research 2022; 11, 169.
[19] FSK amila, Y Annisa, N Rosyadah, FE Hermanto, MH Widyananda, DR Dwijayanti and N Widodo. Evaluation of polyphenol and antioxidant properties of Blumea balsamifera extract as potential therapeutic for breast cancer. BIO Web of Conferences 2025; 154, 03004.
[20] M Kuhn, D Szklarczyk, S Pletscher-Frankild, TH Blicher, C Von Mering, LJ Jensen and P Bork. STITCH 4: Integration of protein–chemical interactions with user data. Nucleic Acids Research 2014; 42(D1), D401-D407.
[21] NM O’Boyle, M Banck, CA James, C Morley, T Vandermeersch and GR Hutchison. Open Babel: An open chemical toolbox. Journal of Cheminformatics 2011; 3(1), 33.
[22] S Dallakyan and AJ Olson. Small-Molecule Library Screening by Docking with PyRx. In: JE. Hempel, CH Williams and CC Hong (Eds.). Chemical biology, Springer, New York, 2015, pp. 243-250.
[23] E Krieger and G Vriend. New ways to boost molecular dynamics simulations. Journal of Computational Chemistry 2015; 36(13), 996-1007.
[24] J Yammine, A Fathima, A Gharsallaoui, MZ Masalmeh, M Hasan, HC Yalcin, L Karam and AA Shaito Nanoencapsulation mitigates the toxicity of thymol in human cells and zebrafish embryos. International Journal of Food Properties 2018; 28(1), 2573183
[25] V Shrihastini, P Muthuramalingam, S Adarshan, M Sujitha, JT Chen and H Shin. M. Ramesh, plant derived bioactive compounds, their anti-cancer effects and In Silico approaches as an alternative target treatment strategy for breast cancer: An updated overview. Cancers 2021; 13(24), 6222.
[26] D Basak, S Arrighi, Y Darwiche and S Deb. Comparison of anticancer drug toxicities: Paradigm shift in adverse effect profile. Life 2021; 12(1), 48.
[27] P Biswas, D Dey, PK Biswas, TI Rahaman, S Saha, A Parvez, DA Khan, NJ Lily, K Saha, M Sohel, MM Hasan, S Al Azad, S Bibi, MDN Hasan, M Rahmatullah, J Chun, MDA Rahman and B Kim. A comprehensive analysis and anti-cancer activities of quercetin in ros-mediated cancer and cancer stem cells. International Journal of Molecular Sciences 2022; 23(19), 11746.
[28] MH Widyananda, SK Pratama, ANM Ansori, Y Antonius, VD Kharisma, AAA Murtadlo, V Jakhmola, M Rebezov, M Khayrullin, M Derkho, E Ullah, RJK Susilo, S Hayaza, AP Nugraha, A Proboningrat, A Fadholly, MT Sibero and R Zainul. Quercetin as an anticancer candidate for glioblastoma multiforme by targeting AKT1, MMP9, ABCB1, and VEGFA: An in silico study. Karbala International Journal of Modern Science 2023; 9(3), 450-459.
[29] E Gottlieb, SM Armour, MH Harris and CB Thompson. Mitochondrial membrane potential regulates matrix configuration and cytochrome c release during apoptosis. Cell Death & Differentiation 2003; 10(6), 709-717.
[30] A Jabłońska-Trypuć, M Matejczyk and S Rosochacki. Matrix metalloproteinases (MMPs), the main extracellular matrix (ECM) enzymes in collagen degradation, as a target for anticancer drugs. Journal of Enzyme Inhibition and Medicinal Chemistry 2016; 31(1), 177-183.
[31] SW Marseti, FE Hermanto, MH Widyananda, N Rosyadah, FS Kamila, Y Annisa, DR Dwijayanti, SM Ulfa and N Widodo. Pharmacological potential of Clinacanthus nutans: Integrating network pharmacology with experimental studies against lung cancer. Journal of Biologically Active Products from Nature 2024; 14(3), 343-358.
[32] KM Knights, A Rowland and JO Miners. Renal drug metabolism in humans: The potential for drug-endobiotic interactions involving cytochrome P450 (CYP) and UDP ‐glucuronosyltransferase (UGT). British Journal of Clinical Pharmacology 2013; 76(4), 587-602.
[33] S Deng, HC Leong, A Datta, V Gopal, AP Kumar and CT Yap. PI3K/AKT signaling tips the balance of cytoskeletal forces for cancer progression. Cancers 2022; 14(7), 1652.
[34] Y Hayashi, Y Toyomasu, SA Saravanaperumal, MR Bardsley, JA Smestad, A Lorincz, ST Eisenman, G Cipriani, MH Nelson Holte, FJ Al Khazal, SA Syed, GB Gajdos, KM Choi, GJ Stoltz, KE Miller, ML Kendrick, BP Rubin, SJ Gibbons, AE Bharucha, DR Linden, LJ Maher, G Farrugia and T Ordog. Hyperglycemia increases interstitial cells of cajal via MAPK1 and MAPK3 signaling to etv1 and kit, leading to rapid gastric emptying. Gastroenterology 2017; 153(2), 521-535.
[35] F Wang, H Zhang, H Wang, T Qiu, B He and Q Yang. Combination of AURKA inhibitor and HSP90 inhibitor to treat breast cancer with AURKA overexpression and TP53 mutations. Medical Oncology 2022; 39(12), 180.
[36] K Sargsyan, C Grauffel and C Lim. How molecular size impacts RMSD applications in molecular dynamics simulations. Journal of Chemical Theory and Computation 2017; 13(4), 1518-1524.
[37] AMD Fonseca, BJ Caluaco, JMC Madureira, SQ Cabongo, EM Gaieta, F Djata, RP Colares, MM Neto, CFC Fernandes, GS Marinho, HSD Santos and ES Marinho. Screening of potential inhibitors targeting the main protease structure of SARS-CoV-2 via molecular docking, and approach with molecular dynamics, RMSD, RMSF, H-Bond, SASA and MMGBSA. Molecular Biotechnology 2023; 66, 1919-1933.
[38] D Chen, N Oezguen, P Urvil, C Ferguson, SM Dann and TC Savidge. Regulation of protein-ligand binding affinity by hydrogen bond pairing. Science Advances 2016; 2(3), e1501240.
[39] MM Madkour, WS Ramadan, E Saleh and R El-Awady. Epigenetic modulations in cancer: Predictive biomarkers and potential targets for overcoming the resistance to topoisomerase I inhibitors. Annals of Medicine 2023; 55(1), 2203946.
[40] T Chen, S Mahdadi, M Vidal and S Desbène-Finck. Non-nucleoside inhibitors of DNMT1 and DNMT3 for targeted cancer therapy. Pharmacological Research 2024; 207, 107328.
[41] Z Andrysik, KD Sullivan, JS Kieft and JM Espinosa. PPM1D suppresses p53-dependent transactivation and cell death by inhibiting the Integrated Stress Response. Nature Communication 2022, 13(1), 7400.
[42] S Noshita, Y Kubo, K Kajiwara, D Okuzaki, S Nada and M Okada. A TGF-β-responsive enhancer regulates SRC expression and epithelial–mesenchymal transition-associated cell migration. Journal of Cell Science 2023; 136(15), jcs261001.
[43] CFL Gonçalves, F Hecht, J Cazarin, RS Fortunato, M Vaisman, DPD Carvalho and ACF Ferreira. The flavonoid quercetin reduces cell migration and increases NIS and E-cadherin mRNA in the human thyroid cancer cell line BCPAP. Molecular and Cellular Endocrinology 2021; 529, 111266.
[44] XZ Xiao, LY Lin, MK Zhuang, CM Zhong and FL Chen. Roles of AKR1C3 in malignancy. Chinese Medical Journal 2021; 134(9), 1052-1054.
[45] IWY Mak, RW Cowan, RE Turcotte, G Singh and M Ghert. pthrp induces autocrine/paracrine proliferation of bone tumor cells through inhibition of apoptosis. PLoS ONE 2011; 6(5), e19975.