
Trends
Sci.
2026;
23(7):
12467
Identification and Prediction of Bioactive Peptides from Anchovy (Engraulidae) Based on Processing Methods: Sub-Fragment Analysis via LC-HRMS Proteomics
Artika
Dewie1,
Susetyowati Susetyowati
2,*,
![]() and Diah
Rumekti Hadiati3
and Diah
Rumekti Hadiati3
1Doctoral Program in Medicine and Health Sciences, Faculty of Medicine, Public Health and Nursing,
Gadjah Mada University, Yogyakarta, Indonesia
2Department of Nutrition and Health, Faculty of Medicine, Gadjah Mada University, Yogyakarta, Indonesia
3Faculty of Medicine, Public Health and Nursing, Gadjah Mada University, Yogyakarta, Indonesia
(*Corresponding author’s e-mail: [email protected])
Received: 7 November 2025, Revised: 22 December 2025, Accepted: 29 December 2025, Published: 20 February 2026
Abstract
This exploratory study aimed to identify and compare peptide profiles and predicted bioactivities of anchovies (Engraulidae) in 3 forms - Raw, processed, and fortified processed - using LC HRMS based proteomics combined with BIOPEP UWM and PeptideRanker analyses. Peptides generated by tryptic digestion were analyzed by LC HRMS, matched to an Engraulidae protein database, and evaluated in silico to annotate potential biological activities and estimate intrinsic bioactivity scores. In this non replicated dataset, raw anchovies showed the highest proteomic complexity (254 protein IDs and 1,304 peptide sequences), whereas processed and fortified samples exhibited fewer proteins and peptides, particularly with chain lengths ≥ 4 amino acids. Across all samples, the most frequent predicted activities were Dipeptidyl Peptidase IV (DPP IV) (27% - 31%) and Angiotensin Converting Enzyme (ACE) inhibition (± 24%), supported by antioxidant, neprilysin inhibitory (3% - 6%), and other enzyme inhibitory annotations. Short peptides (2 - 3 amino acids) dominated the predicted bioactive profiles, especially in processed and fortified products, indicating a shift toward shorter chains. Although total peptide numbers decreased after processing, the proportion of peptides with multiple annotated activities and high PeptideRanker scores (≥ 0.8) was descriptively higher in fortified samples (14.58%). These findings provide a preliminary molecular map suggesting that anchovy proteins can act as precursors of peptides with putative multifunctional bioactivities and may represent candidates for anchovy based functional foods or nutraceuticals. However, all functional inferences are based solely on sequence level, in silico predictions from single run LC HRMS data; the patterns observed are hypothesis generating and require confirmation through replicated experiments and targeted in vitro and in vivo validation.
Keywords: Anchovies, Bioactive peptides, LC-HRMS, Fortification, BIOPEP, PeptideRanker
Introduction
Functional foods are defined as foods that are consumed as part of the daily diet and provide health benefits beyond their basic nutritional functions [1]. These effects are primarily mediated by bioactive components that work through specific physiological mechanisms, including the prevention and management of various chronic conditions [2,3]. In the last 2 decades, scientific attention has intensively focused on bioactive peptides, which are short fragments of amino acids
stored in parent protein structures and can be released through proteolytic processes [4].
Bioactive peptides generally consist of 2 - 20 amino acid residues, and their biological activity is strongly influenced by their sequence, chain length, and chemical structure [5]. Specific interactions between peptides and molecular targets - such as enzymes or cell receptors [6] - underlie their various biological activities. In the context of animal-based foods, marine fish, particularly anchovies, represent a rich and promising source of bioactive peptides, as numerous studies have reported the antioxidant, anti-inflammatory, antihypertensive, antimicrobial, antidiabetic, and anticancer properties of hydrolyzed fish proteins (7 - 9). The release and stability of these peptides can occur during gastrointestinal digestion, in vitro hydrolysis, or as a consequence of food processing [10].
In the context of protein-based foods, particularly fish and fishery products, the identification of bioactive peptides is increasingly being carried out using a mass spectrometry-based proteomics approach [11]. Techniques such as liquid chromatography-tandem mass spectrometry (LC-MS/MS) [12] and liquid chromatography-high-resolution mass spectrometry (LC‑HRMS) [13] enable the identification of amino acid sequences, molecular masses, and protein origins with high precision from complex food matrices [12]. LC-HRMS specifically offers the advantages of exact mass accuracy and high resolving power, making it highly relevant for peptide mapping in real food systems [13]. This approach has been applied to various marine organisms and fish, including mesopelagic fish (14), anchovies [15], herring [16], and octopus [17], generally in combination with bioinformatics analysis to predict the potential biological activity of the identified peptides.
However, most fish proteomics studies still focus on enzymatic hydrolysates [18,19] or specially conditioned fermentation products [20]. These approaches provide important insights, but do not fully represent the conditions of ready-to-eat fish products produced through household processing, such as frying. These processes are known to modulate peptide release patterns during digestion and proteomic analysis [21]. Therefore, direct peptide mapping from processed fish products is important to understand the real impact of processing on bioactive peptide profiles. This limitation is particularly relevant for ready-to-eat fish products commonly consumed by the public.
Anchovies (Engraulidae) are small pelagic fish that are abundant in Indonesian waters and are known to be rich in macro- and micronutrients [22,23], and contain complete and easily absorbed essential and non-essential amino acids [15, 24]. Several previous studies have reported the presence of bioactive peptides in fresh anchovies [15] and in traditional fermented products [24,25]. However, these studies were generally conducted separately and focused on one form of material, without systematically comparing the peptide profiles of raw anchovies, processed anchovies, and fortified anchovies in an integrated research design. Micronutrient fortification also has the potential to modify the matrix properties of the product and the interactions between its components. Fortification with lipophilic micronutrients such as cholecalciferol (vitamin D3) is generally carried out using a carrier or encapsulation system to increase stability against heat, oxygen, and light [26].
The selection of cholecalciferol as a fortifier also has a nutritional basis, given that anchovies are known as a high source of calcium [27], while vitamin D3 plays a role in supporting calcium absorption in the body [28,29]. Fortification is primarily seen as a factor that has the potential to modify the food matrix and protein environment. The presence of carriers and accompanying changes in processing conditions may affect the stability and profile of peptides formed or detected. To date, studies that comparatively assess the effects of processing and fortification on the bioactive peptide profile of fish at the molecular level are still very limited.
Food processing methods such as heating, drying, and frying can alter the structure of fish protein through denaturation, aggregation, and oxidation [30,31,32]. These structural changes can affect the exposure of protein cleavage sites [31,32], thereby altering the release pattern and stability of peptides [30]. Variations in the sequence and length of the resulting peptides then become the basis for predicting the potential for certain biological activities, such as enzyme inhibition activity, through amino acid sequence-based in silico analysis. However, the predicted biological activity does not necessarily reflect the bioavailability and physiological effects of peptides in the body, as the processes of digestion, absorption, and metabolism have not been evaluated in this study. Therefore, this study focuses on molecular-level peptide mapping as a preliminary step prior to further biological evaluation.
Based on this, there is still a clear knowledge gap regarding the mapping of bioactive peptides in Indonesian anchovies in various forms of processing - raw, processed, and fortified - using the LC-HRMS approach. In particular, there are no reports that systematically compare the number and types of proteins and peptides, the distribution of peptide chain lengths, the variety of predicted bioactive activities, and in silico bioactivity scores within a consistent analytical framework.
Therefore, this study aims to identify and compare the bioactive peptide profiles in raw anchovies, processed anchovies, and fortified anchovies using an LC-HRMS proteomics approach integrated with bioinformatics analysis (BIOPEP and PeptideRanker). This study was designed as an in silico-based exploratory mapping to provide an initial molecular overview of the impact of processing and fortification on anchovy peptide profiles. The findings are expected to provide a rational basis for further research, including in vitro and in vivo validation and process optimization, before health implications and functional food applications can be empirically confirmed.
Materials and methods
Study design
To address the research gap regarding changes in the profile of bioactive peptides caused by anchovy processing and fortification, this study used a comparative and exploratory proteomic approach based on LC-HRMS (Liquid Chromatography-High Resolution Mass Spectrometry), which enables high-precision peptide identification. Three sample groups were analyzed: raw anchovies, processed anchovy products, and processed and fortified anchovy products. This approach enabled a systematic comparison of changes in peptide complexity between sample groups, as well as an evaluation of the predicted bioactivity of peptides in silico as a result of processing and fortification at the molecular level. The study design was non-inferential and did not include biological or technical replication, thus focusing on preliminary molecular mapping - including the number of identified proteins and peptides, peptide length distribution, and predicted bioactivity diversity and scores - without intending to test for statistical differences between groups.
Samples and treatment
Anchovies (family Engraulidae) were obtained from a local market in Palu City, Central Sulawesi (Mamboro Market), which originated from the coastal fishing area of Mamboro Beach. All samples were obtained at the same time of purchase to minimize raw material variation. Three groups of samples were analyzed, namely: (1) raw anchovies, (2) Product processed anchovies, and (3) Fortified processed anchovies.
Raw samples were stored at −18 °C, while processed samples were packaged in airtight containers and stored at room temperature away from direct sunlight for one day before being sent to the laboratory for LC-HRMS analysis. Processed products were prepared by mixing anchovies with flour and spices.
The proportions of flour and spices were not quantitatively determined but were adjusted until an even coating was obtained and the anchovies did not stick together. Next, drying was carried out using a food dehydrator at 40 °C for 1 h to reduce the surface moisture content. Frying was carried out in hot cooking oil until golden brown and crispy in texture.
The fortified product follows the same procedure, but is formulated with 600 IU (15 µg) of cholecalciferol (vitamin D₃) powder per serving (± 40 g of product). This study is not intended to evaluate the effectiveness of fortified nutrition, but rather to compare the peptide profiles at the molecular level between sample groups.
LC-HRMS preparation and analysis
Proteomic analysis was performed at the Integrated Research and Testing Laboratory (LPPT), Gadjah Mada University, Yogyakarta. Peptide identification was performed using a Thermo Scientific™ Dionex Ultimate 3000 RSLCnano UHPLC system connected to a Thermo Scientific™ Q Exactive™ (Orbitrap) high-resolution mass spectrometer. Ionization was performed in positive mode using an EASY-NanoSpray ionization source. Each sample (± 15 g) was analyzed using a bottom-up proteomics approach.
Proteins were first extracted from solid samples, then underwent reduction and alkylation before enzymatic digestion using trypsin (Thermo Scientific), according to standard laboratory procedures. The digest filtrate was then filtered using a 0.22 µm membrane to remove particulate contaminants. A 5 µL aliquot of the filtrate was injected into the LC-HRMS system for analysis.
Peptide separation was performed using an EASY-Spray PepMap C18 column (15 cm×75 µm ID, particle size 3 µm). The mobile phase consisted of MS-grade water containing 0.1% formic acid (phase A) and MS-grade acetonitrile containing 0.1% formic acid (phase B). Peptides were eluted using the following linear gradient: 5% phase B for 1 min, followed by a gradual increase from 5% to 50% phase B over 50 min, then returned to the initial conditions, with a total analysis time of 60 min. The nanoLC flow rate was set at 100 nL/min, and the column temperature was maintained at room temperature.
Mass spectra were collected using data-dependent acquisition (DDA) mode. Full MS scanning was performed at a resolution of 70,000 (FWHM), followed by MS/MS scanning at a resolution of 17,500 (FWHM). The mass scan range was set at m/z 150 - 2,000. Ion fragmentation was performed using higher-energy collisional dissociation (HCD) with a normalized collision energy (NCE) of 27. The spray voltage was set to approximately 3.5 kV, with a capillary temperature of 250 °C. The automatic gain control (AGC) and maximum injection time parameters were run using the Orbitrap instrument default settings.
The mass tolerance was set at ±10 ppm for precursor ions and ± 0.02 Da for fragment ions, in accordance with the Orbitrap instrument specifications. Some highly specific instrument parameters (e.g., target AGC, maximum number of ions selected per cycle, and maximum injection time) follow the standard settings of the Q Exactive instrument in the service laboratory and are not fully listed in the technical report, but are within the range commonly used for LC-HRMS-based peptide analysis.
Protein identification and reference database
Protein identification was performed using Proteome Discoverer 2.2 software (Thermo Fisher Scientific). Protein sequence matching was performed against a database downloaded from UniProt (available at the time of analysis), with trypsin set as the proteolytic enzyme and up to 2 missed cleavages allowed. Validation of protein identification results was performed using the Percolator module, with a false discovery rate (FDR) < 1% applied at the peptide spectrum match (PSM) and protein group levels. Search parameters included a mass tolerance of approximately ±10 ppm for precursor ions and ±0.02 Da for fragment ions, in accordance with Orbitrap instrument specifications and standard proteomics practices. Since the analysis was performed in an external laboratory, highly detailed instrument parameters were not fully available in the technical report, but the settings used followed software standards and MIAPE (Minimum Information About a Proteomics Experiment) guidelines.
Analysis bioinformatics
Identified peptides were analyzed in silico using 2 bioinformatics platforms: BIOPEP-UWM and PeptideRanker. BIOPEP-UWM was used to identify bioactive motifs and map their potential biological activities, such as antihypertensive, antioxidant, antidiabetic, and immunomodulatory effects. PeptideRanker was used to predict peptide bioactivity scores based on a machine-learning model, with values ranging from 0 to 1. Peptides with scores ≥ 0.8 were categorized as highly bioactive, those between 0.5 - 0.79 as moderately bioactive, and those < 0.5 as having low bioactivity. In this study, the BIOPEP-UWM and PeptideRanker results were used solely as exploratory in silico indicators of potential bioactivity and were not treated as confirmatory proof of real biological function.
Criteria peptides and classification
Peptides were classified based on 5 main parameters: (1) Number of Proteins and Peptides identified, (2) Types of biological activity (BIOPEP), (3) Peptide length (number of amino acids residues), (4) Number of biological activities per peptide and (5) Predicted bioactivity score (PeptideRanker).
The distribution of peptides according to these parameters was analyzed for each treatment (raw, processed, and fortified). The main results are presented in the primary tables of this article, while the complete peptide list is available in the supplementary data file.
Data analysis and visualization
This study represents an exploratory effort to establish an initial map of bioactive peptides in anchovies across 3 forms - raw, processed, and fortified. Data were presented in tabular form for each sample, covering the number of identified peptides, types of biological activities, peptide length distribution, number of activities per peptide, and predicted bioactivity scores.
This study was exploratory and non-inferential, as the proteomic analyses were conducted externally, and replication data for biological or technical approaches were unavailable. Such an approach is common in the initial stages of untargeted proteomics research, where the goal is to generate a preliminary mapping of bioactive peptide profiles. To ensure data reliability, protein identification parameters such as sequence coverage, number of unique peptides, and total PEP scores were applied as criteria for internal validation. The interpretation of results focused on peptide trends, distribution, and qualitative-quantitative characteristics rather than inferential statistical differences between treatments.
Research ethics
All research procedures, including sample collection, processing, and laboratory analyses, received ethical approval from the Research Ethics Committee of Gadjah Mada University, Yogyakarta, Indonesia (Ref. No: KE/FK/0618/EC/2025). The study was conducted in accordance with ethical principles for scientific research to ensure data integrity and the responsible use of biological materials.
Identification of parent peptides in raw, processed, and fortified anchovy samples
Proteomic analysis was conducted to describe the peptide profiles of raw anchovies and explore how processing and fortification might affect peptide composition, without intending to inferential test treatment effects. Three types of samples - raw anchovies, processed anchovies, and fortified processed anchovies - were analyzed using the LC-HRMS approach to identify parent peptides resulting from protein degradation. These peptides are considered as molecular components that potentially contribute to the functional characteristics and predicted bioactivity of each sample in the context of this exploratory analysis.
Identification was performed by matching peptide sequences against the Engraulidae Coilia grayii FASTA database, producing quantitative data in the form of protein IDs, total identified peptide sequences, and unique peptides.
These identification results form the descriptive basis for in silico bioactivity analysis in the next stage and are presented in tables and diagrams to illustrate the distribution patterns and differences observed between samples, without implying statistically validated differences.
Table 1 Identification of parent peptides in raw, processed, and fortified anchovy samples.
No |
Material |
Raw |
Processed |
Fortified |
1 |
Accession / ID protein |
254 |
30 |
25 |
2 |
Sequence Peptides identified |
1304 |
134 |
98 |
3 |
Peptides unique |
662 |
51 |
48 |
Data source: LC-HRMS analysis (LPPT UGM, 2025).
Table 1 summarizes the number of protein IDs (accessions), identified peptide sequences, and unique peptides specific to each sample type. The raw sample exhibited the highest proteomic complexity, with 254 protein IDs and 1,304 identified peptide sequences. Among these, 662 were unique peptides not found in either the processed or fortified samples. In contrast, the processed and fortified samples showed a reduction in both protein and peptide counts - 30 and 25 protein IDs, and 134 and 98 peptide sequences, respectively. The number of unique peptides was also lower, with 51 in the processed and 48 in the fortified samples. This pattern is interpreted as a descriptive trend in the dataset without replication, and not to draw conclusions about the effects of processing or fortification.
The identification of protein and peptide numbers from LC-HRMS results provides essential information on the types and characteristics of proteins present in each sample, allowing for the evaluation of both the quantity and quality of identified proteins [33]. This step is crucial for improving data reliability and building a comprehensive proteomic profile, especially when comparing different samples, such as raw, processed, and fortified anchovies.
Different treatments can theoretically affect protein structure and produce variations in in the proteomic composition of each sample [25]. The greater the number of proteins and peptides successfully identified, the more comprehensive the understanding of protein composition within a biological sample becomes. This is crucial for exploratory comparative analysis and for gain initial insights into the potential functional properties of the resulting proteins and peptides [23].
The identified proteins most likely originate from the muscle tissues of anchovies, which is widely recognized as a source of easily digestible protein and is rich in essential amino acids [15]. Previous studies have reported that enzymatic hydrolysis of fish muscle proteins can break peptide bonds and release peptides that have various functional properties [35,36], and bioactive potential such as antihypertensive (ACE inhibitor), antioxidant, antiproliferative, and antimicrobial activities [37]. Bioactive peptides can be derived from several fish parts, including muscle, collagen, gelatin from the skin, and bones [29]. However, in this study, these activities were not evaluated in vitro or in vivo. Therefore, this information is used solely as contextual literature and should not be interpreted as direct evidence of the bioactivity of the identified peptides.
Peptides identified through LC-HRMS analysis form the basis for protein identification [39]. Among these, unique peptides - those associated with only one protein or specific protein group without ambiguity - are particularly important [40], because they increase confidence in protein identification, reduces the risk of false positives, and strengthens automated validation. Conversely, proteins identified solely from non-unique peptides require more careful interpretation, as they may lead to less accurate identification [39].
However, not all identified proteins provide relevant information on bioactivity. Therefore, the selection of representative protein IDs was based on key criteria: A minimum of 2 unique peptides, sequence coverage, and a cumulative peptide score ≥ 10%. The presence of at least 2 unique peptides is a stronger indicator of protein identification validity, improving analytical accuracy and reliability [41]. Meanwhile, a cumulative peptide score ≥ 10% reflects high confidence in the mass spectrum matching, and sequence coverage between 20% - 30% or higher is considered sufficient for exploratory proteomics studies [32]. These thresholds help minimize reporting that proteins identified solely due to partial or coincidental matches.
Table 2 summarizes several representative proteins identified in the 3 sample types - raw, processed, and fortified anchovies - which were selected for their predicted major contributions to peptide complexity and potential bioactivity.
Table 2 Identification of Featured Protein IDs based on LC-HRMS analysis on raw, processed, and fortified anchovies.
No |
Sample |
Unique peptides |
Coverage (%) |
SUM Peptide score |
1 |
Raw Anchovies |
|
|
|
|
|
22 |
32.24 |
94.54 |
|
|
13 |
59.51 |
93.06 |
|
|
10 |
35.43 |
107.06 |
|
|
8 |
31.96 |
117.01 |
|
|
14 |
25.18 |
88.82 |
|
|
7 |
20.97 |
31.99 |
|
|
16 |
28.57 |
51.62 |
|
|
8 |
21.97 |
51.6 |
|
|
3 |
26.13 |
56.41 |
|
|
3 |
29.97 |
47.15 |
2 |
Product processed anchovies |
|||
|
|
8 |
33.80282 |
27.161 |
|
|
3 |
13.43511 |
61.096 |
|
|
2 |
10.30397 |
58.738 |
3 |
Fortified processed anchovies |
|||
|
|
8 |
21.12676 |
14.701 |
|
|
2 |
10.2799 |
32.157 |
|
|
4 |
9.47965 |
30.765 |
Data source: LC-HRMS analysis (LPPT UGM, 2025).
Based on the selection criteria described above, ten key proteins were identified in raw anchovies, and 3 representative proteins each were identified in the processed and fortified products. The raw samples showed the highest proteomic complexity, with 3 - 22 unique peptides per protein. Among these, protein A0ABD1K6P1 had the highest number of unique peptides, A0ABD1JQU7 exhibited the highest sequence coverage, and A0ABD1KC28 showed the highest cumulative peptide score. In the processed and fortified samples, the number of protein IDs decreased, but A0ABD1JQU7 remained detectable, although with lower unique peptide counts and sequence coverage. Proteins A0ABD1JWL9 and A0ABD1J939 were identified with more limited representation.
The reduction in unique peptide counts, sequence coverage, and peptide scores in the processed and fortified samples may reflect protein degradation or structural modifications occurring during processing and fortification, which in turn could affect the detectable peptides. Previous studies have shown that treatments such as heating, fermentation, and fortification can lead to protein denaturation, aggregation, and fragmentation, thereby reducing proteome complexity [33]. Other studies also noted that suboptimal processing can diminish peptide stability and biological activity through degradation or modification [44]. Descriptively, the patterns observed in this study are consistent with these findings, although they cannot be used to establish a causal relationship between the treatments and changes in peptide quantity, diversity, or potential bioactivity.
Therefore, protein A0ABD1JQU7 can be tentatively considered as a candidate protein marker that
remains relatively stable under the various processing and fortification conditions tested in this study. Several other proteins were also detected across all sample types but were not included in Table 2 as they did not meet the established threshold for representation. Complete identification data are provided in Supplementary File to ensure transparency and allow further evaluation.
Distribution of biological activity types in bioactive peptides
Following the identification of primary protein IDs through LC-HRMS analysis, the resulting peptides were further examined to evaluate their potential bioactivities in silico. Peptide sequences were matched against the BIOPEP database to identify previously reported bioactive motifs, allowing each peptide to be classified into established biological activity categories. The analysis focused on the distribution of these predicted biological activity types, including antihypertensive, antioxidant, antidiabetic, immunomodulatory, and enzymatic inhibitory activities.
To streamline data presentation, the nine most abundant bioactive peptides from each sample were listed in the main table, along with one combined category labeled “other.” These bioactive peptides were selected based on their frequency of occurrence and activity type identified via BIOPEP matching. The distribution of bioactive peptide activities among raw, processed, and fortified anchovy samples is presented in Table 3 to illustrate both quantitative and proportional differences patterns observed between sampels in this exploratory, non‑replicated dataset, without statistical testing.
Table 3 Distribution of biological activity peptides bioactives identified in raw, processed, and fortified anchovy samples.
No |
Activity |
Sample |
|||||
Raw |
Processed |
Fortified |
|||||
Σ |
% |
Σ |
% |
Σ |
% |
||
1 |
Dipeptidyl Peptidase IV Inhibitor (DPP- IVi) |
8,872 |
31.47 |
865 |
27.48 |
703 |
27.48 |
2 |
ACE Inhibitor (ACEi) |
6,877 |
24.39 |
766 |
24.00 |
614 |
24.00 |
3 |
Antioxidant (Aox) |
1,455 |
5.16 |
167 |
5.36 |
137 |
5.36 |
4 |
Neprilysin Inhibitor (NEPi) |
1,215 |
4.31 |
216 |
6.53 |
167 |
6.53 |
5 |
Glutamate Carboxypeptidase II Inhibitor (GCP- IIi) |
1,062 |
3.77 |
205 |
5.75 |
147 |
5.75 |
6 |
Stimulating (Stim) |
933 |
3.31 |
130 |
3.48 |
89 |
3.48 |
7 |
Dipeptidyl Peptidase III Inhibitor (DPP-IIIi) |
913 |
3.24 |
80 |
2.89 |
74 |
2.89 |
8 |
Glutamate Carboxypeptidase Inhibitor (GCP- i) |
688 |
2.44 |
143 |
3.28 |
84 |
3.28 |
9 |
Inhibitor Of Tripeptidyl Peptidase II (TPP- IIi) |
596 |
2.11 |
43 |
1.76 |
45 |
1.76 |
10 |
Miscellaneous (Misc) |
5,582 |
19.80 |
682 |
19.47 |
498 |
19.47 |
|
Total |
28,193 |
100 |
3,297 |
100 |
2,558 |
100 |
Data source: LC-HRMS analysis (LPPT UGM, 2025).
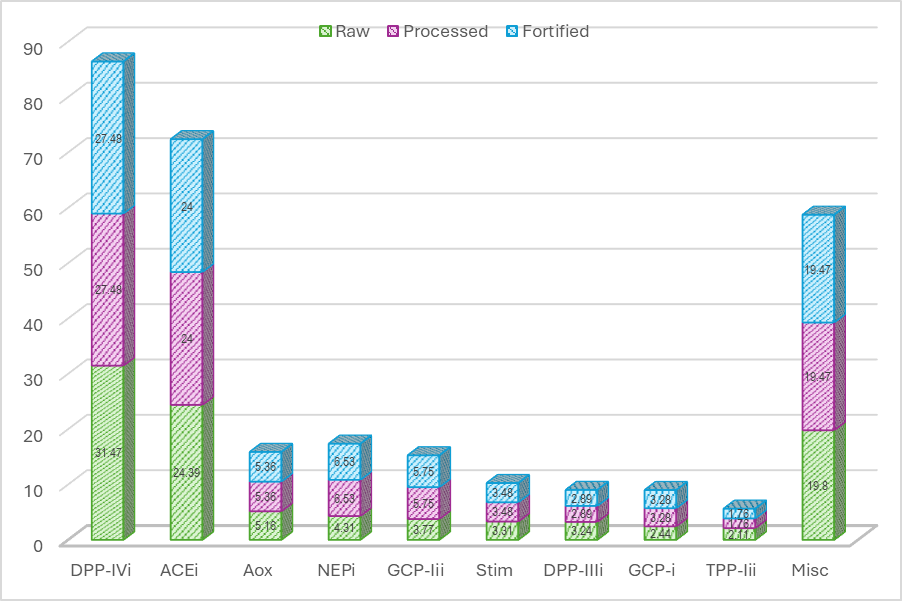
Figure 2 Distribution of biological activity peptides bioactives identified in raw, processed, and fortified anchovy samples.
Data source: LC-HRMS analysis (LPPT UGM, 2025).
The results show that although the proportional distribution of predicted bioactive activities remained broadly similar across the 3 samples, the total peptide counts decreased following processing and fortification. The dominant activity category across all groups was Dipeptidyl Peptidase IV (DPP-IV) inhibition, with the highest proportion in raw anchovies (31.47%), followed by processed and fortified anchovies (27.48%). In previous studies, DPP-IV inhibitory peptides have been reported to regulate glucose metabolism by preventing the degradation of incretin hormones that stimulate insulin secretion, thereby improving glycaemic control and offering potential benefits for the prevention and management of type 2 diabetes [45-47].
ACE inhibitor activity was also constituted a consistently large proportion in all samples, ranging from 24.0% to 24.39%, suggesting that, at the descriptive level, processing and fortification did not markedly alter the proportion of peptides annotated as ACE inhibitors in this dataset. Anchovies, as marine organisms, have been widely reported as source of ACE inhibitory peptides with potential applications in pharmaceutical and nutraceutical development [48]. ACE (angiotensin-converting enzyme) converts angiotensin I into angiotensin II, a vasoconstrictor that raises blood pressure. By inhibiting ACE, these peptides help reduce angiotensin II formation, thereby lowering blood pressure [49,50].
Neprilysin inhibitor activity showed moderate levels, with a slightly higher proportion in fortified samples (6.53%) compared to raw samples (4.31%). Neprilysin is involved in the metabolism of natriuretic peptides, insulin, and enkephalins, and its inhibition has been linked in the literature to potential benefits for cardiovascular regulation and protection [51]. Previous studies have suggested that combined inhibition of neprilysin and the angiotensin II receptor system is associated with improved glycemic control and insulin sensitivity potentially through increased levels of beneficial peptides such as bradykinin and natriuretic peptides, which play crucial roles in glucose metabolism and cardiovascular health [52].
Other activities, including glutamate carboxypeptidase II inhibition (3.77% - 5.75%), antioxidative activity (5.16% - 5.36%), and stimulating activities (3.31% - 3.48%), were detected across all sample types. The moderate yet relatively consistent proportions of these predicted activities indicate that core categories of bioactive peptide annotations are retained despite differences in peptide composition and abundance among samples. According to previous reports, peptides annotated as Glutamate carboxypeptidase II inhibitors have been associated with potential therapeutic relevance in neurological disorders [53], while peptides annotated with antioxidant activity have been reported to contribute to cellular protection against oxidative stress through free radical scavenging and inflammation-related pathways [54]. In addition, peptides classified as stimulating have been described in the literature as being involved in regulatory processes related to gene expression, cell proliferation, differentiation, tissue repair, and the modulation of degenerative processes [55].
The lowest proportions were observed for glutamate carboxypeptidase inhibitors, dipeptidyl peptidase III inhibitors, and tripeptidyl peptidase II inhibitors, each contributing less than 3.5%. The “other” category accounted for approximately 19% - 20% of total activity, encompassing enzymatic or biological functions outside the main categories.
Overall, the findings indicate that peptides annotated as DPP-IV and ACE inhibitors represent the predominant predicted bioactive categories in anchovy samples, supported by peptides annotated with antioxidant and other enzyme-inhibitory activities. Processing and fortification were associated with only minor reductions in the relative proportions of some predicted activities, without eliminating the overall profile of annotated bioactive potential.
Previous studies have noted that food processing can influence peptide composition and activity. Certain treatments may enhance peptide release, while excessive heat or chemical fortification can mildly reduce peptide stability and function [56]. Overall, descriptive analysis based on sequence annotation shows that anchovies maintain the diversity of peptide categories predicted to have bioactive potential after processing and fortification. Although there was a decrease in the number and composition of certain peptides, the annotation patterns observed remained consistent across samples. This decrease is likely related to protein denaturation, structural changes, or peptide degradation during thermal and chemical processes, as reported in previous studies [43,44]. These findings emphasize the importance of controlling processing conditions in the context of preserving the peptide profile, although biological function stability was not experimentally evaluated in this study.
Overall, all reported bioactivities were inferred solely from sequence-based annotation and literature references. Given the exploratory and non-replicated nature of this study, these findings are presented descriptively and are not intended to indicate direct biological efficacy.
Peptide length distribution of bioactive peptides identified in raw, processed, and fortified anchovy samples
Bioactive peptides identified from each sample were analyzed according to peptide length to characterize the distribution of predicted bioactive peptide profiles. Peptide length is an important parameter because it can influence stability, absorption, and biological relevance - particularly in food, functional, and nutraceutical contexts.
The distribution of peptide lengths was determined based on matching results from the BIOPEP database and is presented separately for raw, processed, and fortified anchovy samples. Table 4 and Figure 2 summarize the length distribution of predicted bioactive peptides across these samples.
Table 4 Peptide length distribution of bioactives peptides identified in raw, processed, and fortified anchovy samples.
No |
Peptide length (aa) |
Samples |
|||||
Raw |
Processed |
Fortified |
|||||
Σ |
% |
Σ |
% |
Σ |
% |
||
1 |
2 aa |
309 |
48.97 |
187 |
75.10 |
181 |
73.88 |
2 |
3 aa |
278 |
44.06 |
52 |
20.88 |
52 |
21.22 |
3 |
4 aa |
23 |
3.65 |
8 |
3.21 |
10 |
4.08 |
4 |
≥ 5 aa |
21 |
3.33 |
2 |
0.80 |
2 |
0.82 |
|
Total |
631 |
100 |
249 |
100 |
245 |
100 |
Data source: LC-HRMS analysis (LPPT UGM, 2025).
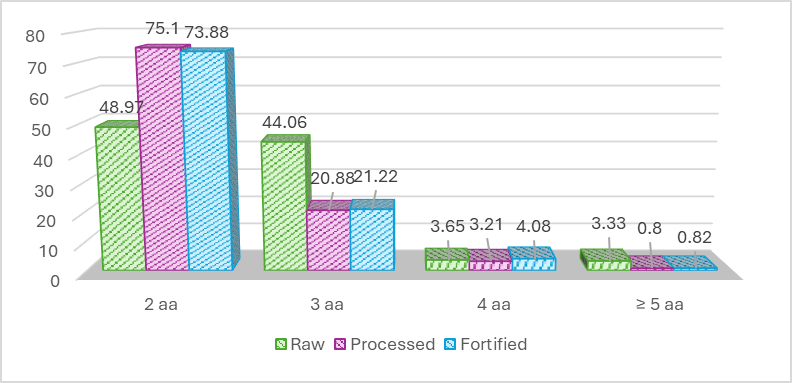
Figure 2 Peptide length distribution of bioactives peptides identified in raw, processed, and fortified anchovy samples.
Data source: LC-HRMS analysis (LPPT UGM, 2025).
In the raw anchovy samples, peptide length distribution was the most diverse, with dipeptides (2 amino acids) comprising 48.97%, tripeptides (3 amino acids) 44.06%, and peptides ≥ 4 amino acids reaching a cumulative 6.98%. This distribution reflects a broader range of peptide chain lengths and greater variation in the identified peptide profiles.
In contrast, the processed and fortified products displayed a narrower peptide length distribution dominated by dipeptides, accounting for 75.10% and 73.88% respectively, while peptides ≥ 4 amino acids constituted ≤ 4%. This pattern suggests that processing and fortification may be associated with more extensive protein fragmentation, leading to shorter peptide chains. According to previous reports, short peptides, despite their smaller size, can still retain or even exhibit enhanced biological functionality [32]. The bar chart visualization highlights differences in peptide length distributions among samples and supports a descriptive interpretation that processing and fortification are associated with shifts in peptide length profiles, rather than direct alterations of peptide structure.
The short-chain peptides identified in this study can be understood as the result of a combination of the intrinsic characteristics of the raw materials and the effects of the processing. Raw anchovies that have not undergone any treatment already contain a number of short-chain peptides, but there appears to be an increase in the proportion of short peptides in samples that have undergone processing. In line with previous reports, processing - particularly thermal treatments such as drying and frying - can cause peptide chain scissions, whereby longer protein chains are fragmented into shorter peptides. Heating and processing can trigger protein denaturation and expose previously hidden enzymatic cleavage sites, making it easier for proteolytic enzymes such as trypsin to break peptide bonds. As a result, proteins tend to degrade into shorter peptides, contributing to the dominance of short-chain peptides observed in the peptide profile of this study [57].
Peptide length plays a critical role in determining potential bioactivity. Short peptides are often reported to exhibit higher activity because they can interact more efficiently with biological targets such as enzymes or cell membranes. Their small size enhances penetration and reduces enzymatic degradation during digestion or metabolism [58]. Tripeptides and tetrapeptides with high bioactivity scores are also easier to synthesize and serve as promising candidates for nutraceutical and therapeutic development [59], due to their relatively simple structures and synthetic accessibility.
Peptides with lengths of 2 - 4 amino acids are commonly associated with antihypertensive and antioxidant functions, while antimicrobial peptides typically have longer sequences (≥ 5 amino acids) [60]. Other studies report that antioxidant peptides often range from 5 to 16 residues, with length influencing both stability and interaction with biological targets [61]. Previous research has shown that short-chain peptides such as Phenylalanine-Tryptophan (FW), Phenylalanine-Phenylalanine (FF), and Leucine-Proline-Phenylalanine (LPF) possess anti-inflammatory properties by inhibiting key enzymes involved in inflammation and prostaglandin synthesis, which are associated with pain and swelling [62]. These examples are cited here to contextualize the possible functional implications of peptide length; nevertheless, in this study, the bioactivities of the identified peptides were not experimentally validated and remain predicted based on sequence homology and literature evidence.
Distribution of bioactive peptides based on the number of biological activities in raw, processed, and fortified anchovy samples
After determining peptide activity types and peptide lengths, the next step involved grouping peptides according to the number of biological activity categories with which they were annotated. Table 5 presents a comparative distribution across the 3 sample types-raw, processed, and fortified anchovies.
Table 5 Distribution of bioactive peptides based on the number of biological activities in raw, processed, and fortified anchovy samples.
No |
Number of biological activities |
Samples |
|||||
Raw |
Processed |
Fortified |
|||||
Σ |
% |
Σ |
% |
Σ |
% |
||
1 |
1 activity |
406 |
64.65 |
141 |
56.63 |
135 |
55.10 |
2 |
2 activities |
115 |
18.31 |
57 |
22.89 |
54 |
22.04 |
3 |
3 activities |
51 |
8.12 |
19 |
7.63 |
22 |
8.98 |
4 |
4 activities |
31 |
4.94 |
15 |
6.02 |
15 |
6.12 |
5 |
≥ 5 activities |
25 |
3.98 |
17 |
6.83 |
19 |
7.76 |
|
Total |
628 |
100 |
249 |
100 |
245 |
100 |
Data source: LC-HRMS analysis (LPPT UGM, 2025).
The analysis revealed that the majority of peptides in all samples were annotated with only one biological activity: 64.65% in raw anchovies, 56.63% in processed anchovies, and 55.10% in fortified anchovies. However, the proportion of peptides annotated with 2 or more biological activity types increased in the processed and fortified samples. Peptides annotated with ≥ 5 biological activity categories increased from 3.98% in raw samples to 6.83% in processed and 7.76% in fortified samples. These descriptive patterns suggest that, although the total number of peptides decreased as a result of processing, the functional complexity of the remaining peptides actually increased.
In previous studies, peptides annotated with multiple biological activities are often considered superior because they can influence various physiological mechanisms simultaneously, potentially producing broader and more synergistic therapeutic effects [4], than peptides with single activities [63]. For instance, the peptides Alanine-Tryptophan (AW) and Tryptophan-Tyrosine (WY) have been reported as multifunctional bioactive peptides, acting as ACE inhibitors, antioxidants, and DPP-IV inhibitors, each with a PeptideRanker bioactivity score ≥ 0.9.
This distribution supports a descriptive interpretation that although processing and fortification reduce the overall number of peptides, they do not eliminate bioactivity potential. processed and fortification samples tends appear to contain yield peptides with more complex and diverse biological functions. In other words, while the total peptide quantity decreases, the diversity of biological activity per peptide may increase [64], thereby preserving - or even concentrating - the predicted bioactivity of anchovy‑derived peptides.
A reduction in peptide quantity does not necessarily imply a loss of function, as certain peptides with higher molecular stability may exhibit stronger or more specific biological effects when confirmed experimentally [65]. In the context of this exploratory, non‑replicated study, the observed patterns therefore suggest that the predicted bioactivity potential of anchovy peptides can remain largely preserved - and in some cases may appear to be concentrated in a smaller number of multifunctional sequences - following controlled processing and fortification. These observations highlight the promise of anchovy‑derived peptides as candidates for functional food and nutraceutical applications, while also underscoring the need for further in vitro and in vivo studies to validate their actual biological efficacy.
Distribution of peptides based on PeptideRanker bioactivity scores in raw, processed, and fortified anchovy samples
After identifying the quantity and types of bioactive peptides and their biological activities, the next step was to evaluate the intrinsic bioactivity potential of each peptide using predictive scores from PeptideRanker. This score reflects the likelihood that a peptide may exhibit biological activity, based on the sequence order and amino acid motif patterns statistically recognized by predictive machine-learning models.
PeptideRanker score distributions were analyzed comparatively among the 3 sample types - raw anchovies, processed anchovies, and fortified anchovies - to describe whether processing and fortification were associated with changes in the proportion of peptides with high bioactivity potential (score ≥ 0.8). The complete list of peptides, along with their individual scores, is provided in the supplementary file, while the following table summarizes the score distribution in each sample as the basis for functional interpretation and peptide candidate mapping.
Table 6 Distribution of peptides based on PeptideRanker bioactivity scores in raw, processed, and fortified anchovy samples.
No |
Score Range |
Samples Type |
|||||
Raw |
Processed |
Fortified |
|||||
Σ |
% |
Σ |
% |
Σ |
% |
||
1 |
High Bioactivity (≥ 0.8) |
3,854 |
13.67 |
382 |
11.59 |
373 |
14.58 |
2 |
Moderate Bioactivity (0.5 - 0.79) |
3,303 |
11.72 |
336 |
10.19 |
293 |
11.45 |
3 |
Low Bioactivity (< 0.5) |
21,036 |
74.61 |
2,579 |
78.22 |
1,892 |
73.96 |
|
Total |
28,193 |
100 |
3,297 |
100 |
2,558 |
100 |
Data source: LC-HRMS analysis (LPPT UGM, 2025).
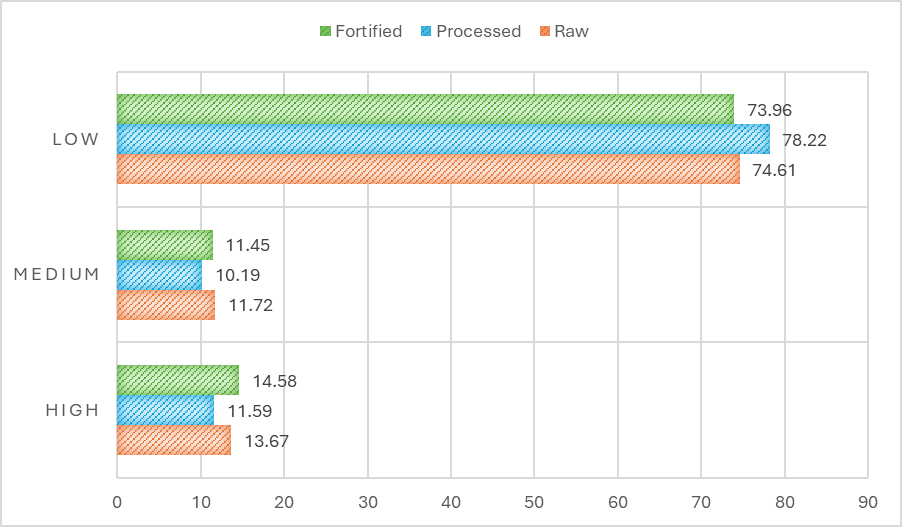
Figure 2 Distribution of peptides based on PeptideRanker bioactivity scores in raw, processed, and fortified anchovy samples.
Data source: LC-HRMS analysis (LPPT UGM, 2025).
In silico analysis using PeptideRanker revealed distinct predicted bioactivity score distributions among the 3 anchovy samples. The majority of peptides exhibited low bioactivity scores (< 0.5), accounting for 74.61% in raw anchovies, 78.22% in processed anchovies, and 73.96% in fortified anchovies - indicating that most peptides had low or suboptimal predicted intrinsic bioactivity. Nevertheless, peptides with high bioactivity scores (≥ 0.8) were still detected in meaningful proportions, particularly in fortified samples (14.58%), followed by raw (13.67%) and processed (11.59%) samples.
These descriptive findings suggest that fortification with cholecalciferol (vitamin D₃) does not eliminate, and may be associated with a higher proportion of peptides predicted to have high bioactivity scores in this dataset. This pattern could reflect structural modifications of proteins during processing and potential interactions with fortifying agents that promote the formation of bioactive peptide sequences. The observed trend - where fortified samples exhibited the highest proportion of high-score peptides - should therefore be regarded as hypothesis generating.
Peptides with high PeptideRanker scores are often considered, in the literature, to have greater biological significance, with potential roles such as angiotensin-converting enzyme inhibition (ACE-I), dipeptidyl peptidase IV inhibition (DPP-IV), antioxidant effects, and immunomodulatory functions [63,66]. Such peptides have been reported to be more promising candidates because they are more resistant to enzymatic digestion in the gastrointestinal tract and have a greater likelihood of reaching systemic circulation to exert biological effects [59].
Not all peptides produced during proteolysis exhibit the same biological potential; therefore, evaluating peptides based on their predicted bioactivity scores provides more meaningful insight for the selection and prioritization of candidates for health‑oriented and functional food applications than merely counting total peptide numbers without considering their biological relevance [66]. However, such scoring‑based evaluations should be regarded as a screening tool that guides hypothesis generation and candidate selection, rather than as a substitute for experimental assessment of actual bioactivity.
Conclusions
This exploratory study descriptively identified and mapped peptides and their predicted bioactivities from raw anchovies, processed anchovies, and fortified processed anchovies using LC-HRMS proteomics combined with BIOPEP-UWM and PeptideRanker bioinformatics analysis. The results showed that raw anchovies exhibited the highest proteomic complexity, while processing and fortification led to a reduction in the number of proteins and peptides, particularly those with a chain length of ≥ 4 amino acids, in this non‑replicated dataset. Nevertheless, processing did not entirely reduce bioactivity potential; in fact, peptides with high bioactivity scores (≥ 0.8) and multifunctional properties increased notably in fortified anchovy samples at a descriptive level. These patterns should be considered as hypotheses and not as evidence of actual increased bioactivity.
The predominant predicted bioactivities in anchovy‑derived peptides were Dipeptidyl Peptidase IV (DPP-IV) inhibition and Angiotensin-Converting Enzyme (ACE) inhibition, followed by antioxidant, neprilysin inhibitory, and Glutamate Carboxypeptidase II inhibitory (GCPII) activities, which are considered relevant in the literature to glucose and blood‑pressure regulation as well as oxidative stress and neurological pathways. The distribution of peptide lengths revealed a dominance of short peptides (2 - 3 amino acids) in processed and fortified anchovies, indicates greater protein fragmentation, likely as a result of heat treatment and fortification. Further research is needed.
Overall, the findings from these 3 sample types provide preliminary information suggesting that anchovy proteins may serve as promising precursors of peptides with putative multifunctional bioactivities and could be explored as candidates for anchovy‑based functional foods or nutraceutical products once their effects are confirmed through replicated experiments and targeted in vitro and in vivo validation.
Limitations of the study
This study was exploratory and descriptive and did not include biological or technical replication. Consequently, the peptide profiles reported here should be interpreted with caution, as they may represent treatment‑specific trends rather than reproducible biological effects. Without replicates, statistical comparisons cannot be performed, and causal interpretations - such as whether processing or cholecalciferol fortification truly influenced specific peptide patterns - cannot be confirmed. Nevertheless, these findings provide an initial foundation for future in vitro and in vivo studies to validate the predicted biological activities of the identified peptides.
The functional interpretation is based exclusively on in silico tools (BIOPEP‑UWM and PeptideRanker), which generate motif‑based predictions and ranking scores but do not account for gastrointestinal digestion, peptide degradation, absorption barriers, or real physiological conditions. Therefore, all reported functions should be regarded as hypothetical potentials rather than validated biological activities. The predominance of very short peptides (2 - 3 amino acids) may also reflect intrinsic material properties, processing‑induced fragmentation, or technical artefacts, so their predicted bioactivities must be interpreted with particular caution.
Future work should optimize processing and fortification conditions to enhance peptide stability and bioactivity, incorporate replicated LC-MS/MS analyses, and apply targeted validation strategies (e.g., PRM or SRM) together with functional assays to confirm the biological relevance and safety of prioritized peptides. In addition, the limited availability of annotated Engraulidae protein sequences in public databases may reduce identification completeness and increase the risk of misassignment; expanding proteomic resources for small pelagic fish will be essential to improve depth and accuracy in subsequent studies.
Acknowledgements
The author sincerely expresses gratitude to Universitas Gadjah Mada, Indonesia for funding support through the Doctoral Dissertation Research Grant (PDD) from Direktorat Penelitian dan Pengabdian kepada Masyarakat, Direktorat Jenderal Riset dan Pengembangan, Kementerian Pendidikan Tinggi, Sains, dan Teknologi in 2025, which made this study possible. Deep appreciation is also extended to the Laboratorium Penelitian dan Pengujian Terpadu (LPPT), Universitas Gadjah Mada, Yogyakarta, Indonesia for their technical assistance in the implementation and analysis of LC-HRMS proteomics, which was an essential component of this research. The author also conveys heartfelt thanks to the promoters, co-promoters, and all contributors who assisted in sample collection, bioinformatics data processing, and scientific discussions during the preparation of this manuscript.
Declaration of Generative AI in Scientific Writing
The author declares that artificial intelligence (AI) tools, including ChatGPT and Gemini AI, were used solely to refine the scientific language, enhance sentence structure, and check grammar to ensure the manuscript adheres to academic standards. All content, data interpretation, and scientific conclusions are entirely the author’s own work, without any AI-generated content. The author assumes full responsibility for the authenticity and scientific accuracy of this manuscript.
CRediT Author Statement
Artika Dewie: Conceptualization Ideas, design of methodology, Formal analysis, Investigation, Data Curation, Writing - Original Draft, Visualization; Susetyowati Susetyowati: Project administration Management and coordination, Validation, Resources, Data Curation, Writing - Review & Editing, Visualization, Supervision, Funding acquisition; Diah Rumekti Hadiati: Validation, Resources, Data Curation, Writing - Review & Editing, Visualization, Supervision.
References
[1] JYH Toy, LL Yu and D Huang. Functional foods for health: Evidence and insights. In: D Huang and LL Yu (Eds.). Evidence-based nutraceuticals and functional foods. Royal Society of Chemistry, London, 2025. p. 1-15.
[2] M Fekete, A Lehoczki, A Kryczyk-Poprawa, V Zábó, JT Varga, M Bálint, V Fazekas-Pongor, T Csípő, E Rząsa-Duran and P Varga. Functional foods in modern nutrition science: Mechanisms, evidence, and public health implications. Nutrients 2025; 17(13), 2153.
[3] A Vignesh, TC Amal, A Sarvalingam and K Vasanth. A review on the influence of nutraceuticals and functional foods on health. Food Chemistry Advances 2024; 5, 100749.
[4] TR Nuringtyas, L Hidayati, Z Rohmah, DK Paramita, A Suparmin, HH Prinanda, QN Febryzalita, SL Utami, LF Zulfa, BK Ardiansyah, SN Dewi and YA Purwestri. Bioactive peptides from sea cucumbers and sea urchins: Therapeutic roles and mechanistic insights. Trends in Sciences 2025; 22(5), 9513.
[5] RJS de Castro and HH Sato. Biologically active peptides: Processes for their generation, purification and identification and applications as natural additives in the food and pharmaceutical industries. Food Research International 2015; 74, 185-198.
[6] J Stadnik and P Kęska. Meat and fermented meat products as a source of bioactive peptides. Acta Scientiarum Polonorum, Technologia Alimentaria 2015; 14(3), 181-190.
[7] F Shahidi and A Saeid. Bioactivity of marine-derived peptides and proteins: A review. Marine Drugs 2025; 23(4), 157.
[8] V Manikkam, T Vasiljevic, ON Donkor and ML Mathai. A review of potential marine-derived hypotensive and anti-obesity peptides. Critical Reviews in Food Science and Nutrition 2016; 56(1), 92-112.
[9] L Xing, Z Wang, Y Hao and W Zhang. Marine products as a promising resource of bioactive peptides: Update of extraction strategies and their physiological regulatory effects. Journal of Agricultural and Food Chemistry 2022; 70(10), 3081-3095.
[10] E Daliri, D Oh and B Lee. Bioactive peptides. Foods 2017; 6(5), 32.
[11] I Perkons, MS Varunjikar and JD Rasinger. Unveiling the potential of proteomics in addressing food and feed safety challenges. EFSA Journal 2023; 21, e211013.
[12] SA Tadesse and SA Emire. Production and processing of antioxidant bioactive peptides: A driving force for the functional food market. Heliyon 2020; 6(8), e04765.
[13] C Aydoğan. Recent advances and applications in LC-HRMS for food and plant natural products: A critical review. Analytical and Bioanalytical Chemistry 2020; 412(9), 1973-1991.
[14] M Hayes, A Naik, L Mora, B Iñarra, J Ibarruri, C Bald, T Cariou, D Reid, M Gallagher, R Dragoy, J Galino, A Deya, S Albrektsen, L Thoresen and RG Solstad. Generation, characterisation and identification of bioactive peptides from mesopelagic fish protein hydrolysates using in silico and in vitro approaches. Marine Drugs 2024; 22(7), 297.
[15] NM Kari, SH Azmi, TC Tuan Zainazor, AI Zamri and F Ahmad. Anchovy’s protein as a potential precursor of Angiotensin-I Converting Enzyme (ACE) inhibitory peptide and Dipeptidyl Peptidase-IV (DPP-IV) inhibitory peptide by an in silico approach. Food Research 2023; 7(2), 248-261.
[16] DM Pampanin, E Larssen, F Provan, M Sivertsvik, P Ruoff and MO Sydnes. Detection of small bioactive peptides from Atlantic herring (Clupea harengus L.). Peptides 2012; 34(2), 423-426.
[17] MAS Imran, M Carrera, S Pérez-Polo, J Pérez, L Barros, S Dios and C Gestal. Insights into common octopus (Octopus vulgaris) ink proteome and bioactive peptides using proteomic approaches. Marine Drugs 2023; 21(4), 206.
[18] M Nemati, SR Shahosseini and P Ariaii. Review of fish protein hydrolysates: Production methods, antioxidant and antimicrobial activity and nanoencapsulation. Food Science and Biotechnology 2024; 33(8), 1789-1803.
[19] J Du, M Xiao, N Sudo and Q Liu. Bioactive peptides of marine organisms: Roles in the reduction and control of cardiovascular diseases. Food Science & Nutrition 2024; 12(8), 5271-5284.
[20] ES Okeke, RE Ita, EJ Egong, LE Udofia, CL Mgbechidinma and OD Akan. Metaproteomics insights into fermented fish and vegetable products and associated microbes. Food Chemistry: Molecular Sciences 2021; 3, 100045.
[21] C Litaay, Indriawati, L Nuraini, CEW Anggara, Sriharti, A Triyono and T Hidayat. The value of the nutritional composition of fresh fish and flour-anchovy. IOP Conference Series: Earth and Environmental Science 2023; 1273, 012074
[22] RW Kusumaningtyas, T Sudiarti and N Laily. Development and nutritional value of instant anchovy (Stolephorus indicus) soup as vitamin D-rich meal for pregnant women. Journal of Food Technology Research 2025; 12(2), 102--113.
[23] PA Aneesh, J Varkey, R Anandan, S Mathew, KK Asha, PT Lakshmanan and VT Shankar. Proximate composition, amino acid and fatty acid profiles of commerson’s anchovy (Stolephorus commersonii) and Indian mackerel (Rastrelliger Kanagurta). Biochemical and Cellular Archives 2012; 12, 371-376.
[24] L Najafian L and AS Babji. Fractionation and identification of novel antioxidant peptides from fermented fish (pekasam). Journal of Food Measurement and Characterization 2018; 12(3), 2174-2183.
[25] MA Kurnianto, F Syahbanu, H Hamidatun, Yushinta and A Sanjaya. Prediction and mapping of potential bioactive peptide from traditional fermented anchovy (Rusip) using bioinformatics approaches. Squalen Bulletin of Marine and Fisheries Postharvest and Biotechnolog 2023; 18(2), 93-105.
[26] D Jelić, M Vraneš, S Zeljković, S Veličković, F Veljković, A Cvetkovski, B Pejova and S Papovic. Advancing food fortification with kinetics: Stabilizing vitamin D3 through calcium carbonate-based vehicles. Food Chemistry 2025; 487, 144811.
[27] C Litaay, TA Mutiara, A Indriati, F Novianti, L Nuraini and N Rahman. Fortifikasi tepung ikan teri (Stolephorus sp.) terhadap karakteristik fisik dan mikrostruktur mi berbasis sagu (in Indonesian). Jurnal Pengolahan Hasil Perikanan Indonesia 2023; 26(1), 127-138.
[28] MEL Diogenes, FF Bezerra, EP Rezende, MF Taveira, I Pinhal and CM Donangelo. Effect of calcium plus vitamin D supplementation during pregnancy in Brazilian adolescent mothers: A randomized, placebo-controlled trial. The American Journal of Clinical Nutrition 2013; 98(1), 82-91.
[29] M Mogan and E Trisnawati. Pengaruh suplemen kalsium dan vitamin D Pada ibu hamil dengan hipertensi di puskesmas sentani (in Indonesian). Gema Kesehatan 2022; 14, 188-195.
[30] V Kumar, RJ Shakila, AU Muzaddadi, G Jeyasekaran, D Sukumar and P Padmavathy. In vitro stability of ACE-Inhibitory peptides of rohu fish waste to heat, pH and gastrointestinal enzymes. Turkish Journal of Fisheries and Aquatic Sciences 2024; 24(3), TRJFAS23788.
[31] Y Gong, M Feng and J Sun. Effect of different thermal processing methods and thermal core temperatures on the protein structure and in vitro digestive characteristics of beef. Food Chemistry 2025; 464, 141751.
[32] GJ Fadimu, TT Le, H Gill, A Farahnaky, OO Olatunde and T Truong. Enhancing the biological activities of food protein-derived peptides using non-thermal technologies: A review. Foods 2022; 11(13), 1823.
[33] JM Jacobs, HM Mottaz, LR Yu, DJ Anderson, RJ Moore, WNU Chen, KJ Auberry, EF Strittmatter, ME Monroe, BD Thrall, DG Camp and RD Smith. Multidimensional proteome analysis of human mammary epithelial cells. Journal of Proteome Research 2004; 3(1), 68-75.
[34] MR Hoopmann, GE Merrihew, PD von Haller and MJ MacCoss. Post analysis data acquisition for the iterative MS/MS sampling of proteomics mixtures. Journal of Proteome Research 2009; 8(4), 1870-1875.
[35] B Ryu, KH Shin and SK Kim. Muscle protein hydrolysates and amino acid composition in fish. Marine Drugs 2021; 19(7), 377.
[36] M Senevirathne and S Kim. Production and health effects of peptides from fish proteins. In: SK Kim (Ed.). Marine proteins and peptides: Biological activities and applications. Wiley, New Jersey, 2013, p. 737-751.
[37] JT Ryan, RP Ross, D Bolton, GF Fitzgerald and C Stanton. Bioactive peptides from muscle sources: Meat and fish. Nutrients 2011; 3(9), 765-791.
[38] M Senevirathne and SK Kim. Development of bioactive peptides from fish proteins and their health promoting ability. Advances in Food and Nutrition Research 2012; 65, 235-248.
[39] Y Zhao and YH Lin. Whole-cell protein identification using the concept of unique peptides. Genomics Proteomics Bioinformatics 2010; 8(1), 33-41.
[40] DL Swaney, CD Wenger and JJ Coon. Value of using multiple proteases for large-scale mass spectrometry-based proteomics. Journal of Proteome Research 2010; 9(3), 1323-1329.
[41] Y Zhao and YH Lin. Whole-cell protein identification using the concept of unique peptides. Genomics Proteomics Bioinformatics 2010; 8(1), 33-41.
[42] M Chevallet, H Diemer, S Luche, A van Dorsselaer, T Rabilloud and E Leize-Wagner. Improved mass spectrometry compatibility is afforded by ammoniacal silver staining. Proteomics 2006; 6(8), 2350-2354.
[43] R Li, N Raak and L Roman. Recent advances in processing-induced changes in the structure, techno-functional properties and nutritional quality of animal- and plant-based food proteins. Foods 2025; 14(5), 764.
[44] W Ogugua. Role of processing conditions in enhancing peptide stability and bioactivity in foods. Journal of Food Processing & Technology 2023; 14, 1001068.
[45] S Mäkinen, J Hiidenhovi, X Huang, ADS Lima, L Azevedo, J Setälä, AL Valimaa, P Mattila and D Granato. Production of bioactive peptides from baltic herring (Clupea harengus Membras): Dipeptidyl peptidase-4 inhibitory, antioxidant and antiproliferative properties. Molecules 2022; 27(18), 5816.
[46] SJ Chen and WT Jiaang. Current advances and therapeutic potential of agents targeting dipeptidyl peptidases-IV, -II, 8/9 and fibroblast activation protein. Current Topics in Medicinal Chemistry 2011; 11, 1447-1463.
[47] L Fleury, B Deracinois, C Dugardin, AB Nongonierma, RJ FitzGerald, C Flahaut, B Cudennec and R Ravallec. In vivo and in vitro comparison of the DPP-IV inhibitory potential of food proteins from different origins after gastrointestinal digestion. International Journal of Molecular Sciences 2022; 23(15), 8365.
[48] DY Pujiastuti, MNG Amin, MA Alamsjah and JL Hsu. Marine organisms as potential sources of bioactive peptides that inhibit the activity of angiotensin i-converting enzyme: A review. Molecules 2019; 24(14), 2541.
[49] CD Cinq-Mars and ECY Li-Chan. Optimizing angiotensin I-converting enzyme inhibitory activity of pacific hake (Merluccius productus) fillet hydrolysate using response surface methodology and ultrafiltration. Journal of Agricultural and Food Chemistry 2007; 55(23), 9380-9388.
[50] W Cao, C Zhang, H Ji and J Hao. Optimization of peptic hydrolysis parameters for the production of angiotensin I‐converting enzyme inhibitory hydrolysate from Acetes chinensis through Plackett-Burman and response surface methodological approaches. Journal of the Science of Food and Agriculture 2012; 92(1), 42-48.
[51] X Zhang, C Hu, E Tian, Y Shen, W Liu and J Li. Comprehensive review on neprilysin (NEP) inhibitors: Design, structure-activity relationships, and clinical applications. Frontiers in Pharmacology 2024; 15, 1501407.
[52] N Esser and S Zraika. Neprilysin inhibition: A new therapeutic option for type 2 diabetes? Diabetologia 2019; 62(7), 1113-1122.
[53] DV Ferraris, K Shukla and T Tsukamoto. Structure-activity relationships of glutamate carboxypeptidase II (GCPII) inhibitors. Current Medicinal Chemistry 2012; 19(9), 1282-1294.
[54] KT Jegani, A Balde and RA Nazeer. A review on anti-inflammatory and antioxidant peptides derived from marine organisms: Mechanism of action and therapeutic applications. Food Bioscience 2025; 63, 105745.
[55] VK Khavinson. Peptides, genome, aging. Advances in Gerontology 2014; 4(4), 337-345.
[56] DJ Daroit and A Brandelli. In vivo bioactivities of food protein-derived peptides - a current review. Current Opinion in Food Science 2021; 39, 120-129.
[57] ZF Bhat, JD Morton, AEA Bekhit, S Kumar and HF Bhat. Thermal processing implications on the digestibility of meat, fish and seafood proteins. Comprehensive Reviews in Food Science and Food Safety 2021; 20(5), 4511-4548.
[58] S Avilés-Gaxiola, I García-Aguiar, LA Jiménez-Ortega, EP Gutiérrez-Grijalva and JB Heredia. Bioactive plant peptides: Physicochemical features, structure-function insights and mechanism of action. Molecules 2025; 30(18), 3683.
[59] S Windarto, VE Herawati, YJ Wijaya and DA Indriati. Exploring the bioactive potential of peptides derived from the RuBisCO protein in Caulerpa racemosa: An in silico approach. Biotech Studies 2025; 34(2), 71-85.
[60] B Tamam, D Syah, HN Lioe, MT Suhartono and WA Kusuma. Beberapa penciri berbasis sekuens untuk mengenali sifat fungsional peptida bioaktif: Studi eksplorasi (in Indonesian). Jurnal Sains dan Teknologi Pangan 2018; 29(1), 1-9.
[61] K Purohit, N Reddy and A Sunna. Exploring the potential of bioactive peptides: From natural sources to therapeutics. International Journal of Molecular Sciences 2024; 25(3), 1391.
[62] Suwarjoyowirayatno, C Hidayat, TD Wahyuningsih and R Indrati. Identification of bioactive peptide inhibitors of cyclooxygenase-2 from peanut worm (Siphonosoma australe): An in silico studies. Trends in Sciences 2025; 22(5), 9565.
[63] L Amigo, D Martínez-Maqueda and B Hernández-Ledesma. In silico and in vitro analysis of multifunctionality of animal food-derived peptides. Foods 2020; 9(8), 991.
[64] M Akbarian, A Khani, S Eghbalpour and VN Uversky. Bioactive peptides: Synthesis, sources, applications, and proposed mechanisms of action. International Journal of Molecular Sciences 2022; 23(3), 1445.
[65] BP Singh, PF Santos, S Luna-Suárez and CJ Martínez. Editorial: Advances in bioactive peptides and functional properties of foods. Frontiers in Chemistry 2025; 12, 1538944.
[66] P Kęska, KM Wójciak and J Stadnik. Bioactive peptides from beef products fermented by acid whey - in vitro and in silico study. Scientia Agricola 2019; 76(4), 311-320.