
Trends
Sci.
2026;
23(6):
12463
New Onoceranoid-Triterpenoids from the Stem Bark of Lansium domesticum Corr. cv. Piedjietan: Structural Characterization and Their Cytotoxic Activity Evaluated by In Vitro and In Silico Approaches
Rika Septiyanti1, Selvi Apriliana Putri1, Dilla Mardyana1,
Iman Permana Maksum1, Sofa Fajriah3, Mohamad Azlan Nafiah2,
Kindi
Farabi1,
Rani Maharani1
and Tri Mayanti1,4,*,
![]()
1Department of Chemistry, Faculty of Mathematics and Natural Sciences, Universitas Padjadjaran,
Jatinangor 45363, Indonesia
2Department of Chemistry, Faculty of Sciences and Mathematic, Sultan Idris Education University,
Perak 35900, Malaysia
3Research Center for Pharmaceutical Ingredients and Traditional Medicine, National Research and Innovation Agency (BRIN), Cibinong Science Center Complex - BRIN, Cibinong 16911 Indonesia
4PKR Universitas Padjadjaran-BRIN: Drug Discovery from Indonesian Meliaceae Plants, Nanomaterial Microbes, West Java 45363, Indonesia
(*Corresponding author: e-mail: [email protected])
Received: 29 October 2025, Revised: 4 December 2025, Accepted: 11 December 2025, Published: 20 February 2026
Abstract
This study presents the isolation, characterization and cytotoxic potential of four onoceranoid triterpenoids from the stem bark of Lansium domesticum Corr. cv. Piedjietan, including a newly identified compound, methyl lansic (1), and three known compounds: α,γ-onoceradienedione (2), methyl ester lansiolate (3), and lansiolic acid (4), but reported for the first time in this cultivar. Methyl lansic was tested for cytotoxicity against MCF-7 breast cancer cells. The Results showed that methyl lansic exhibited strong cytotoxic activity with an IC₅₀ value of 7.78 µg/mL, while its IC₅₀ on normal CV-1 cells was substantially higher (620.30 µg/mL), indicating good selectivity toward cancer cells. Molecular docking analysis indicated that methyl lansic and α,γ-onoceradienedione had strong binding affinities to ERα, suggesting their potential as estrogen receptor inhibitors. Molecular dynamics, RMSD/RMSF profiles, persistent key interactions, and favourable MM/GBSA binding energy collectively demonstrate that the methyl lansic–ERα complex is structurally stable and exhibits characteristics consistent with a promising candidate for further evaluation in ERα-positive breast cancer models. These findings expand the phytochemical and pharmacological understanding of Lansium domesticum cv. Piedjietan and highlight its promise as a source of bioactive compounds for the development of anticancer agents.
Keywords: Onoceranoid-triterpenoid, Lansium domesticum Corr. cv. Piedjietan, In silico, In vitro, Breast cancer, MCF-7
Introduction
The family Meliaceae has been reported to consist of 52 genera and more than 1.400 species, with one of its genera being Lansium [1]. According to Hasskarl (1844), there are three cultivars of Lansium domesticum Corr. found in Indonesia: Duku (duku Hasskl), kokosan
(kokossan Hasskl), and pisitan (piedjietan Hasskl) [2,3]. Lansium genus is also known for its rich secondary metabolite content, particularly from stem bark [4]. Triterpenoids in Lansium genus have been reported to exhibit a wide range of bioactivities [5], such as antifeedant [6,7], antioxidant [8], antidiabetic [9], anticancer [10-12], antibacterial [13] and antimutagenic [14].
This study is particularly significant as no previous reports have described the isolation of compounds from Lansium domesticum Corr. cultivar piedjietan. Considering that onoceranoid triterpenoids are well known for a wide range of biological activities, especially their strong cytotoxic effects, the absence of any data on the presence or activity of such compounds in this species. Moreover, the structural characteristics of onoceranoid triterpenoids, particularly the presence of oxygenated carbon atoms at C-3 and C-21, provide functional groups capable of forming hydrogen bonds with key residues within the estrogen receptor alpha binding site. In addition, their predominantly hydrophobic skeleton facilitates favorable interactions within the receptor’s hydrophobic pocket, thereby enhancing their potential as modulators or inhibitors of estrogen receptor activity. Therefore, this research aims to identify the secondary metabolites contained within this plant, with particular attention to potential onoceranoid structures, and to evaluate their cytotoxic properties through comprehensive in silico and in vitro studies. In silico analysis was performed through molecular docking between the isolated compounds and the estrogen alpha receptor, as the estrogen alpha receptor is commonly involved in breast cancer cases [15,16]. Furthermore, drug-likeness was assessed to determine the suitability of the compounds for development into drugs, while the Rule of Five was applied to evaluate their oral bioavailability potential. The Rule of Five was used to test whether the molecular properties of the compounds meet the criteria for favorable absorption and permeability, which are essential factors for drug development.
In vitro testing was conducted to evaluate the toxicity of the compounds against MCF-7 cells dan normal cells (CV-1). The MCF-7 cell line was selected because it is widely used as a model for studying estrogen receptor-positive breast cancer and is suitable for evaluating the effects of compounds on breast cancer cells [17]. From the stem bark of the Piedjietan cultivar, we have successfully isolated four onoceranoid triterpenoids; methyl lansic (1), α, γ-onoceradienedione (2), and methyl ester lansiolate (3) were isolated from the n-hexane extract, whereas lansiolic acid (4) was isolated from the ethyl acetate extract. Notably, methyl lansic (1) exhibited significant cytotoxic activity against MCF-7 breast cancer cells.
Experimental
General
The equipment used in this study includes glassware commonly employed in the Organic Chemistry Laboratory of Natural Products and Synthesis Universitas Padjadjaran. Additionally, other supporting equipment such as distillation apparatus, maceration, partitioning tools, and a Buchi R-215 rotary evaporator equipped with a V-700 vacuum system, a B-491 water bath, and an F-100 Buchi cooling circulator were utilized for the concentration of macerates and fractions. Compound separation was guided by thin-layer chromatography with a UV detector lamp (λ254 and λ365), as well as 10% sulfuric acid in ethanol as a spotting reagent. Partitioning and purification were carried out using a separatory funnel and a chromatography column. For the analysis and characterization of isolates, infrared absorption was performed using KBr plates with an FTIR spectrometer (Perkin Elmer One). NMR spectra were measured with a JEOL JNM-ECZ700/S1 700 MHz NMR spectrometer for 1H-NMR and 175 MHz for 13C NMR with TMS as the standard, conducted at BRIN (Badan Riset dan Inovasi Nasional) Serpong Indonesia. Additionally, 500 MHz for 1H-NMR and 125 MHz for 13C-NMR with TMS as the standard were performed at Institut Teknologi Bandung. The relative molecular mass was determined using an HR-TOF-MS mass spectrometer (Waters LCT Premier XE) at Institut Teknologi Bandung.
Plant material
The sample used in this study is the stem bark of Lansium domesticum Corr. cultivar piedjietan, obtained from Cililin, West Java, Indonesia and identified at the Laboratory of the Department of Biology, Universitas Padjadjaran, with specimen code 10188.
Cytotoxic bioassay
The materials used in the cytotoxicity activity test include antibiotics (Sigma Aldrich P4333), cisplatin (EDQM C2210000), dimethyl sulfoxide (DMSO), FBS (fetal bovine serum) (Gibco 18912-014), PBS (phosphate-buffered saline), resazurin, RPMI (Roswell Park Memorial Institute), trypsin-EDTA, and trypan blue. The cytotoxicity test of the pure compound against MCF-7 breast cancer cells and normal cell CV-1 was conducted by incubating the cells in a 96-well plate at 37 °C and 5% CO2 until cell growth reached 70%. The cells were then treated with the sample and incubated for 48 h, followed by the addition of resazurin reagent to measure absorbance using a multimode reader. The media preparation included RPMI, FBS, and antibiotics, with cisplatin as the positive control. The cells, which reached 70% confluence, were treated with trypsin-EDTA and centrifuged, then seeded into a 96-well plate. The assay was performed in duplicate, and both samples and controls were tested across a various of concentrations. For MCF-7 cells (breast cancer cells), the concentrations used were 1.17, 2.34, 4.69, 9.38, 18.75, 37.50, 75.00, and 150.00 µg/mL, while for CV-1 cells (normal cells), the concentrations were 7.81, 15.63, 31.25, 62.50, 125.00, 250.00, 500.00, and 1000.00 µg/mL. Absorbance was measured at a wavelength of 570 nm to determine the IC50 value. The absorbance data obtained were converted into percentage cell viability at each concentration, and the slope of the resulting dose–response curve was used to determine the IC50 value, expressed in µg/mL.
Extraction and isolation
The separation process was initiated by macerating a 2.7 kg sample (stem bark of Lansium domesticum Corr. cultivar piedjietan) using 97% ethanol for 8×24 h at room temperature, followed by concentration using a rotary evaporator at 40 °C. Partitioning was then carried out using n-hexane, ethyl acetate, and butanol as solvents, with the n-hexane extract selected for isolation due to its characteristic response to the Liebermann-Burchard reagent, indicating the presence of terpenoids.
The
n-hexane
extract (145.1 g) was separated using vacuum liquid chromatography
with silica gel and a gradient elution system of n-hexane:ethyl
acetate:methanol (10%), resulting in ten fractions (A
-
J).
Fraction E (44.1 g) was further separated using silica gel column
chromatography with a gradient system of
n
-hexane:ethyl acetate (1%), yielding four sub-fractions (E1
-
E4).
Sub-fraction E3 (5.7 g) was further purified by silica gel column
chromatography with gradient elution of n-hexane:ethyl
acetate (2.5%), resulting in six
sub-fractions (E3a
-
E3f).
Sub-fraction E3b (0.7577 g) was re-separated using silica gel column
chromatography with a gradient system of
n-hexane:ethyl
acetate (1%), followed by reverse-phase chromatography using ODS
with a methanol:water (10:1) isocratically, resulting in compounds 2
(1.7 mg) and 3 (2 mg). Fraction E3c (1.0826 g) was further separated
by silica gel column chromatography with a gradient system of n
-hexane:ethyl acetate (1%), yielding compound 1 (13 mg).
The ethyl acetate extract (18.9 g) was separated using vacuum liquid chromatography with silica gel and a gradient elution system of n-hexane:ethyl acetate:methanol (10%), resulting in six fractions (A-F). Fraction D was separated using vacuum liquid chromatography with a gradient elution system of n-hexane:ethyl acetate:methanol (5%), yielding six sub-fractions (D1-D6). Sub-fraction D5 was further purified using open-column chromatography with ODS and a mobile phase of chloroform:methanol (9.6:0.4) isocratically, yielding compound 4.
Molecular docking (in silico)
This study utilizes molecular docking to assess the interactions between test compounds and the alpha estrogen receptor using AutoDock Vina. The 3D structures of the test ligands were retrieved from PubChem and subjected to geometry minimization in Chem3D using the MM2 method to ensure that the compounds adopt a stable, low-energy conformation. The 3D structure of the alpha estrogen receptor (PDB ID: 3ERT, https://www.rcsb.org/structure/3ERT) was obtained from the Protein Data Bank and prepared by removing water molecules and the native ligand bound to the active site using BIOVIA Discovery Studio, to eliminate potential interference from other molecules. Both the ligand and receptor structures were then processed in AutoDockTools, where Gasteiger charges were applied to the ligand and Kollman charges to the receptor, ensuring proper charge distribution. In addition, all hydrogen atoms were added to the ligands, while only polar hydrogen atoms were added to the receptor. The prepared structures were saved in pdbqt format for docking simulations.
Docking simulations were performed using AutoDock Vina, with the grid centered on coordinates x = 30.010, y = –1.913, z = 24.206, corresponding to the position of the native ligand Tamoxifen within the receptor. The grid box size was set to 40×40×40 to cover the entire active site. The docking simulations were repeated 100 times (runs) to ensure consistency and accuracy, with parameters such as energy range and exhaustiveness optimized to yield the best binding conformations. To verify the reliability of the docking protocol, a re-docking validation step was performed. The native ligand Tamoxifen was removed from the receptor and docked back into the same binding site using the exact protocol applied to the test ligands. The top ranked pose from Vina was then compared with the original crystallographic pose. The degree of agreement between the predicted binding pose and the crystallographic pose was assessed using the Root Mean Square Deviation (RMSD). A protocol is considered valid when the RMSD between the redocked pose and the native pose is less than 2.0 Å, which indicates that the method can reproduce the experimental binding orientation with acceptable accuracy.
The results were analyzed based on binding affinity (in kcal/mol) and the types of interactions, including hydrogen bonds and hydrophobic interactions, between the ligand and receptor. This analysis aimed to evaluate the potential of the test compounds to inhibit the activity of the alpha estrogen receptor, contributing to the development of therapeutic agents for the treatment of estrogen-dependent breast cancer.
Molecular dynamics
Molecular dynamics simulations were carried out with the Amber20 suite. The protein ligand complex for each system was built in the LEaP module. The protein followed the ff14SB force field parameters and the ligands were assigned parameters from the gaff2 force field. Each complex was placed in a tetrahedral periodic TIP3P water box that extended 10 Å from the solute, and the systems were neutralized with Na⁺ ions. Before running the simulations, the systems underwent several preparation steps. Energy minimization was performed in three phases. The first 2,000 cycles minimized only the solvent while keeping the solute fixed. The second 2,000 cycles applied restraints to the protein backbone. The final 4,000 cycles minimized the entire system without restraints. The minimized systems were then heated from 0 to 310 K for 50 ps with weak harmonic restraints of 5.0 kcal mol⁻¹ Å⁻² on solute atoms, followed by a 200 ps equilibration under NPT conditions at 1 atm. Production simulations were performed for 100 ns under the NPT ensemble at 310 K and 1 atm using the Langevin thermostat and the Berendsen barostat. Binding free energies were evaluated using the MM/GBSA approach implemented in the MMPBSA.py program in AmberTools. The calculations were based on 5,000 snapshots taken at regular intervals from the production trajectories, and water molecules and ions were removed before the analysis.
Pharmacokinetics properties
To analyze the Rule of Five and drug-likeness of the isolated compounds, tamoxifen, and estrogen, computational tools such as pkCSM (https://biosig.lab.uq.edu.au/pkcsm/) and ADMETlab 3.0 (https://admetlab3.scbdd.com/) were used. These platforms were employed to assess the pharmacokinetic properties, including absorption, distribution, metabolism, excretion, and toxicity profiles of the compounds, providing valuable insights into their potential as drug candidates. The Rule of Five, a guideline for evaluating the drug-likeness of compounds based on molecular properties, was also applied to determine whether the compounds meet the criteria for oral drug absorption
Results and discussion
Isolation and characterization
The
research workflow began with sample preparation, using the bark of
Lansium
domesticum
Corr. cv. Piedjietan,
which was collected from Cililin, West Java Indonesia, and
identified at the Laboratory of the Department of Biology,
Universitas Padjadjaran (specimen code: 10188). The sample was first
air-dried at room temperature without direct sunlight and then
finely ground using a grinder until 2.7 kg of fine powder was
obtained. The powdered bark sample was then macerated with ethanol
as the solvent, yielding 543.8 g of ethanol extract.
The
obtained crude ethanol extract was then fractionated using n-hexane,
ethyl acetate, and n-butanol
as solvents. Each resulting fraction was concentrated using a rotary
evaporator, yielding the following amounts of crude extract:
n-hexane
extract (145.1 g), ethyl acetate extract (54.1 g), and n-butanol
extract (9.8 g).
The
n-hexane
extract from the bark of
L.
domesticum
Corr. was further separated using various chromatographic
techniques, guided by spot visualization with 10% sulfuric acid in
ethanol, leading to the isolation of compounds 1, 2, and 3.
Meanwhile, from the ethyl acetate extract, compound 4 was
successfully isolated.
Compound 1 characterization
Compound 1 is a colorless oil with a TOF-MS spectrum showed a molecular ion peak at m/z 485.3642 [M+H]⁺ (Figure S1), corresponding to the molecular formula C₃₁H₄₈O₄ with a degree of unsaturation of eight. Its IR spectrum (KBr) exhibited absorption bands at 3,079, 2,956, 1,738, 1,078 and 1,644 cm⁻¹ (Figure S2). Compound 1 did not exhibit fluorescence under UV light at λ254 nm and λ365 nm, indicating the absence of electronic conjugation involving n → π* or π → π* transitions (no conjugated double bonds).
The ¹H-NMR spectrum (CDCl₃, 500 MHz) of compound 1 displayed characteristic signals of an aliphatic triterpenoid, marked by six singlet methyl resonances at δH 0.73 (3H, s), 0.82 (3H, s), 1.72 (3H, s), 1.78 (6H, s), and 3.67 ppm (3H, s). The signal at δH 3.67 ppm (3H, s) is characteristic of an oxygenated methyl group. Additionally, the spectrum revealed three olefinic methylene (sp²) signals at δH 4.86 ppm (s, 2H), 4.69 ppm (s, 2H), and 4.82 ppm (s, 2H), along with one olefinic methine (sp²) proton at δH 5.38 ppm (s, 1H). Based on the ¹H-NMR data, compound 1 is proposed to belong to the onoceranoid triterpenoid class, with a seco modification in its three core rings. It consists of six singlet methyl groups, one olefinic methine (sp²), and three olefinic methylene groups (sp²) (Figure S3).
The
¹³C-NMR spectrum analysis (CDCl₃, 125 MHz) and DEPT of compound
1
revealed the presence of 31 carbon atoms, consisting of six methyl
groups, including two sp³
methyl groups [δC
17.7 ppm (C-25); 16.3 ppm (C-28)], three sp²
methyl groups [δC
23.6 ppm (C-23); 23.4 ppm (C-27); 22.9 ppm (C-30)], and one
oxygenated methyl group [δC
51.1 ppm (C-31)]. The spectrum also showed twelve methylene groups,
consisting of nine sp³
methylene groups [δC
28.3 ppm (C-1); 32.4 ppm (C-2); 29.5 ppm (C-6); 38.1 ppm (C-7);
27.25 ppm (C-11); 27.4 ppm (C-12); 30.5 ppm (C-16); 29.1 ppm (C-19);
32.8 ppm (C-20)], and three sp²
methylene groups [δC
113.7 ppm (C-24); 107.6 ppm (C-26); 114.0 ppm (C-29)]. Additionally,
five methine groups were identified, consisting of four sp³
methine groups [δC
47.9 ppm (C-5); 50.8 ppm (C-9); 51.8 ppm (C-13); 48.9 ppm (C-17)]
and one sp²
methine group [δC
121.9 ppm (C-15)]. Eight
quaternary carbon atoms were detected, including two sp³
quaternary carbons [δC
41.6 ppm (C-10); 38.8 ppm (C-18)], four sp²
quaternary carbons [δC
147.1 ppm (C-4); 147.9 ppm (C-8); 135.9 ppm (C-14); 147.6 ppm
(C-22)], and two carbonyl quaternary carbons [δC
178.4 ppm (C-3); 175.5 ppm (C-21)] (Figure
S4).
Based on 1D and 2D NMR (Figures
S5 -
S7)
analysis and comparison with literature data, compound 1 is a
bicyclic onoceranoid with seco
structures on rings A, C, and E, bearing a carboxylic acid group at
C-3 and an ester group at C-21. This structure has not been
previously reported and represents a
new compound isolated
from Lansium
domesticum
Corr. cv. Piedjietan.
The compound has been named methyl lansic. This compound
exhibited significant cytotoxic activity against MCF-7 breast cancer
cells, with an IC₅₀ value of 7.78 μg/mL (Figure
S30),
whereas its IC₅₀ value against normal CV-1 cells was 620.30
μg/mL (Figure
S31),
indicating selective cytotoxicity toward breast cancer cells. The
structure of the compound is shown in Figure
1.
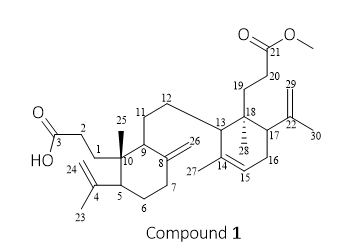
Figure 1 Structure of compound 1 (Methyl Lansic).
Compound 2 characterization
Compound 2 is a colorless crystalline substance that is soluble in chloroform and acetone. It does not exhibit fluorescence under UV light at λ254 nm and λ365 nm. Its IR spectrum (KBr) exhibited absorption bands at 2,936, 2,854, 1,708, 1,645, 1,457 cm−1 (Figure S8). The ¹H-NMR spectrum (CDCl₃, 700 MHz) of compound 2 displayed characteristics of an aliphatic triterpenoid, as indicated by the presence of seven singlet methyl signals resonating at δH 0.81; 0.95; 1.02; 1.05; 1.09; 1.10 and 1.72 ppm (3H, s). Additionally, the spectrum revealed the presence of an olefinic methylene (sp²) at δH 4.93 and 4.62 ppm (s, 1H), as well as an olefinic methine (sp²) at δH 5.43 ppm (s, 1H), which are characteristic of the onoceranoid group [18]. Based on ¹H-NMR data (Figure S9), compound 2 is suggested belonging to onoceranoid triterpenoid class, possessing seven singlet methyl groups, one olefinic methine (sp²), and one olefinic methylene (sp²).
Based on the ¹H-NMR, ¹³C-NMR (CDCl₃, 175 MHz), DEPT, and HMBC data (Figures 10 - 11), compound 2 is proposed to have the molecular formula C₃₀H₄₆O₂ with eight degrees of unsaturation (DBE). The presence of eight DBE is attributed to two double bonds at δC 147.3 ppm (C-8) and δC 107.6 ppm (C-26), as well as δC 135.4 ppm (C-14) and δC 121.9 ppm (C-15); two carbonyl (ketone) groups at δC 216.8 ppm (C-21) and δC 217.0 ppm (C-3); and four cyclic rings. The presence of 30 carbon atoms, seven methyl signals, ten methylene groups, five methine groups, eight quaternary carbons, and four cyclic rings suggests that the structure of compound 2 follows the onoceranoid triterpenoid skeleton. The ¹³C-NMR spectrum shows characteristic carbon signals, including four sp² carbons, consisting of two quaternary sp² carbons at δC 135.4 ppm (C-14) and 147.3 ppm (C-8), one sp² methine carbon at δC 121.9 ppm (C-15) substituted at C-14, and one sp² methylene carbon at δC 107.6 ppm (C-26) substituted at C-8. These shifts are distinctive features of onoceranoid triterpenoids, as they indicate the cleavage of ring C at carbon atoms C-8 and C-14. The quaternary carbon signals at δC 216.8 ppm (C-21) and 217.0 ppm (C-3) are also characteristic of oxygenated carbons, specifically ketone functional groups. Based on 1D and 2D NMR (Figures S12 - S14) analyses and comparison with literature data, compound 2 was identified as α,γ-onoceradienedione. This structure has previously been reported from the fruit peel extract of Lansium domesticum Corr. cultivar kokossan [10], but is reported here for the first time from the stem bark of Lansium domesticum Corr. cultivar Piedjietan. The compound has been named α, γ-onoceradienedione, and its structure is depicted in Figure 2.
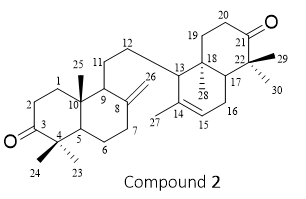
Figure 2 Structure of compound 2 (α, γ-onoceradienedione).
Compound 3 characterization
Compound
3
is a colorless crystalline substance that is soluble in chloroform
and acetone with
a TOF-MS spectrum showed a molecular ion peak at m/z 469.3663
(Figure
S15),
corresponding to the molecular formula C₃₁H₄₈O3
with a degree of unsaturation of eight.
Its
IR spectrum (KBr) exhibited absorption bands at 2,950, 1,739, 1,708,
1,644, 1,436 and 1,328 cm⁻¹ (Figure
S9).
Compound 3 did not exhibit fluorescence under UV light at λ254
nm and λ365
nm, indicating the absence of electronic conjugation involving n →
π* or π → π* transitions (no conjugated double bonds).
The
1H-NMR
(CDCl₃, 700 MHz) spectrum of compound 3 exhibited characteristic
signals of an aliphatic triterpenoid, as indicated by the presence
of six singlet methyl proton signals resonating at δH
0.82;
0.85;
1.02;
1.10;
1.74
and
1.78 ppm
(3H, s).
Additionally, two olefinic methylene groups (sp²)
were observed at δH
4.63 ppm (1H,
s),
4.91 ppm (1H, s), and 4.82 ppm (2H, s), along with one olefinic
methine (sp²)
proton at δH
5.38 (1H, s), a feature typical of the onoceranoid class of
compounds. Based on the 1H-NMR
data
(Figure
S17),
compound 3 is suggested to be an onoceranoid-type triterpenoid
possessing six methyl singlets, one sp²
methine, and two sp²
methylene groups.
The
13C-NMR
(CDCl₃, 125 MHz) and DEPT spectra of compound 3 revealed the
presence of 31 carbon atoms, including seven methyls [δC
26.0 ppm
(C-23), 21.7 ppm (C-24), 14.1 ppm (C-25), 22.9 ppm
(C-27), 16.4 ppm (C-28), 22.9 ppm (C-30), and 22.9 ppm (C-31)], nine
sp³
methylenes [δC
37.6 ppm (C-1), 34.8 ppm (C-2), 25.1 ppm (C-6), 37.8 ppm (C-7),
36.2 ppm (C-11), 32.8 ppm (C-12), 27.2 ppm (C-16), 29.5 ppm
(C-19), and 28.9 ppm (C-20)], two sp²
methylenes [δC
107.6 ppm (C-26), 113.9 ppm (C-29)], four sp³
methines [δC
55.1 ppm (C-5), 57.5 ppm (C-9), 48.2 ppm (C-13), and 49.1 ppm
(C-17)], one sp²
methine [δC
121.8 ppm (C-15)], and eight quaternary carbons, including three sp³
quaternary carbons [δC
47.7 ppm
(C-4), 39.3 ppm (C-10), 38.6 ppm (C-18)], three sp²
quaternary carbons [δC
147.5 ppm (C-8), 135.8 ppm
(C-14), 147.6 ppm (C-22)], and two
carbonyl quaternary carbons [δC
216.8 ppm (C-3), 174.5 ppm (C-21)] (Figure
S18).
These data suggest that compound 3 has a molecular formula of
C₃₁H₄₈O3
with eight degrees of unsaturation (DBE). The eight DBE are presumed
to arise from three double bonds (C-8/C-26, C-14/C-15, and
C-22/C-29), two carbonyl groups (C-3 and C-21), and three rings.
Based on 1D and 2D NMR
(Figures
S19 -
S22)
analyses and comparison with literature data, compound 3 was
identified as methyl ester lansiolate. This structure has previously
been reported from the fruit peel extract of Lansium
domesticum
Corr. cultivar duku
[8],
but is
reported here for the first time from the stem bark of Lansium
domesticum
Corr. cultivar Piedjietan.
The compound has been named methyl ester lansiolate, and its
structure is depicted in Figure
3.
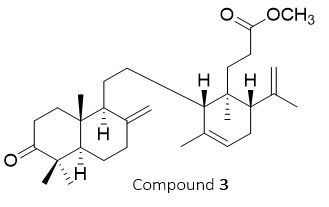
Figure 3 Structure of compound 3 (methyl ester lansiolate).
Compound 4 characterization
Compound 4 is a colorless oil with a TOF-MS m/z of 457.3426 [M+H]⁺ (Figure S23), corresponding to the molecular formula C₃₀H₄₈O₃ with a degree of unsaturation of seven. Its IR spectrum (KBr) exhibited absorption bands at 3,500, 2,965, 1,710, 1,442 and 1,381 cm⁻¹ (Figure S24). The ¹H-NMR spectrum (CDCl₃, 500 MHz) shows the presence of six singlet signals from methyl groups at δH 0.67; 0.76; 0.90; 0.81; 1.77 and 1.73 ppm (3H, s), which correspond to sp³ protons. Signals at δH 4.50 and 4.83 ppm, 4.70 and 4.82 ppm, and 5.36 ppm originate from sp² protons, along with a signal at δH 3.26 ppm attributed to an oxygenated carbon (Figure S25). Based on the ¹H-NMR analysis, compound 4 contain 48 protons, consisting of five sp² protons, one oxygenated sp³ proton, 40 sp³ protons, and two hydroxyl protons.
The
¹³C-NMR and DEPT spectrum (CDCl₃, 125 MHz) indicates the
presence of 30 carbon signals (Figure
S26),
including two sp²
methylene carbons at
δC
106.9 and 114.1 ppm, one sp²
methine carbon at
δC
121.8 ppm, and four sp²
quaternary carbons at
δC
136.1, 147.8, 148.4 ppm, and one carbonyl carbon
at δC
178.0 ppm. The proton H-23 (δH
0.98 ppm) and the proton H-24 (δH
0.76 ppm) exhibit three-bond correlations with C-3 (δC
79.4 ppm), which is an oxygenated methine carbon bonded to a
hydroxyl group, as well as with C-5 (δC
54.6 ppm), and show two-bond correlations with C-4 (δC
39.2 ppm). This suggests that the two methyl groups are attached to
carbon number four and are adjacent to the hydroxyl group. The
proton H-24 exhibits a four-bond correlation with C-6
(δC
27.9 ppm), while the proton H-23 shows a four-bond correlation with
C-24 (δC
15.6 ppm), indicating that the methyl groups at C-23 and C-24 are
attached to the same carbon. The position of the olefinic groups in
this compound’s structure is confirmed by the HMBC (Figure
28).
The olefinic group is identified through three-bond correlations
between H-26 (δH
4.5 & 4.83 ppm) and C-9 (δC
58.5 ppm) and C-7 (δC
38.2 ppm). Meanwhile, the olefinic position at C-22 and C-29 is
confirmed by three-bond correlations between H-29 (δH
4.70 & 4.82 ppm) and C-17 (δC
49.2 ppm), as well as between H-29 and C-30 (δC
23.1 ppm). These correlations indicate the neighboring relationship
between C-30 and C-29. Based on 1D and 2D NMR (Figures
S27
-
S29)
analyses and comparison with literature data, compound 4 was
identified as lansiolic acid. This structure has previously been
reported from the fruit peel extract of Lansium
domesticum
Corr. cultivar duku
[19],
but is reported here for the first time from the stem bark of
Lansium
domesticum
Corr. cultivar Piedjietan.
The compound has been named lansiolic
acid,
and its structure is depicted in Figure
4.
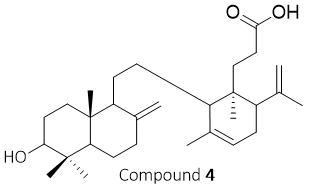
Figure 4 Structure of compound 4 (lansiolic acid).
The chemical shift of the compounds 1-4, which was first isolated from Lansium domesticum Corr cv. Piedjietan, can be seen in Table 1, and the HMBC and COSY correlations are shown in Figure 5.
Table 1 1H-NMR and 13C-NMR methyl lansic (1), α,γ-onoceradienedione (2), methyl ester lansiolate (3) and lansiolic acid (4).
Carbon position |
1 |
2 |
3 |
4 |
||||
δC (125 MHz, CDCl3) |
δH (500 MHz, CDCl3)
|
δC (175 MHz, CDCl3) |
δH (700 MHz, CDCl3) |
δC (175 MHz, CDCl3) |
δH (700 MHz, CDCl3) |
δC (175 MHz, CDCl3) |
δH (500 MHz, CDCl3)
|
|
1 |
28.3 |
1.47 (2H, m) |
37.7 |
2.01 (2H, m) |
37.6 |
2.44 (2H, m) |
37.1 |
1.81 (2H, m) |
2 |
32.4 |
1.73 (2H, m) |
34.7 |
2.25 (2H, m) |
34.8 |
2.44 (2H, m) |
29.6 |
1.24 (2H, m) |
3 |
178.4 |
- |
217.0 |
- |
216.85 |
- |
79.4 |
3.26 (1H, dd) |
4 |
147.1 |
- |
47.8 |
- |
47.7 |
- |
39.2 |
- |
5 |
47.9 |
1.85 (1H, m) |
55.2 |
1.67 (1H, m) |
55.1 |
1.63 (1H, m) |
54.6 |
1.08 (1H, m) |
6 |
29.5 |
1.22 (2H, m) |
25.2 |
1.72(2H, m) |
25.1 |
1.70, (2H, m) |
27.9 |
1.39 (2H, m) |
7 |
38.1 |
1.99 (2H, m) |
37.9 |
2.45 (2H, m) |
37.8 |
2.04 (2H, m) |
38.2 |
1.98; 2.37 (1H, m) |
8 |
147.9 |
|
147.3 |
- |
147.5 |
- |
148.4 |
- |
9 |
50.8 |
2.22 (1H, m) |
56.0 |
60 (1H, m) |
57.5 |
1.68 (1H, m) |
58.5 |
1.56 (1H, m) |
10 |
41.6 |
- |
39.1 |
- |
39.3 |
- |
38.7 |
- |
11 |
27.3 |
1.22 (2H, m) |
24.0 |
1.94; 2.08 (1H, m) |
26.2 |
1.45 (2H, m) |
26.1 |
1.09 (2H, m) |
12 |
27.4 |
1.21 (2H, m) |
25.1 |
1.42 (2H, m) |
32.8 |
1.68 (2H, m) |
24.1 |
1.36 (2H, m) |
13 |
51.8 |
1.8 (1H, m) |
54.3 |
1.65 (1H, m) |
48.2 |
1.86, (1H, m) |
48.4 |
1.85 (1H, m) |
14 |
135.9 |
- |
135.4 |
- |
135.8 |
- |
136.1 |
- |
15 |
121.9 |
5.38 (1H, brs) |
121.9 |
5.43 (1H, brs) |
121.8 |
5.38 (1H, brs) |
121.8 |
5.36 (1H, brs) |
16 |
30.5 |
1.74 (2H, m) |
25.3 |
1.40; 1.70 (1H, m) |
27.2 |
1.22 (2H, m) |
27.4 |
1.62 (2H, m) |
17 |
48.9 |
2.22 (1H, m) |
51.5 |
1.50 (1H, m) |
49.1 |
2.22 (1H, m) |
49.2 |
2.19 (1H, m) |
18 |
38.8 |
- |
36.4 |
- |
38.6 |
- |
39.5 |
- |
19 |
29.1 |
1.8 (2H, m) |
38.1 |
1.26 (2H, m) |
29.5 |
1.86 (2H, m) |
32.6 |
1.64 (2H, m) |
20 |
32.8 |
5.16 (2H, m) |
34.7 |
2.46 (2H, m) |
28.9 |
2.38 (2H, m) |
28.9 |
2.43 (2H, m) |
21 |
175.5 |
- |
216.8 |
- |
174.5 |
- |
178.0 |
- |
22 |
147.6 |
- |
47.5 |
- |
147.61 |
- |
147.8 |
- |
23 |
23.6 |
1.73 (3H, s) |
26.0 |
1.10 (3H, s) |
26.0 |
1.10 (3H, s) |
28.4 |
0.98 (3H, s) |
24 |
113.7 |
4.86 (3H, s) |
21.6 |
1.05 (3H, s) |
21.7 |
1.02 (3H, s) |
15.6 |
0.76 (3H, s) |
25 |
17.7 |
0.72 (3H, s) |
14.2 |
0.85 (3H, s) |
14.1 |
0.85 (3H, s) |
14.7 |
0.67 (3H, s) |
26 |
107.6 |
4.69 (2H, s) |
107.6 |
4.62 (2H, s) |
107.6 |
4.91; 4.63 (1 H, s) |
106.9 |
4.5; 4.83 (1 H, s) |
27 |
23.4 |
1.78 (3H, s) |
22.2 |
4.93 (3H, s) |
22.9 |
1.74 (3H, s) |
23.0 |
1.73 (3H, s) |
28 |
16.3 |
0.82 (3H, s) |
13.3 |
1.72 (3H, s) |
16.4 |
0.82(3H, s) |
16.4 |
0.81 (3H, s) |
29 |
114.0 |
4.79 (1H, s); 4.82 (1H, s) |
22.1 |
0.95 (3H, s) |
113.9 |
4.82 (2H, s) |
114 |
4.70 (1H, s); 4.82(1H, s) |
30 |
22.9 |
1.78 (3H, s) |
25.0 |
1.10 (3H, s) |
22.9 |
1.78 (3H, s) |
23.1 |
1.77 (3H, s) |
31 (Methoxy) |
51.1 |
3.67 (3H, s) |
- |
- |
51.6 |
3.63 (3H, s) |
- |
|
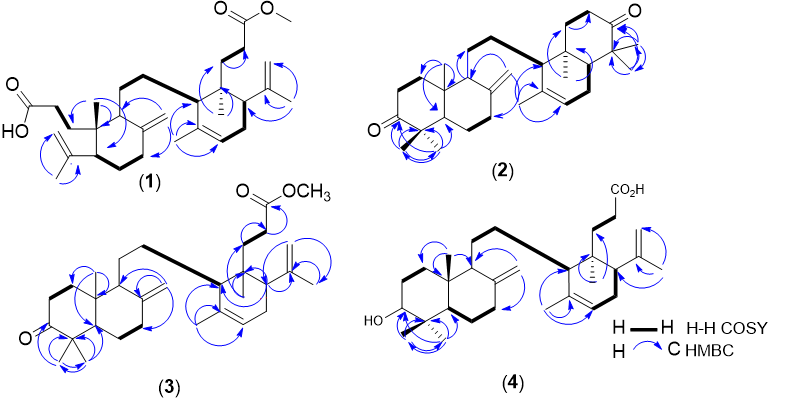
Figure 5 Structure of onoceranoid triterpenoid isolated from Lansium domesticum Corr. cv. Piedjietan; (1) methyl lansic, α, γ-onoceradienedione (2), methyl ester lansiolate (3), and lansiolic acid (4).
Molecular docking
The results from the molecular docking analysis provide comprehensive insights into the binding interactions and potential inhibitory effects of several test compounds, including tamoxifen, estrogen, lansiolic acid (4), methyl ester lansiolate (3), methyl lansic (1), and α, γ-onoceradiene (2), with the alpha estrogen receptor (Figure 6 and Table 2). Among these, tamoxifen (–9.8 kcal/mol) and α, γ-onoceradiene (2) (–9.9 kcal/mol) exhibited the strongest binding affinities, suggesting that these compounds have a high potential for targeting the receptor and modulating its activity. Interestingly, the test compounds did not bind directly to the active site of the receptor, which contrasts with the classical binding modes of reference ligands, such as estrogen and tamoxifen, which typically engage with the receptor’s active site to trigger receptor activation or inhibition [20]. This is likely due to the larger molecular sizes of the test compounds, which potentially hinder direct access to the receptor’s active site. The steric bulk of these molecules, in addition to their size, may prevent their proper orientation within the receptor’s pocket, thus limiting the ability of the ligands to bind directly to the receptor’s agonist-binding region.
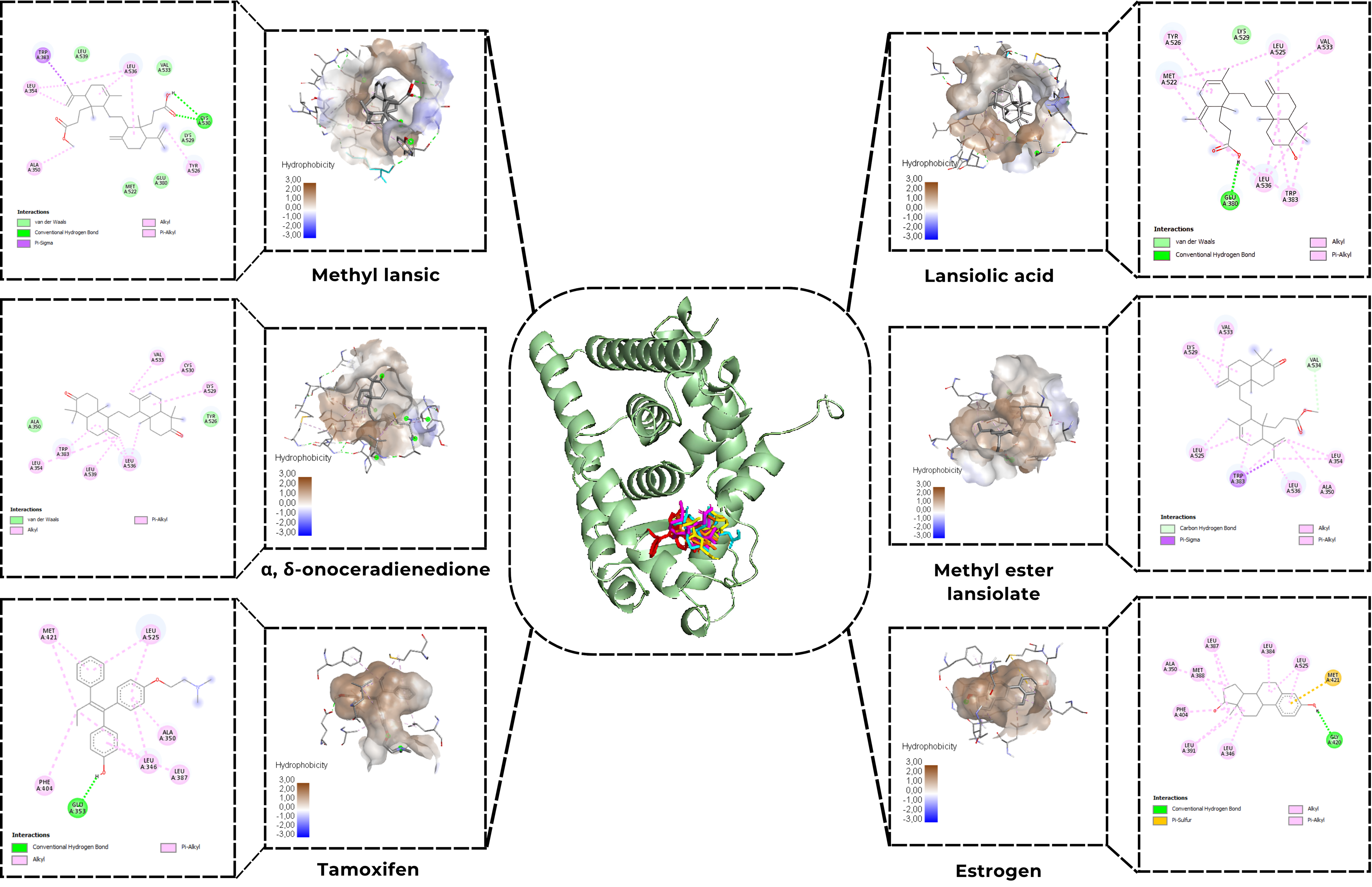
Figure 6 Molecular docking Methyl lansic (1), α,γ-onoceradienedione (2), methyl ester lansiolate (3), lansiolic acid (4), tamoxifen, and estrogen to estrogen receptor alpha.
Despite the absence of direct binding to the active site, the test compounds demonstrated similar binding orientations and interacted extensively with surrounding residues via hydrophobic contacts (Figure 6 and Table 2). These hydrophobic interactions play a pivotal role in stabilizing the ligand-receptor complex [21]. For instance, compounds like tamoxifen and methyl ester lansiolate (3) formed hydrophobic contacts with key residues such as Leu525, Leu346, and Ala350, which are involved in the receptor’s hydrophobic binding pocket. Such interactions are critical because they can effectively block the access of endogenous agonists like estrogen, preventing its binding and subsequent activation of the receptor. The displacement of estrogen from its active binding site due to these hydrophobic interactions implies that these test compounds act through an indirect, competitive inhibition mechanism, where they reduce the receptor’s activation potential by physically occupying the receptor’s binding pocket. The docking profiles indicate that Van der Waals interactions constitute the major stabilizing force for all four onoceranoid triterpenoids within the ERα binding pocket. These interactions arise from extensive hydrophobic contacts with Leu-, Val-, Met-, and Ala-rich regions. Notably, Methyl Lansic (1) relies heavily on these interactions, forming numerous Van der Waals contacts without participating in hydrogen bonding, which explains its relatively weaker binding affinity despite its stable hydrophobic enclosure.
Table 2 Molecular docking analysis methyl lansic (1), α,γ-onoceradienedione (2), methyl ester lansiolate (3) and lansiolic acid (4).
Compounds |
Parameter |
||
Binding Affinity (Kcal/mol) |
Hydrogen bonds |
Hydrophobic contacts |
|
Tamoxifen |
–9.8 |
Glu353 |
Met431, Leu525, Phe404, Leu346, Leu387, Ala350 |
Estrogen |
–9.4 |
Gly420 |
Leu525, Leu384, leu387, Ala350, Met388, Phe404, Leu391, Leu346 |
Lansiolic acid (4) |
–8.0 |
Glu380 |
Tyr526, Met522, Leu525, Val533, Leu536, Trp383, Lys529 |
Methyl ester lansiolic (3) |
–8.2 |
- |
Val533, Lys529, leu525, leu536, Ala350, Leu354 |
Methyl lansic (1) |
–7.3 |
Cys530 |
Leu536, Leu354, Ala350, Tyr526, leu539, Val533, Glu380, Met522 |
α, γ-onoceradienedione (2) |
–9.9 |
- |
Val533, Cys530, Lys529, leu356, Leu539, Trp383, leu354 |
His competitive inhibition mechanism may explain the reduced activity of the estrogen receptor alpha (ERα) in the presence of these test compounds. By hindering the binding of natural ligands, such as Estrogen, the test compounds could effectively lower the likelihood of receptor activation, a key step in the signaling cascade that mediates the proliferation of estrogen-responsive cancer cells, such as in breast cancer [22]. Given that the activation of the ERα is involved in tumor growth and metastasis, the inhibition of receptor activation is a promising strategy for cancer therapy, particularly for estrogen-receptor-positive (ER+) breast cancers. The findings are consistent with previous research on selective estrogen receptor modulators (SERMs), which are known to block or modulate ERα activity by interacting with the receptor’s ligand-binding domain [11]. While the test compounds studied here exhibited moderate binding affinities compared to estrogen and tamoxifen, they still represent viable candidates for further development as anticancer agents. Lansiolic acid (4) (–8.0 kcal/mol), methyl ester lansiolate (3) (–8.2 kcal/mol) and methyl lansic (1) (–7.3 kcal/mol), despite having slightly lower binding affinities, demonstrated significant hydrophobic interactions with residues like Leu525 and Val533, suggesting they might still be effective in impeding the receptor’s activation. This implies that these compounds could potentially serve as less potent, but still valuable, modulators in a therapeutic setting.
Furthermore, the indirect inhibition of ERα activity by the test compounds may offer an advantage over traditional agonistic therapies, as it could avoid some of the unwanted side effects associated with complete receptor activation, such as endocrine resistance and tumor recurrence [23], The potential for these compounds to disrupt ERα-mediated signaling at the molecular level could provide a novel therapeutic avenue in breast cancer treatment, especially in cases where resistance to conventional hormone therapies, such as tamoxifen, has developed. The results from this molecular docking analysis suggest that these compounds, particularly α, γ-onoceradiene (2), methyl ester lansiolate (3), lansiolic acid (4), and methyl lansic (1), hold promise as selective estrogen receptor modulators (SERMs) that could be developed further as therapeutic agents for estrogen-receptor-positive breast cancer. The hydrophobic interactions at the receptor’s binding site, which prevent the binding of natural estrogen, serve as a competitive inhibition mechanism that reduces receptor activation and cell proliferation.
When evaluated from its chemical structure among the isolated compounds, methyl lansic (1) shows notable potential as a breast cancer drug candidate because the opening of the A and E rings creates new functional features that enhance molecular interactions with the target protein. The opening of ring A generates an oxygenated C-3 bearing a carboxylic acid group, while the opening of ring E results in C-21 forming a methyl ester. These newly exposed polar groups provide additional interaction sites within the ERα binding pocket, contributing to stronger and more diverse ligand–receptor contacts, including a key hydrogen bond with Cys530. This structural advantage aligns with its observed in vitro activity, reflected by an IC₅₀ value of 7.78 µg/mL against MCF-7 breast cancer cells. Furthermore, Methyl lansic (1) was also evaluated against normal cells and exhibited no cytotoxic activity, with an IC₅₀ value of 620.30 μg/mL, indicating a highly favorable selectivity profile toward breast cancer cells without affecting healthy cells.
Molecular dynamics simulation
The 100 nanosecond molecular dynamics simulation showed that the complex formed between methyl lansic and the ERα receptor remained structurally stable. The Root Mean Square Deviation (RMSD) values were low, with an average of 1.13 Å for the ligand and 1.44 Å for the receptor. These values indicate limited structural drift during the simulation. The stability of the ligand RMSD suggests that methyl lansic stayed in a consistent pose within the binding pocket. Minor fluctuations were linked to natural thermal motions such as internal sigma bond rotations and did not affect the integrity of the complex. Stability of the complex was supported by the persistence of key molecular interactions at the active site. Hydrogen bonds with CYS225 and LEU231 were consistently maintained throughout the trajectory. These interactions helped preserve the orientation of the ligand and contributed to the maintenance of the binding pocket architecture. MM/GBSA binding free energy analysis yielded a value of –36.0850 kcal/mol, which reflects a strong and spontaneous interaction. A value of this magnitude aligns with the affinity typically associated with lead compounds that display meaningful biological potential.
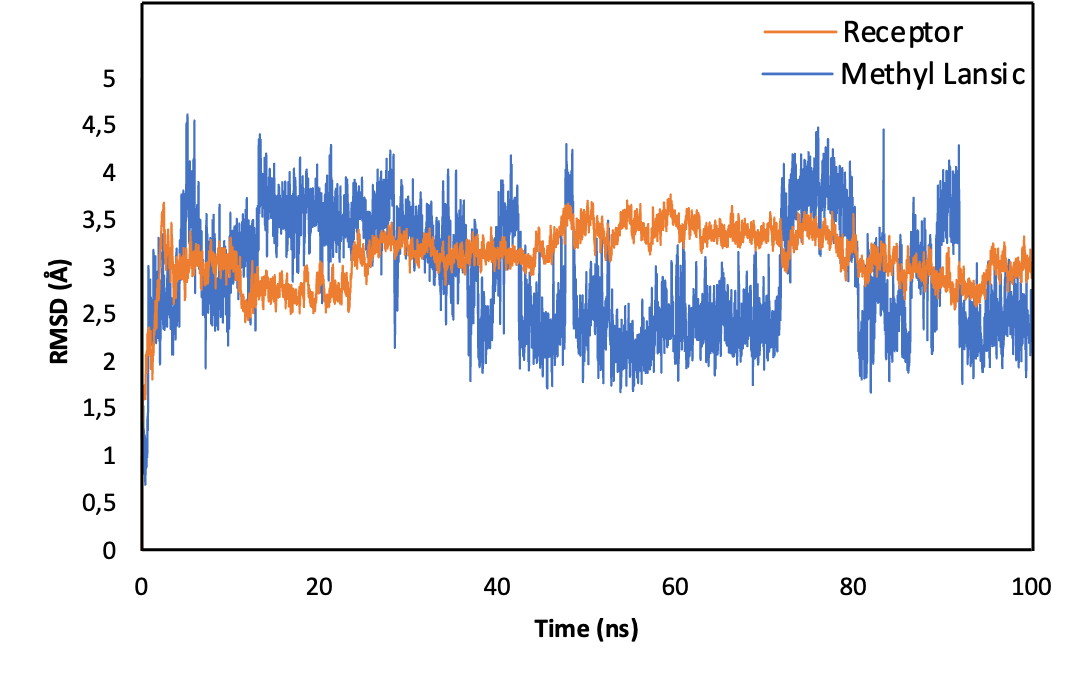
Figure 7 Root Mean Square Deviation (RMSD) profiles of the ERα–methyl lansic complex over a 100 ns molecular dynamics simulation. The receptor backbone (blue) and ligand (orange) show consistently low RMSD values, averaging 1.44 and 1.13 Å, respectively, indicating structural stability throughout the trajectory. Minor fluctuations correspond to natural thermal motions and do not disrupt the binding pose. The stable RMSD supports the persistence of key interactions, including hydrogen bonds with CYS225 and LEU231, maintaining the integrity of the binding pocket.
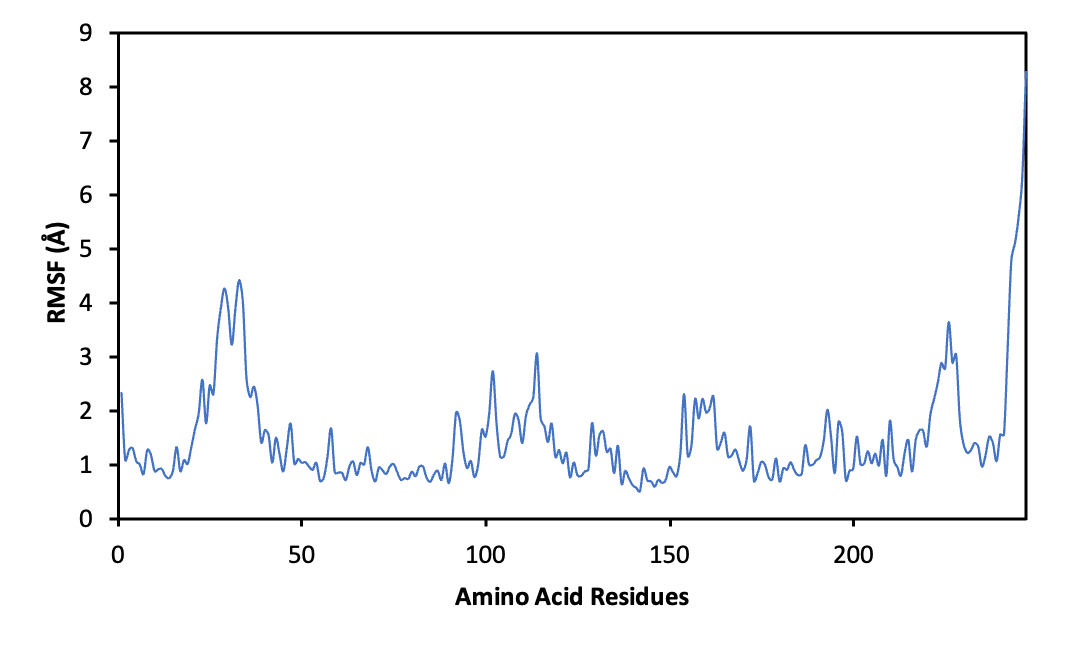
Figure 8 Root Mean Square Fluctuation (RMSF) profile of ERα residues during the 100 ns MD simulation of the ERα–methyl lansic complex. Residues lining the binding pocket exhibit low fluctuation amplitudes, indicating minimal local mobility and absence of ligand-induced destabilization. The overall pattern reflects a stable receptor environment that supports consistent ligand accommodation throughout the trajectory.
Root Mean Square Fluctuation (RMSF) analysis provided additional confirmation. Residues surrounding the binding pocket showed limited fluctuation, and there were no high peaks that would indicate local structural instability or ligand induced perturbation. This pattern supports the conclusion that methyl lansic does not trigger unfavorable conformational changes upon binding. Taken together, the RMSD and RMSF profiles, the interaction pattern with key residues, and the favorable binding free energy highlight the stable nature of the methyl lansic–ERα complex. These findings indicate that methyl lansic is a strong candidate for further biological testing in ERα positive breast cancer models.
Pharmacokinetics
Drug-likeness prediction
Lipinski’s Rule of five is used to assess whether a chemical compound with pharmacological or biological activity can be an orally active drug in humans [24]. This rule defines molecular properties that affect the pharmacokinetics of the drug, such as absorption, distribution, metabolism, and excretion (ADME). Key factors include molecular weight, hydrogen bonds, and the log P value. Larger molecules tend to have lower solubility, reducing absorption. Hydrogen bonds enhance solubility in water but hinder permeation through lipid membranes. A high partition coefficient (LogP) reduces solubility and absorption.
Table 3 Rule of Five Analysis of Methyl Lansic (1), α,γ-onoceradienedione (2), Methyl Ester Lansiolate (3) and Lansiolic Acid (4). All four compounds meet Lipinski’s Rule of Five, each presenting no more than one allowable violation. These parameters reflect favorable drug-likeness characteristics, supporting their potential suitability as orally active therapeutic candidates.
Compound |
MW (g mol−1) |
LogP |
HBA |
HBD |
Ro5 Violations |
Lipinski compliance |
Tamoxifen |
371.524 |
5.9961 |
2 |
0 |
1 |
Yes |
Estrogen |
272.388 |
3.6092 |
2 |
2 |
0 |
Yes |
Methyl lansic (1) |
484.721 |
7.9142 |
3 |
1 |
1 |
Yes |
α, γ-onoceradienedione (2) |
438.696 |
7.7222 |
2 |
0 |
1 |
Yes |
Methyl ester lansiolate (3) |
468.722 |
7.8624 |
3 |
0 |
1 |
Yes |
Lansiolic acid (4) |
456.711 |
7.5658 |
2 |
2 |
1 |
Yes |
As presented in Table 3, all four onoceranoid triterpenoids fulfilled Lipinski’s Rule of Five, each showing no more than one allowable violation. These results indicate that the compounds possess favorable drug-likeness properties and are suitable for consideration as orally active therapeutic candidates. Molecular weight of a compound is directly related to its ability to cross biological membranes, its solubility, and its potential to bind with biological targets, such as enzymes or receptors. an indication of the compound’s size and complexity, with larger molecules generally having more intricate structures. Lansiolic acid (4) (456.711 g/mol) and methyl ester lansiolate (3) (468.722 g/mol) have relatively high molecular weights, which may suggest more complex interactions within biological systems. The LogP value, which measures a compound’s lipophilicity by indicating its partition coefficient between octanol and water, is another key determinant in understanding solubility and permeability. Compounds with a LogP greater than five, such as Lansiloic acid (4) (LogP = 7.5658) and Methyl ester lansiolate (3) (LogP = 7.8624), are considered lipophilic and hydrophobic, meaning they are more soluble in fats than in water. These characteristics often lead to challenges in oral bioavailability, as hydrophobic compounds typically have reduced water solubility.
The number of hydrogen bond acceptors (HBA) and donors (HBD) plays a critical role in determining how compounds interact with their environment, particularly in aqueous solutions. Lower HBA and HBD values suggest that a compound is less likely to form hydrogen bonds with water molecules, thereby enhancing its hydrophobicity. For instance, α, γ-onoceradienedione (2), with two HBA, exhibits a reduced potential for hydrogen bonding, supporting its lipophilic nature. The hydrogen bonding capability also affects the interaction of the compound with biological targets, as hydrogen bonding is often involved in molecular recognition processes such as enzyme-substrate interactions or receptor-ligand binding.
The “Rule of five” is a well-known guideline that helps predict the drug-likeness of compounds based on their physicochemical properties[11]. According to this rule, compounds that violate more than one of the following criteria are less likely to be suitable oral drugs: Molecular weight greater than 500 g/mol, LogP greater than five, more than five hydrogen bond donors (HBD), or more than ten hydrogen bond acceptors (HBA). In the table, compounds like Lansiolic acid, Methyl ester lansiolate, and Methyl lansic violate the Rule of five primarily due to their high LogP values, which are indicative of their reduced solubility in aqueous environments. This violation suggests that these compounds may face challenges in oral drug administration but could still be effective through alternative routes such as injections or transdermal delivery.
ADMET analysis
From Table 4, lansiolic acid (4), methyl ester lansiolic (3), and α,γ-onoceradienedione (2) show interesting characteristics with distinct ADMET profiles. lansiolic acid (4) and methyl ester lansiolic (3) have good absorption and distribution, but their slower excretion rates and lower total clearance suggest they remain in the body longer, providing stability. α,γ-onoceradienedione also display good absorption and distribution, with slow excretion and no toxicity, suggesting a favorable safety profile.
Although Tamoxifen remains a gold standard in breast cancer treatment, its potential toxicity requires ongoing attention. Its advantageous absorption and excretion properties are balanced by the need to address its toxicity. Therefore, exploring alternative compounds is crucial to finding safer and more effective options for breast cancer therapy. Lansic acid (4) and methyl ester lansiolate (3) present more stable and safer profiles, which could be used as alternative treatments or in combination with tamoxifen to improve patient outcomes.
In the comparison of pharmacokinetic parameters of several compounds, tamoxifen and estrogen exhibit relatively rapid elimination, with tamoxifen having the highest clearance plasma (CLₚₗₐₛₘₐ) value (12.138), while estrogen is slightly lower (12.024), but with the longest half-life (T₁/₂) of 1.686, indicating that estrogen remains in the body longer. In contrast, Lansioic acid (4) (CLₚₗₐₛₘₐ 7.335), methyl ester lansioilate (3) (9.309), and methyl lansic (1) (5.774) exhibit lower CLₚₗₐₛₘₐ values, suggesting slower elimination, with methyl ester lansiolate (3) having the fastest half-life (0.289). α, γ-onoceradienedione (2) shows the slowest elimination rate in terms of both CLₚₗₐₛₘₐ (5.774) and half-life (0.242), indicating rapid clearance. Overall, these differences impact the duration of effect and accumulation of the compounds in the body, which is critical in determining their application for therapies that require careful management of circulation levels and effect duration. In the context of breast cancer treatment, tamoxifen, which has proven efficacy, has a relatively fast elimination rate, allowing for continuous use with optimal dose adjustments. On the other hand, candidate compounds like α, γ-onoceradienedione (2), which demonstrate faster elimination, may require further investigation to ensure their therapeutic effectiveness and potential to provide clinical benefits comparable to Tamoxifen, without leading to undesirable side effects.
Overall, while tamoxifen is widely used, the compounds discussed here particularly lansiolic acid (4) and methyl ester lansiolic (3) offer promising ADMET profiles for further development in breast cancer treatment. These compounds could potentially lead to safer, more stable therapies, either as stand-alone treatments or in combination with existing therapies like Tamoxifen, making them valuable candidates for future drug development.
Several limitations of the computational predictions warrant consideration. Docking provides a simplified and largely static view of ligand–ERα interactions, while ADMET and Lipinski’s Rule of Five rely on predictive models that cannot fully mimic in vivo pharmacokinetics. Likewise, the Molecular dynamics simulations are constrained by the simulation time and force-field assumptions. Therefore, these in silico results should be interpreted as supportive predictions and validated in conjunction with the experimental cytotoxicity data.
Table 4 ADME(T) (Absorption, Distribution, Metabolism, Excretion, Toxicity) Analysis of Methyl Lansic (1), α,γ-onoceradienedione (2), Methyl Ester Lansiolate (3) and Lansiolic Acid (4).
Parameter |
Compound |
|
|||||
Tamoxifen |
Estrogen |
Methyl lansic (1) |
α, γ-onoceradienedione (2) |
Methyl ester lansiolate (3) |
Lansiolic acid (4) |
||
Absorption |
Instensial absorption (human) (%) |
97.702 |
93.025 |
97.482 |
96.731 |
97.569 |
94.352 |
Permeability Caco-2 (log Papp in 10-6cm/s) |
1.131 |
1.459 |
0.753 |
1.197 |
1.241 |
1.202 |
|
Distribution |
Volume Distribution human (logL/kg) |
0.58 |
0.62 |
–0.915 |
0.487 |
0.24 |
–0.754 |
BBB |
1.332 |
0.015 |
–0.385 |
0.265 |
–0.177
|
–0.177
|
|
Metabolism |
CYP2D6 substrate |
No |
No |
No |
No |
No |
No |
CYP2D6 inhibitor |
Yes |
No |
No |
No |
No |
No |
|
Excretion |
Total of clearance (logmL/min/kg) |
0.571 |
0.873 |
0.931 |
0.283 |
0.655 |
0.637 |
Renal OCT2 Substrate
|
No |
No |
No |
No
|
No |
No |
|
CLplasma |
12.138 |
12.024 |
5.774 |
9.949 |
9.309 |
7.335 |
|
T1/2 |
1.114 |
1.686 |
0.624 |
0.242 |
0.289 |
0.679 |
|
Toxicity |
Ames Toxicity |
Yes |
No |
No |
No |
No |
No |
Conclusions
Four onoceranoid compounds were successfully isolated from the stem bark of Lansium domesticum Corr. cv. Piedjietan: methyl lansic (1), α,γ-onoceradienedione (2), methyl ester lansiolate (3), and lansiolic acid (4). Among them, methyl lansic (1) is a newly identified compound, while the other three have been previously reported within the Lansium genus but are described here for the first time in the Piedjietan cultivar. Notably, this study represents the first report of secondary metabolites isolated from the L. domesticum cv. Piedjietan, as no previous phytochemical investigations have been documented for this specific cultivar.
Molecular docking, molecular dynamics and ADMET analyses revealed that all four compounds exhibit strong binding affinities to the estrogen receptor alpha (ERα), primarily through hydrophobic interactions, and possess favorable pharmacokinetic properties, highlighting their potential as selective estrogen receptor modulators (SERMs). Molecular dynamics analysis confirms that the methyl lansic–ERα complex is structurally stable and displays properties indicative of a promising candidate for further evaluation in ERα-positive breast cancer models. Methyl lansic (1) also showed strong anticancer potential, exhibiting potent cytotoxicity against MCF-7 cells (IC₅₀ = 7.78 μM) while demonstrating no cytotoxic effect on normal cells (IC₅₀ = 620.30 μg/mL), reflecting an excellent selectivity profile.Although all compounds, exhibit minor violations of Lipinski’s Rule of Five, their ADMET profiles suggest they may offer safer and more stable therapeutic alternatives or adjuncts to tamoxifen in the treatment of estrogen receptor-positive breast cancer.
Given these findings, this study provides a coherent foundation for understanding the structural features that drive both cytotoxic and receptor-binding activities, thereby underscoring the relevance of onoceranoid triterpenoids as prospective anticancer agents. Notably, methyl lansic reported here for the first time both as a new compound and as a constituent of the Piedjietan cultivar, exhibited compelling potential as a therapeutic candidate for estrogen receptor-positive breast cancer. However, this study has several limitations. The biological evaluation was limited to a single cancer cell line, and no in vivo validation was performed to substantiate the pharmacological relevance of the observed activities. Additionally, although molecular docking, molecular dynamics and ADMET predictions provide valuable preliminary insights, they remain computational approximations that require experimental confirmation. To address these limitations, future research is warranted to elucidate their mechanisms of action, assess possible synergistic effects, and establish detailed structure–activity relationships, including in vivo evaluations and expanded pharmacokinetic and toxicity studies aimed at advancing methyl lansic as a viable lead compound.
Acknowledgements
This publication charge is funded by Unpad through the Indonesian Endowment Fund for Education (LPDP) on behalf of the Indonesian Ministry of Higher Education, Science and Technology and managed under the EQUITY Program (Contract No. 4303/B3/DT.03.08/2025 and 3927/UN6.RKT/HK.07.00/2025). This research was funded by Universitas Padjadjaran in the form of BUPP, Number: 993/UN6.3.1/PT.00/2025 and Kemendiksaintek PDD Number: 1692/UN6.3.1/PT.00/2025.
Conflict of interest
The author states that there is no conflict of interest.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on request
Declaration of Generative AI in Scientific Writing
During the writing of this manuscript, the authors used generative AI technologies to assist with language refinement, grammar correction, and the improvement of scientific clarity. The AI tools were employed solely for linguistics; all intellectual content, data interpretation, analysis, and conclusions were developed and critically reviewed independently by the authors. The authors take full responsibility for the integrity and originality of the content presented in this article.
CRediT Author Statement
Tri Mayanti: Conceptualization, methodology, supervision. Rika Septiyanti: Compound purification, structure elucidation, resources, investigation, writing of the original draft, data curation, and visualization. Selvi Apriliana Putri: In silico analysis, molecular docking, data curation, and visualization. Dilla Mardyana: Compound isolation, structure elucidation. Iman Permana Maksum: supervision. Sofa Fajariah: NMR measurement. Mohamad Azlan Nafiah: Review, supervision. Kindi Farabi: Review, supervision. Rani Maharani: Review and supervision. All authors approved the final version of this manuscript.
[1] R Yadav, A Pednekar, A Avalaskar, M Rathi and Y Rewachandani. A comprehensive review on meliaceae family. World Journal of Pharmaceutical Sciences 2015; 3(8), 1572-1577.
[2] K Yulita. Genetic variations of Lansium domesticum Corr. accessions from Java, Bengkulu and ceram based on random amplified ploymorhic DNA. Biodiversitas 2011; 12(3), 125-130.
[3] TK Lim. Edible medicinal and Non-Medicinal plants. Springer, Dordrecht, 2012, p. 265-268.
[4] T Mayanti, SE Sinaga and U Supratman. Phytochemistry and biological activity of Lansium domesticum Corr. species: A review. Journal of Pharmacy and Pharmacology 2022; 74(11), 1568-1587.
[5] R Septiyanti, R Maharani, MA Nafiah and T Mayanti. A review of onoceranoids from the Lansium and Lycopodium genera: phytochemistry and their biological activities. Trends in Sciences 2025; 22(10), 10509.
[6] S Omar, M Marcotte, P Fields, PE Sanchez, L Poveda, R Mata, A Jimenez, T Durst, J Zhang, S MacKinnon, D Leaman, JT Arnason and BJR Philogène. Antifeedant activities of terpenoids isolated from tropical Rutales. Journal of Stored Products Research 2007; 43(1), 92.
[7] T Mayanti, R Tjokronegoro, U Supratman, MR Mukhtar, K Awang and AHA Hadi. Antifeedant triterpenoids from the seeds and bark of Lansium domesticum cv Kokossan (Meliaceae). Molecules 2011; 16(4), 2785-2795.
[8] M Nishizawa, H Nishide and Y Hayashi. Total synthesis of (±)-α,γ-onoceradienedione and lansic acid. Tetrahedron Letters 1984; 25(44), 5071-5074.
[9] R Ramadhan, W Worawalai and P Phuwapraisirisan. Furofuran lignans as a new series of antidiabetic agents exerting α-glucosidase inhibition and radical scarvenging: Semisynthesis, kinetic study and molecular modeling. Bioorganic Chemistry 2019; 87, 783-793.
[10] T Mayanti, SA Azahra, TP Syafriadi, Nurlelasari, R Maharani, E Julaeha, U Supratman, SE Sinaga, S Ekawardhani, and S Fajriah. Onoceranoid triterpenes of Lansium domesticum Corr. cv. Kokossan and their cytotoxicity against MCF-7 breast cancer cells. Trends in Sciences 2025; 22(1), 9000.
[11] A Hardianto, SS Mardetia, W Destiarani, YP Budiman, D Kurnia and T Mayanti. Unveiling the anti-cancer potential of onoceranoid triterpenes from Lansium domesticum Corr. cv. kokosan: An in silico study against estrogen receptor alpha. International Journal of Molecular Sciences 2023; 24(19), 15033.
[12] T Mayanti, LD Ramadhanti, W Safriansyah, Darwati, Nurlelasari, D Harneti, SE Sinaga, E Bachtiar, S Hartati, S Fajriah and U Supratman. Lansioside E, an onoceranoid-type triterpenoid isolated from the fruit peel of Lansium domesticum Corr. cv. Kokossan and its cytotoxic activity against MCF-7 breast cancer cells. Journal of Asian Natural Products Research 2025; 27(11), 1720-1728.
[13] HM Abdallah, GA Mohamed and SRM Ibrahim. Lansium domesticum—A Fruit with Multi-Benefits: Traditional uses, phytochemicals, nutritional value, and bioactivities. Nutrients 2022; 14(7), 1531.
[14] T Matsumoto, T Kitagawa, S Teo, Y Anai, R Ikeda, D Imahori, HSB Ahmad and T Watanabe. Structures and antimutagenic effects of onoceranoid-type triterpenoids from the leaves of Lansium domesticum. Journal of Natural Products 2018; 81(10), 2187-2194.
[15] I Gupta, O Hussein, KS Sastry, S Bougarn, N Gopinath, E Chin-Smith, Y Sinha and HM Koras. Deciphering the complexities of cancer cell immune evasion: Mechanism and therapeutic implications. Advances in Cancer Biology - Metastasis 2023; 8, 100107.
[16] H Qayoom, M Alkhanani, A Almilaibary, SA Alsagaby and MA Mir. Mechanistic elucidation of juglanthraquinone c targeting breast cancer: A network pharmacology-based investigation. Saudi Journal of Biological Sciences 2023; 30(7), 103705.
[17] Mary Ann Liebert, Inc. Antisense & nucleic acid drug development. Mary Ann Liebert, Inc., New Rochelle, 2023.
[18] T Mayanti, Zulfikar, S Fawziah, AA Naini, R Maharani, K Farabi, Nurlelasari, M Yusuf, D Harneti, D Kurnia and U Supratman. New triterpenoids from Lansium domesticum Corr. cv kokossan and their cytotoxic activity. Molecules 2023; 28(5), 2144.
[19] M Nishizawa, H Nishide, S Kosela and Y Hayashi. Structure of lansiosides: Biologically active new triterpene glycosides from Lansium domesticum. The Journal of Organic Chemistry 1983; 48(24), 4462-4466.
[20] LJ Brandes. N,N-diethyl-2-[4-(phenylmethyl) phenoxy] ethanamine (DPPE; tesmilifene), a chemopotentiating agent with hormetic effects on DNA synthesis in vitro, may improve survival in patients with metastatic breast cancer. Human & Experimental Toxicology 2008; 27(2), 143-147.
[21] SA Putri, R Maharani, IP Maksum and TJ Siahaan. Peptide design for enhanced anti-melanogenesis: Optimizing molecular weight, polarity, and cyclization. Drug Design, Development and Therapy 2025; 19, 645-670.
[22] T Barkhem, B Carlsson, Y Nilsson, E Enmark, JÅ Gustafsson and S Nilsson. Differential response of estrogen receptor a and estrogen receptor 13 to partial estrogen agonists/antagonists. Molecular Pharmacology 1998; 54, 105-112.
[23] K Ramasamy, C Samayoa, N Krishnegowda and RR Tekmal. Abstract P1-04-01: Estrogen receptor β agonists inhibits syngeneic mammary tumor growth through cell-cycle arrest by modulating cell-cycle regulators. Cancer Research 2019; 79(S4), P1-04-01.
[24] CA Lipinski. Lead- and drug-like compounds: The rule-of-five revolution. Drug Discovery Today Technologies 2004; 1(4), 337-341.
Supplementary Material
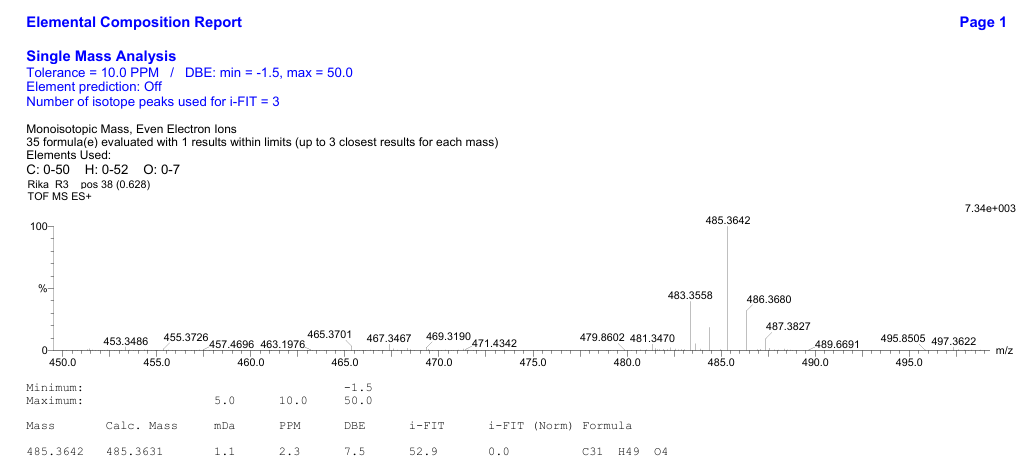
Figure S1 TOF-MS of Methyl lansic (Compound 1).
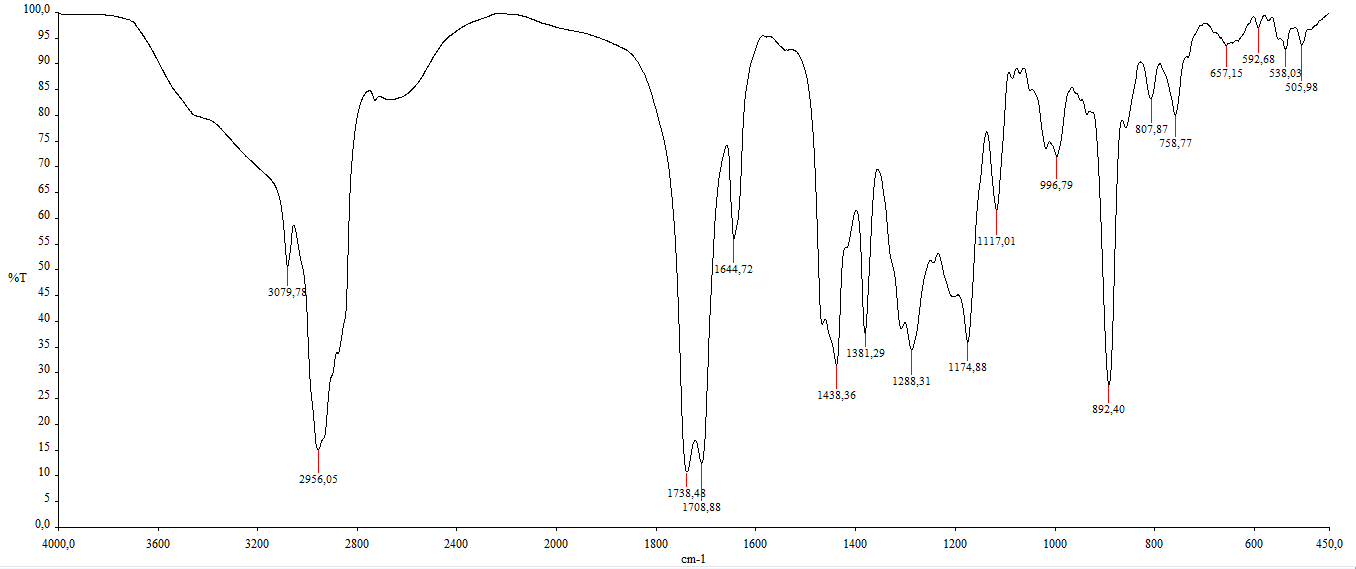
Figure S2 IR of Methyl lansic (Compound 1).
Figure S3 ¹H-NMR spectrum (CDCl₃, 500 MHz) of Methyl lansic (Compound 1).
Figure S4 ¹³C-NMR spectrum (CDCl₃, 125 MHz) and DEPT of Methyl lansic (Compound 1).
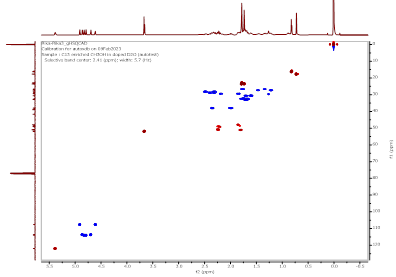
Figure S5 HSQC spectrum of Methyl lansic (Compound 1).
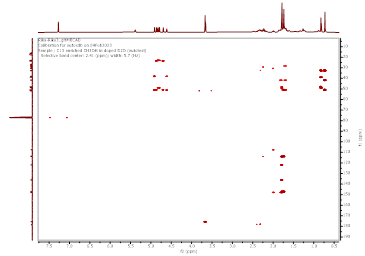
Figure S6 HMBC spectrum of Methyl lansic (Compound 1).
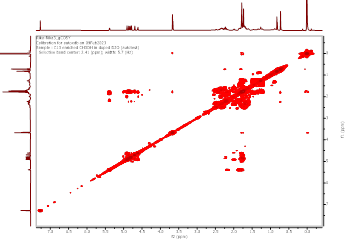
Figure S7 COSY spectrum of Methyl lansic (Compound 1).
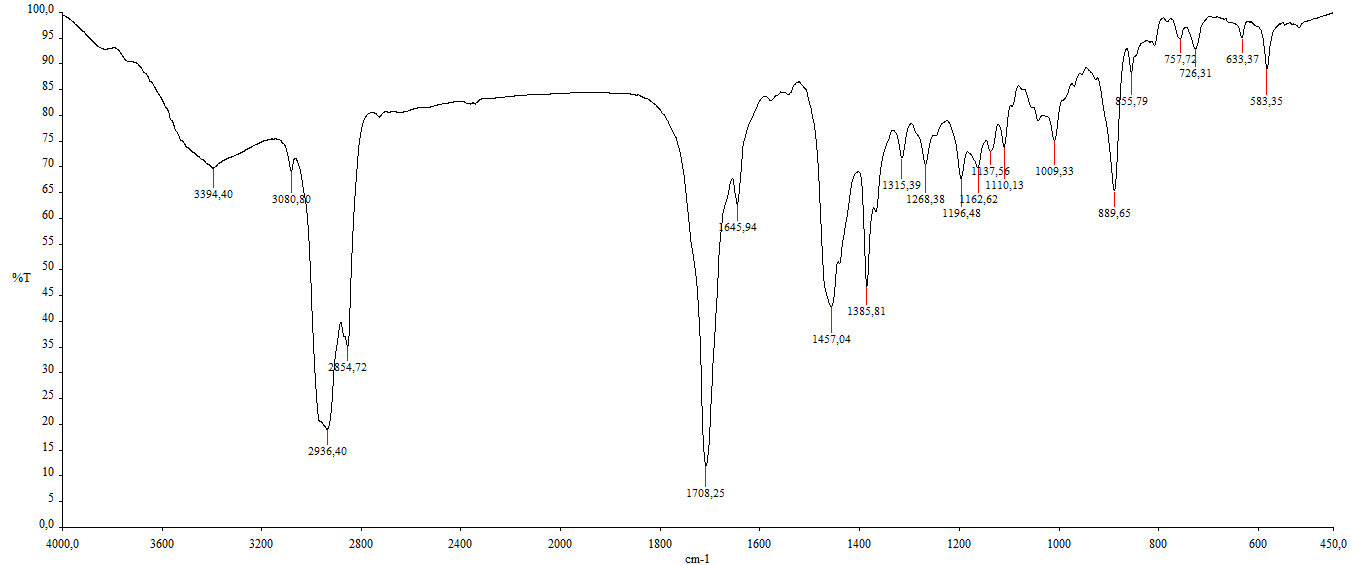
Figure S8 IR of α,γ-onoceradienedione (Compound 2).
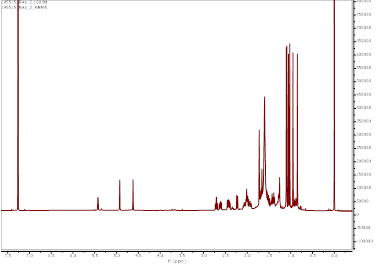
Figure S9 ¹H-NMR spectrum (CDCl₃, 700 MHz) of α,γ-onoceradienedione (Compound 2).
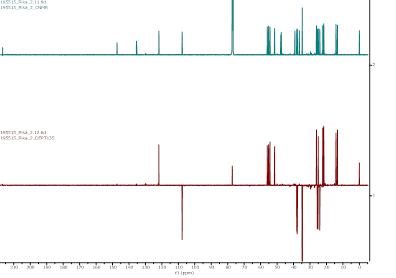
Figure S10 ¹³C-NMR spectrum (CDCl₃, 175 MHz) and DEPT of α,γ-onoceradienedione (Compound 2).
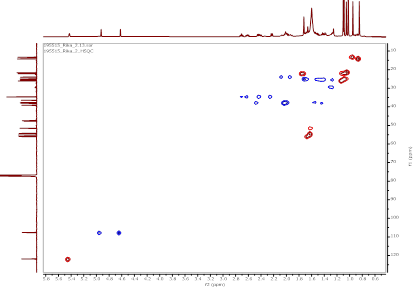
Figure S11 HSQC spectrum of α,γ-onoceradienedione (Compound 2).
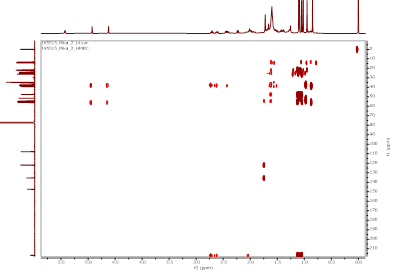
Figure S12 HMBC spectrum α,γ-onoceradienedione (Compound 2).
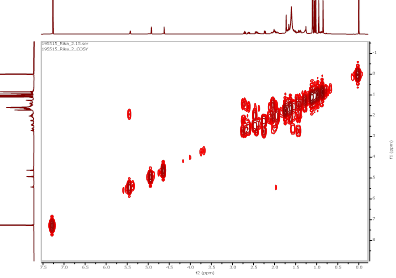
Figure S13 COSY spectrum of α,γ-onoceradienedione (Compound 2).
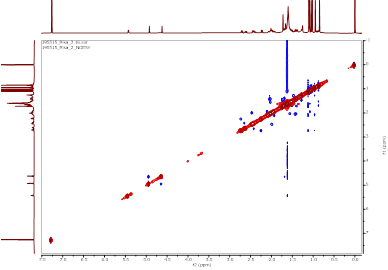
Figure S14 NOESY spectrum of α,γ-onoceradienedione (Compound 2).
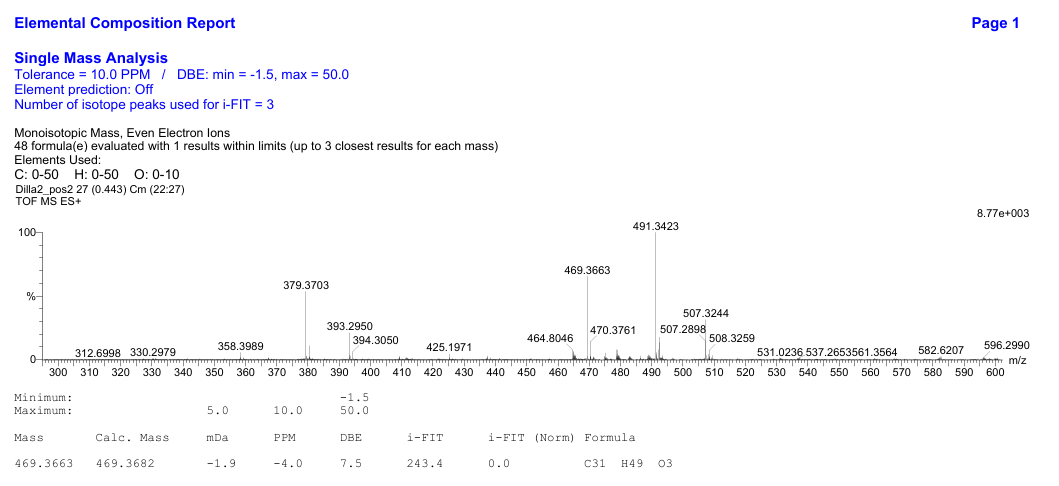
Figure S15 TOF-MS of methyl ester lansiolate (Compound 3).
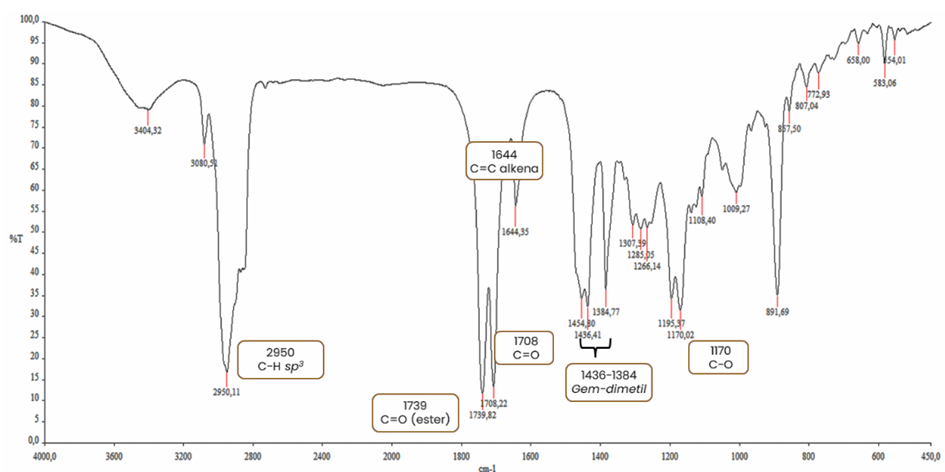
Figure S16 IR of methyl ester lansiolate (Compound 3).
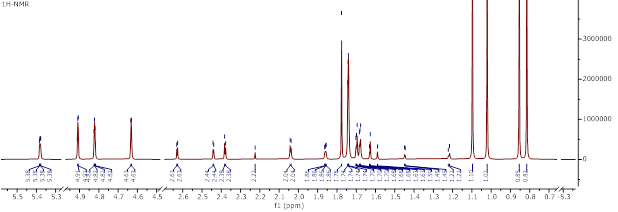
Figure S17 ¹H-NMR spectrum (CDCl₃, 700 MHz) of methyl ester lansiolate (Compound 3).
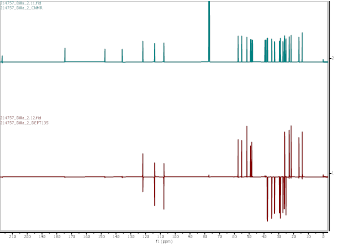
Figure S18 ¹³C-NMR spectrum (CDCl₃, 175 MHz) and DEPT of methyl ester lansiolate (Compound 3).
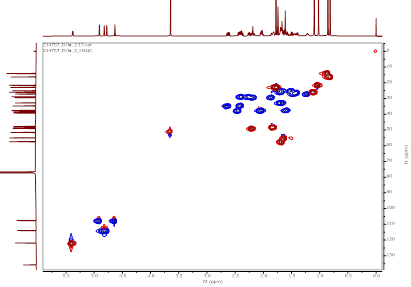
Figure S19 HSQC spectrum of methyl ester lansiolate (Compound 3).
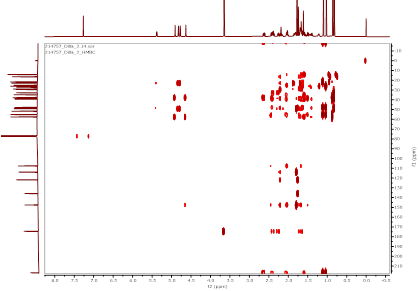
Figure S20 HMBC spectrum methyl ester lansiolate (Compound 3).
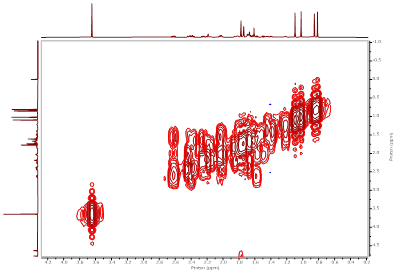
Figure S21 COSY spectrum of methyl ester lansiolate (Compound 3).
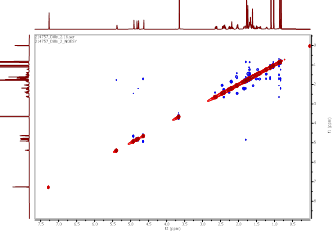
Figure S22 NOESY spectrum of methyl ester lansiolate (Compound 3).
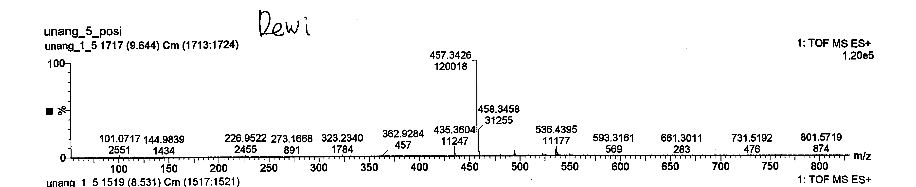
Figure S23 TOF-MS of lansiolic acid (Compound 4).
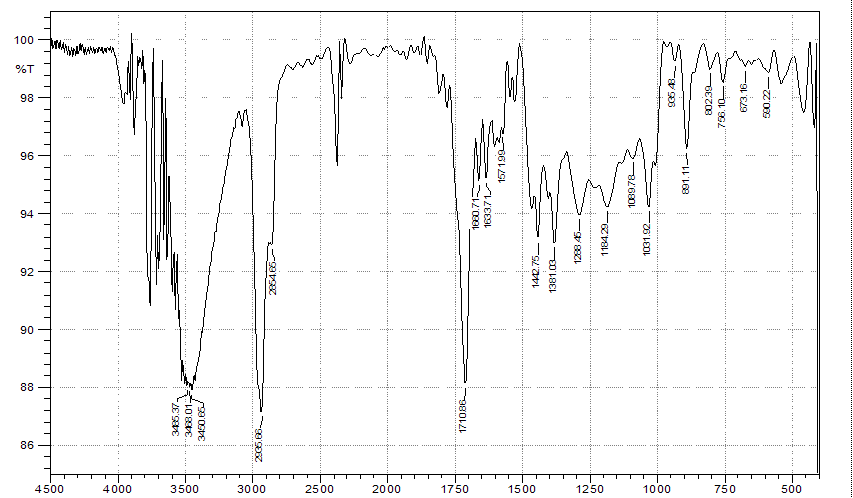
Figure S24 IR of lansiolic acid (Compound 4).
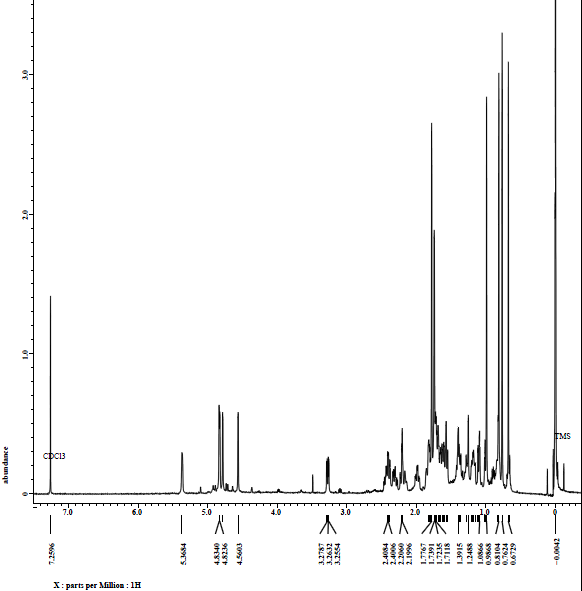
Figure S25 ¹H-NMR spectrum (CDCl₃, 500 MHz) of lansiolic acid (Compound 4).
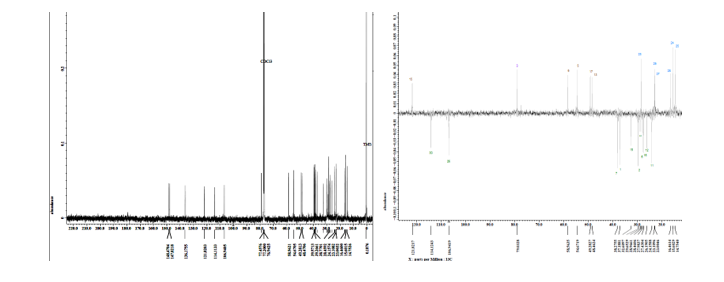 Figure
S26 ¹³C-NMR
spectrum (CDCl₃, 125 MHz) and DEPT
of
lansiolic acid (Compound 4).
Figure
S26 ¹³C-NMR
spectrum (CDCl₃, 125 MHz) and DEPT
of
lansiolic acid (Compound 4).
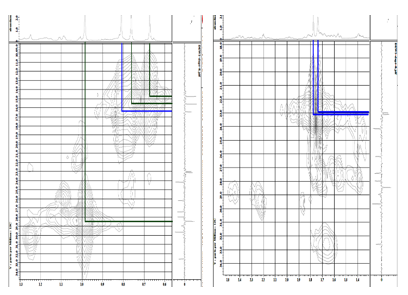
Figure S27 HMQC spectrum of lansiolic acid (Compound 4).
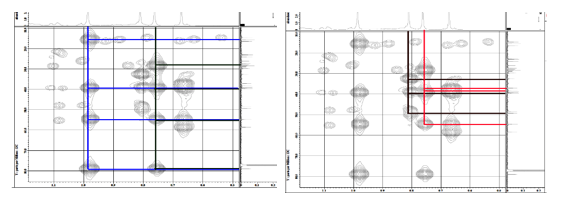
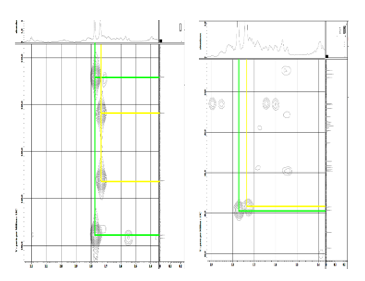
Figure S28 HMBC spectrum lansiolic acid (Compound 4).
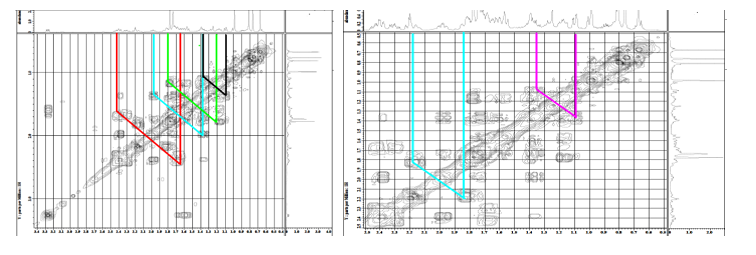
Figure S29 COSY spectrum of lansiolic acid (Compound 4).
In Vitro
Table S1 Cytotoxic activity analysis data of methyl lansic to MCF-7 in vitro
|
Media |
Media + test cell |
Cisplatin |
Solvent |
Concentration |
|||||||
1.17 |
2.34 |
4.69 |
9.38 |
18.75 |
37.50 |
75.00 |
150.00 |
|||||
Absorbance 570 nm
|
0.5776 |
0.9059 |
0.6749 |
0.8902 |
0.7246 |
0.7060 |
0.6958 |
0.5943 |
0.5386 |
0.5042 |
0.4963 |
0.4757 |
0.5746 |
0.9177 |
0.6743 |
0.9063 |
0.7257 |
0.7059 |
0.6959 |
0.5967 |
0.5386 |
0.5010 |
0.4965 |
0.4757 |
|
Absorbance 600 nm
|
0.7360 |
0.2166 |
0.5715 |
0.2675 |
0.2568 |
0.2673 |
0.3274 |
0.3754 |
0.3976 |
0.4077 |
0.4280 |
0.4469 |
0.7288 |
0.2257 |
0.5708 |
0.2778 |
0.2578 |
0.2672 |
0.3275 |
0.3757 |
0.3977 |
0.4078 |
0.4282 |
0.4469 |
|
Difference in absorbance |
−0.1584 |
0.6893 |
0.1034 |
0.6227 |
0.4678 |
0.4387 |
0.3684 |
0.2188 |
0.1410 |
0.0965 |
0.0683 |
0.0288 |
−0.1542 |
0.6920 |
0.1035 |
0.6285 |
0.4679 |
0.4387 |
0.3684 |
0.2210 |
0.1409 |
0.0933 |
0.0683 |
0.0288 |
|
% Live cells |
|
108.15 |
33.22 |
99.63 |
79.81 |
76.09 |
67.11 |
47.98 |
38.02 |
32.33 |
28.72 |
23.68 |
108.49 |
33.22 |
100.37 |
79.83 |
76.10 |
67.11 |
48.26 |
38.01 |
31.92 |
28.72 |
23.68 |
||
Average % of living cells |
|
108.32 |
33.22 |
100,00 |
79.82 |
76.10 |
67.11 |
48.12 |
38.01 |
32.12 |
28.72 |
23.68 |
SEM |
|
0.17 |
0.00 |
0.37 |
0.01 |
0.00 |
0.00 |
0.14 |
0.01 |
0.21 |
0.00 |
0.00 |
Normalization of % live cell data |
|
108.32 |
33.22 |
100.00 |
79.82 |
76.10 |
67.11 |
48.12 |
38.01 |
32.12 |
28. |
23.68 |
Note: The concentration of cisplatin used in the test was 43.18 µM.
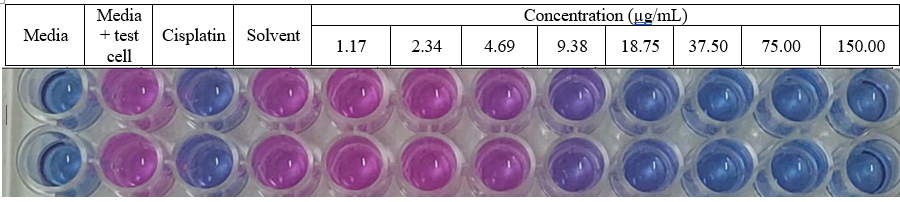
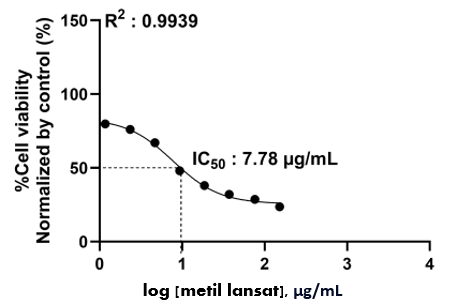

Figure S30 Cytotoxic acticity methyl lansic to MCF-7 (in vitro).
Table S2 Cytotoxic activity analysis data of methyl lansic to CV-1 in vitro
|
Media |
Media + test cell |
Cisplatin |
Solvent |
Concentration |
|||||||
7.81 |
15.63 |
31.25 |
62.50 |
125.00 |
250.00 |
500.00 |
1,000.00 |
|||||
Absorbance 570 nm
|
0.6370 |
0.7768 |
0.6328 |
0.7641 |
0.7798 |
0.7685 |
0.7579 |
0.7473 |
0.7367 |
0.7143 |
0.7013 |
0.6892 |
0.6348 |
0.7758 |
0.6202 |
0.7705 |
0.7797 |
0.7688 |
0.7579 |
0.7473 |
0.7367 |
0.7146 |
0.7013 |
0.6871 |
|
Absorbance 600 nm
|
0.7844 |
0.5678 |
0.6573 |
0.5516 |
0.5495 |
0.5535 |
0.5746 |
0.5954 |
0.6004 |
0.6241 |
0.6429 |
0.6797 |
0.7840 |
0.5537 |
0.6573 |
0.5515 |
0.5498 |
0.5533 |
0.5749 |
0.5953 |
0.6060 |
0.6248 |
0.6428 |
0.6798 |
|
Difference in absorbance |
−0.1474 |
0.2090 |
-0.0245 |
0.2125 |
0.2303 |
0.2150 |
0.1833 |
0.1519 |
0.1363 |
0.0902 |
0.0583 |
0.0095 |
−0.1492 |
0.2221 |
-0.0371 |
0.2190 |
0.2299 |
0.2155 |
0.1830 |
0.1520 |
0.1307 |
0.0898 |
0.0584 |
0.0073 |
|
%Cell Viability Normalized by Control (%) |
|
96.48 |
33.20 |
97.43 |
102.25 |
98.10 |
89.51 |
81.00 |
76.78 |
64.28 |
55.64 |
42.41 |
100.03 |
29.78 |
99.19 |
102.14 |
98.24 |
89.43 |
81.03 |
75.26 |
64.17 |
55.66 |
41.82 |
||
Average % Cell Viability Normalized by Control (%) |
|
98.25 |
31.49 |
98.31 |
102.20 |
98.17 |
89.47 |
81.02 |
76.02 |
64.23 |
55.65 |
42.11 |
SEM |
|
1.78 |
1.71 |
0.88 |
0.05 |
0.07 |
0.04 |
0.01 |
0.7 |
0.05 |
0.01 |
0.30 |
Note: The concentration of cisplatin used in the test was 39.48 µM.
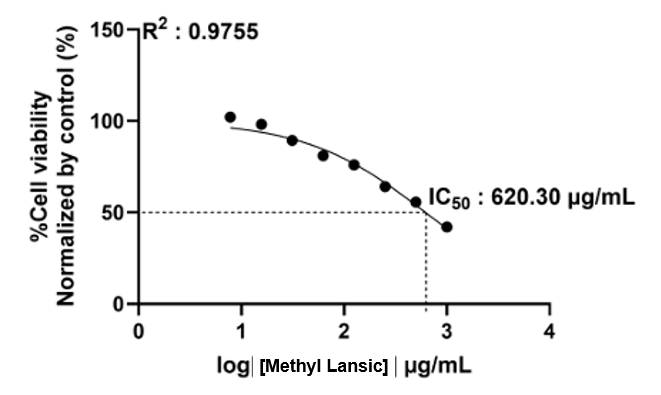
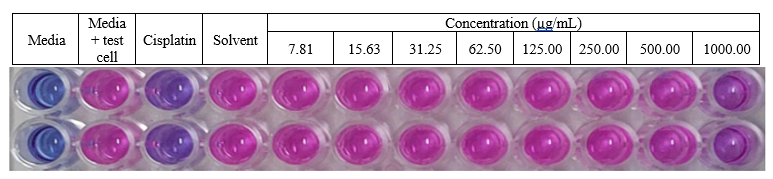
Figure S31 Cytotoxic acticity methyl lansic to normal cell CV-1 (in vitro).