
Trends
Sci.
2026;
23(7):
12559
Dual Inhibition and Safety Profiling of Padina Australis Extract as A Marine-Derived α-Glucosidase Modulator: An Integrated In Vitro and In Silico Study
Ni Made Puspawati1,*, Putu Faraditha Maharani1, I Made Oka Adi Parwata1,
Setyani Budiari2 and Rajesh K Patel3
1Department of Chemistry, Faculty of Mathematics and Natural Sciences, Udayana University, Bali 80361, Indonesia
2Research Center for Chemistry, National Research and Innovation Agency, Building 452, Banten 15314, Indonesia
3Department of Biosciences, Veer Narmad South Gujarat University, Gujarat 395007, India
(*Corresponding author’s e-mail: [email protected])
Received: 6 November 2025, Revised: 14 December 2025, Accepted: 21 December 2025, Published: 25 February 2026
Abstract
Padina australis is a brown seaweed that shows promise as a natural source of α-glucosidase inhibitors for diabetes management. This study evaluated the α-glucosidase inhibitory activity of P. australis extracts and examined how extraction solvents influence this activity. It also identified potential active metabolites using gas chromatography–mass spectrometry (GC–MS), molecular docking, and toxicity prediction, thereby providing new evidence on the α-glucosidase inhibitory potential of P. australis. Ultrasonic-assisted extraction with n-hexane, ethyl acetate, and ethanol revealed that the ethyl acetate extract exhibited the highest inhibitory activity at 500 µg∙mL−1 (94.55 ± 0.16%), comparable to the ethanol extract (93.75 ± 2.56%) and higher than the n-hexane extract (45.37 ± 4.59%). GC–MS analysis identified 23 compounds, including loliolide, 2(4H)-benzofuranone, 5,6,7,7a-tetrahydro-4,4,7a-trimethyl, phytol, neophytadiene, and several fatty acids. Docking against yeast α-glucosidase (3A4A), human maltase–glucoamylase (3L4T), and lysosomal α-glucosidase (5NN6) indicated that loliolide and benzofuranone derivatives showed affinities approaching acarbose on the yeast enzyme, while neophytadiene and polyunsaturated fatty acids exhibited the most favorable interactions with the human intestinal enzyme. Binding to lysosomal α-glucosidase (5NN6) was consistently weaker, suggesting a degree of selectivity away from lysosomal targets. ProTox-3.0 predicted low acute toxicity for most metabolites (Classes V-VI) and moderate toxicity for two compounds (Class IV), whereas loliolide was classified as higher-risk (Class II). These findings support Padina australis as a potential source of α-glucosidase-modulating metabolites at a screening level. However, comprehensive studies including dose-response assays, enzyme kinetic characterization, fractionation, and toxicity testing are required to validate and extend these preliminary observations.
Keywords: Padina australis, α-glucosidase, In vitro, In silico, Molecular docking, Fatty acids, Antidiabetic
Introduction
Indonesia, the world’s largest archipelagic nation, encompasses vast marine ecosystems covering nearly two-thirds of its territory. This richness provides diverse biological resources, including marine macroalgae, which are increasingly recognized as valuable sources of bioactive compounds for pharmaceutical, nutraceutical, and biomedical applications. However, despite being a leading seaweed producer, Indonesia’s marine resources remain underutilized, with most being
exported as raw materials rather than explored for their therapeutic potential [1-3].
Marine macroalgae, particularly brown algae (Phaeophyceae), are known for their diverse secondary metabolites, such as terpenoids, phenolics, steroids, and fatty acids that exhibit pharmacological activities, including antioxidant, anti-inflammatory, anti-obesity, and antidiabetic effects [4-6]. Among these activities, α-glucosidase inhibition has gained attention as a mechanism to delay carbohydrate digestion and reduce postprandial blood glucose spikes, which is beneficial in diabetes management [7,8].
The brown algal genus Padina (family Dictyotaceae), commonly known as fan-shaped seaweed, contains bioactive constituents with multiple therapeutic properties, including hypolipidemic, hypoglycemic, antibacterial, and antioxidant activities [9]. A previous study by Naveen et al. [10] on Padina tetrastromatica revealed significant α-glucosidase inhibitory activity, mainly attributed to its fatty acid components such as palmitic and linoleic acids [10]. Despite its abundance in Indonesian coastal regions, the phytochemical profile and antidiabetic mechanism of Padina australis have not been extensively investigated.
Therefore, this study aimed to evaluate the α‑glucosidase inhibitory potential of P. australis extracts using complementary in vitro and in silico approaches. This study provides new evidence on the α-glucosidase inhibitory activity of P. australis extracts and evaluates how extraction solvents influence the inhibitory activity. The initial screening of crude extracts was performed at a single concentration to identify the most active solvent fraction as a starting point for mechanistic exploration. The volatile and semi‑volatile bioactive compounds in the most active extract were then identified by gas chromatography–mass spectrometry (GC-MS), while molecular docking and toxicity predictions were conducted to assess their binding affinity and putative safety profiles against α‑glucosidase enzymes from S. cerevisiae (PDB: 3A4A) and Homo sapiens (PDB: 3L4T and 5NN6). Although the absence of dose–response curves and kinetic parameters represents an important limitation, the present work provides preliminary evidence supporting the role of fatty acid‑derived and terpenoid compounds from P. australis as candidate α‑glucosidase modulators for antidiabetic applications and defines a rational basis for more detailed future investigations.
Materials and methods
Materials
The study used brown seaweed (Padina australis) collected from Banten waters, West Java, Indonesia, aquadest, technical grade ethanol (C2H5OH), and n-hexane (C6H14) from FULLTIME. Sodium dihydrogen phosphate (NaH2PO4), disodium hydrogen phosphate (Na2HPO4), methanol (CH3OH), and ethyl acetate (C4H8O2) were from Merck. The p-nitrophenyl-α-D-glucopyranoside (pNPG) substrate and α-glucosidase enzyme were from Sigma-Aldrich.
Phosphate buffer solution preparation
Based on the method by G. Gomori [11], 0.1 M phosphate buffer solution at pH 7.0 was prepared according to the Henderson–Hasselbalch equation, using sodium dihydrogen phosphate monohydrate (NaH₂PO₄·H₂O) and disodium hydrogen phosphate heptahydrate (Na₂HPO₄·7H₂O) as buffering components. Based on the desired pH and the second dissociation constant of phosphoric acid (pKa₂ = 7.21), the molar ratio between (HPO₄²⁻) and (H₂PO₄⁻) was calculated to be 0.617. For a total phosphate concentration of 0.1 M, the resulting concentrations of (HPO₄²⁻) and (H₂PO₄⁻) were 0.062 M and 0.038 M, respectively. The corresponding reagent masses for 100 mL buffer were 0.524 g of NaH₂PO₄·H₂O (Mᵣ = 137.99 g·mol⁻¹) and 1.662 g of Na₂HPO₄·7H₂O (Mᵣ = 268.06 g·mol⁻¹). Both salts were accurately weighed using an analytical balance and dissolved in approximately 70 mL of deionized water under gentle stirring until completely dissolved. The solution pH was measured using a calibrated pH meter at 25 °C and, if necessary, adjusted to exactly pH 7.00 using 1.0 M NaOH or HCl. The solution was then transferred quantitatively to a 100 mL volumetric flask and diluted to the mark with deionized water. After homogenization, the final pH was verified. The buffer was stored at 4 °C and used within four weeks.
Sample preparation
Adapted and modified from the method by Widwiastuti et al. [12], brown seaweed (Padina australis) was collected from Banten waters, West Java, Indonesia. Morphological identification was based on thallus shape, blade width, and the presence of characteristic hairline zones, which were compared with standard taxonomic references. Samples were washed thoroughly with aquadest to remove debris and epiphytes, oven-dried at 50 °C until constant weight, and ground into fine powder using a grinder. The powder was sieved through a 32-mesh filter to ensure uniform particle size. The dried powder was stored in airtight containers at room temperature until further analysis.
Moisture content determination
Moisture content was determined according to AOAC 930.04. Approximately 3 g of the powdered sample was heated at 105 °C for 4 h until a constant weight was achieved. Moisture content was calculated as shown below:
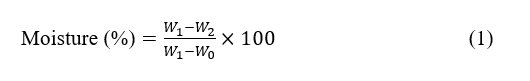
The W0 means the weight of the empty crucible, W1 for the weight of the crucible with the sample before heating, and W2 for the weight of the crucible with the sample weight after heating.
Extraction procedure
Powdered Padina australis (20 g per extraction) was subjected to sonication-assisted maceration. For each solvent, 20 g of powder was separately mixed with 200 mL of ethanol, ethyl acetate, or n-hexane, sonicated for 15 min at 45 °C, and then macerated for 24 h with occasional stirring. The mixtures were filtered, and the filtrates were evaporated to dryness. The extraction yield for each solvent was calculated as follows:
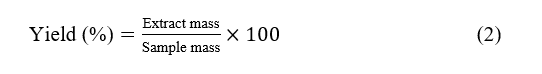
α-Glucosidase inhibitory assay
Modified from a study by Efriani et al. [13], the α‑glucosidase inhibitory activity was measured using p‑nitrophenyl‑α‑D‑glucopyranoside (pNPG) as substrate. The reaction mixture contained 25 µL of α‑glucosidase from Saccharomyces cerevisiae with enzyme activity of ≥ 10 U∙mg−1 of protein; working solution 0.04 U∙mL−1 in 0.1 M phosphate buffer (pH 7.0), 5 µL of extract (10 mg/mL in methanol; final test concentration 500 µg∙mL−1), and 25 µL of pNPG (0.5 mM) in 0.1 M phosphate buffer. The total reaction volume in each well was 200 µL, adjusted with buffer. After pre‑incubation of the enzyme and sample at 37 °C for 10 min, the reaction was initiated by the addition of substrate and incubated at 37 °C for 20 min. The reaction was stopped by adding 100 µL of 5 %(w/v) Na2CO3. Absorbance was measured at 405 nm using a microplate reader, and p‑nitrophenol formation was monitored using an extinction coefficient of 18,300 M−1cm−1 at 405 nm under the assay conditions. Inhibition percentage was calculated as:
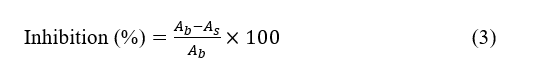
The Ab and As are the absorbance of blank and sample, respectively. Due to limited sample availability, each condition was measured as three technical replicates within a single experimental run. The results are therefore presented as mean ± standard deviation of these technical replicates and interpreted as preliminary screening data rather than definitive potency estimates.
GC-MS analysis
Volatile compounds of the ethyl acetate extract were analyzed using gas chromatography-mass spectrometry (Agilent, USA). The GC-MS used splitless injection mode, with 19091 S-433 type column (HP-5MS UI 5% Phenyl Methyl Silox with 30 m column length, 250 µm internal diameter and 0.25 µm film thickness) with Electron Impact (EI) ionization mode with 70 eV ionization energy. One milligram of extract was dissolved in methanol, filtered through microfilters, and injected (1 μL) into the GC–MS system. Helium was used as the carrier gas with a flow rate of 1 mL∙min−1. The oven temperature was programmed from 40 °C for 1 min to 300 °C at 10 °C∙min−1, then 325 °C at 10 °C∙min−1 for 4 min. The injector and detector (interface) temperatures were set at 250 °C and 230 - 250 °C, respectively. Compounds were identified by comparing retention times and fragmentation patterns with NIST spectral libraries. Chromatogram peak areas were used to estimate relative abundance, and compounds with a similarity index ≥ 80% were selected for further analysis.
Molecular docking
Molecular
docking simulations were carried out using AutoDock Vina 1.2.7.
Crystal structures of α‑glucosidase from S.
cerevisiae
(3A4A), human intestinal maltase–glucoamylase (3L4T), and human
lysosomal acid α‑glucosidase (5NN6) were retrieved from the
RCSB Protein Data Bank (https://www.rcsb.org/). All water molecules
and non‑essential ligands were removed using PyMOL. Polar
hydrogens and Kollman charges were added, and proteins were kept
rigid during docking. Ligand structures (23 compounds identified by
GC–MS, acarbose, and miglitol) were downloaded from PubChem,
energy‑minimized, and converted to pdbqt format using Open
Babel 2.4.1. For each receptor,
the grid box was centered on
the catalytic site to
encompass the active‑site cavity,
with dimensions of
20×20×20 Å3.
The grid centers were set as follows: 3A4A (center_x = 21.549,
center_y = 7.654, center_z = 23.540 Å), 5NN6 (center_x = –14.230,
center_y = 32.011, center_z = 95.780 Å), and 3L4T (center_x =
44.491, center_y = 90.509, center_z = 34.391 Å). Docking was
performed with an exhaustiveness value of 16, generating eight
binding poses for each ligand-receptor pair. The Vina scoring
function (an AutoDock4‑based empirical free‑energy
model) was used to estimate binding free energy (ΔG, kcal∙mol–1).
The reliability of the protocol was verified by redocking the native
ligands (e.g. acarbose, miglitol, and glucose), which yielded RMSD
values of 0.730, 1.499 and 2.553 Å, respectively, indicating
acceptable reproduction of the crystallographic binding modes. The
best‑scoring pose for each ligand was subjected to interaction
analysis using LigPlot+ and BIOVIA Discovery Studio, focusing on
hydrogen bonds, hydrophobic contacts, and interactions with
catalytic residues.
In silico toxicity prediction
The toxicity of identified compounds was evaluated using the ProTox-3.0 webserver (https://tox.charite.de/protox3/). Predictions included acute toxicity (LD₅₀) and organ-specific effects. Compounds were classified into toxicity classes according to the Globally Harmonized System (GHS), where Class I indicates highly toxic (LD₅₀ ≤ 5 mg∙kg−1) and Class VI indicates non-toxic compounds (LD₅₀ > 5,000 mg∙kg−1).
Results and discussion
Identification and moisture content of padina australis
Morphological observation confirmed that the brown seaweed sample collected from Banten waters in March 2025 corresponded to Padina australis Hauck. The specimen exhibited a fan-shaped thallus with regular concentric hair-line scars and distally arranged sori, features consistent with diagnostic descriptions reported by Win et al. [14]. These characteristics clearly distinguish P. australis from other Padina species. The moisture content of the dried simplicia was 7.67 %(w/w), remaining below the maximum limit of 10% specified by the Indonesian Herbal Pharmacopeia. This indicates that the sample met the required quality standard for further extraction and ensured the stability of its bioactive constituents.
Extraction
A
combined ultrasonic–maceration approach was applied to 20 g of
Padina
australis
powder to optimize the recovery of bioactive metabolites.
Ultrasonic-assisted extraction at 45 °C for 15 min promoted
mechanical disruption of cellular structures through cavitation,
thereby enhancing solvent penetration and minimizing thermal
degradation of sensitive compounds, consistent with the optimal
conditions reported by Fu et
al.
[15]. The subsequent 24 h maceration utilized ethanol, ethyl
acetate, and n-hexane to target metabolites across a broad polarity
range. Ethanol, a polar protic solvent, effectively extracts
hydrogen-bonding compounds; ethyl acetate, a
semi-polar
aprotic solvent, selectively solubilizes medium-polarity phenolics
and terpenoids; while
n-hexane, a non-polar hydrocarbon,
primarily extracts lipophilic constituents such as fatty acids
[16,17].
The resulting yields were 2.75% (ethanol), 2.70%
(n-hexane), and 2.27% (ethyl acetate), which were lower than
the 4.62 - 6.01% yields reported by Sari et
al.
[18], likely reflecting differences in solvent strength,
species-specific metabolite composition, and extraction conditions.
Overall, these findings underscore the critical influence of solvent
polarity and extraction parameters on the efficiency and
phytochemical profile of Padina
australis
extracts.
α-glucosidase inhibitory activity
The results showed that the ethyl acetate extract exhibited the highest α-glucosidase inhibitory activity of 94.55 ± 0.16% at a concentration of 500 µg∙mL−1 compared to the other solvents. The ethanol and n-hexane extracts had inhibitory activities of 93.75 ± 2.56% and 45.37 ± 4.59% respectively, as shown in Table 1 The n-hexane extract showed markedly lower inhibition, likely because this non-polar solvent primarily extracts lipophilic constituents and recovers fewer of the semi-polar phenolic and terpenoid inhibitors that interact favorably with α-glucosidase [19-21]. One-way ANOVA confirmed significant differences between solvents (F = 258.586, p-value < 0.001), with Tukey’s post-hoc test revealing that both ethyl acetate and ethanol extracts were significantly more active than n-hexane (p-value < 0.001), while not significantly different from each other (p-value = 1.000). Given its slightly higher inhibition and notably lower variability (SD = 0.16%), the ethyl acetate extract, rich in semi-polar terpenoid, flavonoid, and phenolic compounds known for α-glucosidase inhibition, was selected for further GC–MS profiling and in silico docking studies.
Table 1 α-Glucosidase inhibitory activity of Padina australis extracts.
Extract |
Inhibition ± SD (%) |
Padina ethanol |
93.75 ± 2.56a |
Padina ethyl acetate |
94.55 ± 0.16a |
Padina n-hexane |
45.37 ± 4.59b |
Data presented as mean ± SD of three technical replicates. Different superscript letters indicate statistically significant differences (p-value < 0.001 by one-way ANOVA with Tukey post-hoc test). Although α‑glucosidase inhibition was evaluated only at a single concentration (500 µg/mL), the clear difference between the ethyl acetate and n‑hexane extracts, together with the low variability of the ethyl acetate data (SD = 0.16%, n = 3 technical replicates), supports its selection as the most active fraction for subsequent chemical and in silico analyses. We acknowledge that the absence of full concentration–response curves and IC50 values, including 95% confidence intervals and parallel testing of a standard inhibitor such as acarbose, is a significant limitation that precludes rigorous potency ranking and direct quantitative comparison with clinically used drugs. However, the amount of P. australis material available for the present study was insufficient to perform repeated experiments at multiple concentrations. The current data should therefore be considered as a preliminary activity screen.
The assay, as shown in Figure 1, was conducted using p-nitrophenyl-α-D-glucopyranoside (p-NPG) as a synthetic substrate, which is hydrolyzed by α-glucosidase to release p-nitrophenol (yellow chromophore). The intensity of the yellow color, measured spectrophotometrically at 405 nm, is inversely proportional to the inhibitory effect of the extract. Thus, a reduction in absorbance reflects the ability of the metabolites to bind the enzyme’s active site and prevent substrate hydrolysis.
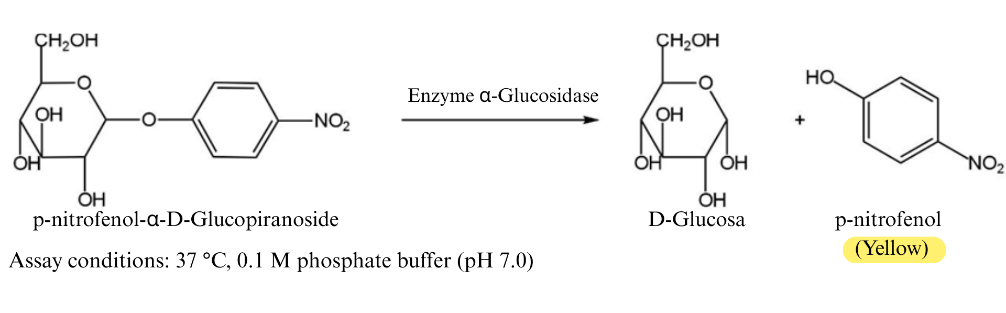
Figure 1 Schematic representation of the α-glucosidase inhibition assay using p-nitrophenyl-α-D-glucopyranoside (p-NPG) as a substrate. In the absence of inhibitors, α-glucosidase hydrolyzes p-NPG to release p-nitrophenol, producing a yellow color measurable at 405 nm.
The inhibition suggests that semi-polar bioactive compounds such as phenolics, terpenoids, and fatty acids play a major role in the extract’s activity [22,23]. The ability of the crude extract to achieve over 90% inhibition implies the presence of dominant active constituents or synergistic effects among multiple metabolites. These findings are consistent with a study by Tuan et al. [24] reporting significant α-glucosidase inhibitory activity in brown algae extracts, highlighting P. australis as a promising source of natural antidiabetic agents.
GC–MS analysis of ethyl acetate extract
Given the α-glucosidase inhibition exhibited by the ethyl acetate extract of Padina australis, a GC–MS analysis was conducted to identify the bioactive metabolites potentially responsible for this activity, as shown in Figure 2. The chromatographic profile from Figure 2 revealed 23 compounds representing diverse chemical classes, including saturated and unsaturated fatty acids, terpenoids, aldehydes, and heterocyclic aromatics. The most abundant constituent was n-hexadecanoic acid (palmitic acid, 49.11%), followed by 9-octadecenoic acid (oleic acid, 19.18%) and tetradecanoic acid (myristic acid, 16.41%). These long-chain fatty acids, commonly found in brown algae, have been extensively reported to exhibit antioxidant, anti-inflammatory, and glucose metabolism–modulating effects [25].
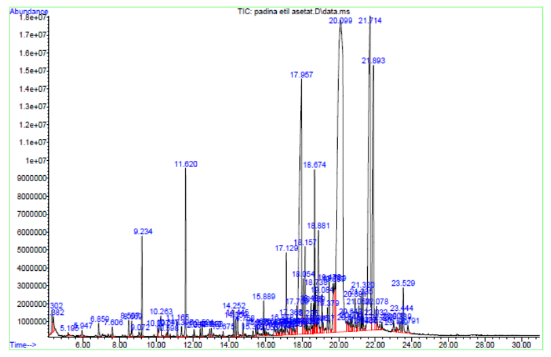
Figure 2 GC-MS chromatogram of the ethyl acetate extract of Padina australis.
Several minor compounds, such as loliolide, 2(4H)-benzofuranone,5,6,7,7a-tetrahydro-4,4,7a-trimethyl, neophytadiene, and unsaturated aldehydes (e.g., 2-decenal, 2,4-decadienal), were also detected. Despite their low abundance, these metabolites may contribute synergistically to the overall inhibitory activity through complementary biological mechanisms. The dominance of fatty acids and terpenoid derivatives observed here is consistent with previous studies on Padina species, where similar profiles were correlated with potent α-glucosidase inhibition [26,27]. Collectively, the GC–MS findings highlight the chemical diversity of P. australis and provide a molecular basis for its antidiabetic potential. These results serve as a foundation for subsequent in silico analyses aimed at elucidating the interaction mechanisms between the identified compounds and α-glucosidase active sites.
Molecular docking
To harmonize the in vitro and in silico components of this work, we selected three α‑glucosidase‑related structures that reflect both the assay conditions and physiologically relevant human targets. The yeast α‑glucosidase isomaltase (PDB ID: 3A4A) corresponds to the S. cerevisiae enzyme used in the inhibition assay and is widely employed for screening natural product inhibitors. For the human system, we used the N‑terminal domain of human maltase–glucoamylase (PDB ID: 3L4T), a well‑established model of intestinal α‑glucosidase involved in luminal starch digestion and a clinically relevant target for postprandial glucose control. In addition, the human lysosomal acid α‑glucosidase (PDB ID: 5NN6) was included as a comparative receptor because it represents another physiologically important α‑glucosidase isoform implicated in intracellular glycogen catabolism. Together, these three structures allow evaluation of ligand binding across yeast and human enzymes with distinct biological roles, while maintaining a direct link to the in vitro assay.
To further elucidate the mechanism underlying the α-glucosidase inhibition observed in vitro, molecular docking simulations were performed using enzyme models from Homo sapiens (PDB ID: 5NN6, 3L4T) and Saccharomyces cerevisiae (PDB ID: 3A4A), as shown in Figure 3. The use of three receptors aimed to evaluate ligand selectivity across biological systems, with S. cerevisiae representing the in vitro assay enzyme, while the human enzymes provided therapeutic relevance. Receptor structures were prepared by removing water molecules and native ligands to optimize the binding cavity and ensure docking accuracy [28]. Validation by redocking the native ligands, glucose, miglitol, and acarbose produced RMSD values of 0.730, 2.553, and 1.499 Å, respectively, confirming the reliability of the docking protocol.
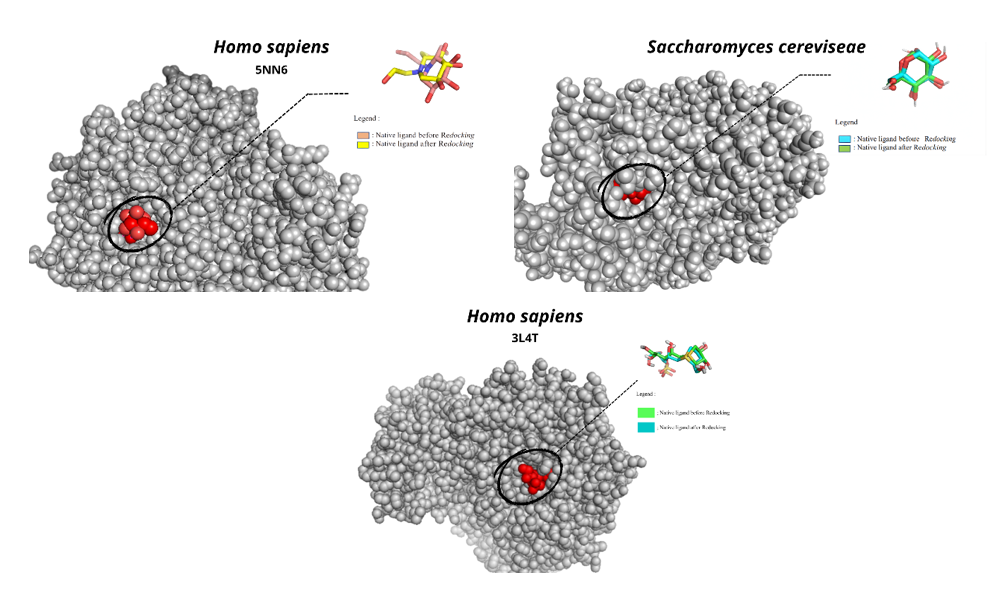
Figure 3 Validation of the docking method by redocking of native ligands in α-glucosidase from Homo sapiens (5NN6, 3L4T) and S. cerevisiae (3A4A). The low RMSD values (2.553, 1.499 and 0.730 Å) confirm the accuracy and reliability of the docking protocol.
Docking analysis of compounds identified in the ethyl acetate extract of Padina australis revealed clear variations in binding affinity across the three α-glucosidase enzymes. To better relate the chemical composition of the extract with its inhibitory activity, all identified compounds were docked against 3 representative α-glucosidase models, yeast isomaltase (3A4A), human intestinal maltase-glucoamylase (3L4T), and human lysosomal α-glucosidase (5NN6). The resulting Vina scores for each compound–enzyme pair are summarized in Table 2, providing a comparative overview of their predicted inhibitory potential.
Table 2 Comparison of molecular docking results (ΔG, kcal/mol) across three α-glucosidase receptors.
No |
Compound |
Binding affinity (Kcal/mol) |
||
3A4A |
5NN6 |
3L4T |
||
1 |
2-Hexenal, (E)- |
–3.885 |
–3.983 |
–3.903 |
2 |
2-Heptenal, (Z)- |
–4.366 |
–4.492 |
–4.149 |
3 |
Octanal |
–4.184 |
–4.183 |
–4.351 |
4 |
Nonanal |
–4.299 |
–4.27 |
–3.905 |
5 |
2-Nonenal, (E)- |
–4.376 |
–4.581 |
–4.339 |
6 |
Octadecanoic acid |
–5.127 |
–4.741 |
–4.425 |
7 |
2-Decenal, (Z) |
–4.793 |
–4.468 |
–4.577 |
8 |
2,4-Decadienal, (E,Z) |
–4.953 |
–4.929 |
–4.889 |
9 |
2,4-Decadienal |
–4.931 |
–4.713 |
–4.591 |
10 |
n-Decanoic acid |
–5.005 |
–4.586 |
–4.739 |
11 |
Tetradecanoic acid |
–4.881 |
–4.855 |
–4.732 |
12 |
Loliolide |
–6.351 |
–4.887 |
–4.523 |
13 |
Pentadecanoic acid |
–5.578 |
–4.668 |
–4.648 |
14 |
Neophytadiene |
–6.019 |
–5.424 |
–5.655 |
15 |
2-Pentadecanone, 6,10,14-trimethyl |
–5.659 |
–5.075 |
–5.104 |
16 |
Palmitoleic acid |
–5.765 |
–4.617 |
–4.67 |
17 |
n-Hexadecanoic acid |
–5.249 |
–5.078 |
–4.551 |
18 |
Phytol |
–6.092 |
–5.101 |
–5.134 |
19 |
9-Octadecenoic acid, (E) |
–5.570 |
–4.877 |
–4.738 |
20 |
10E,12Z-Octadecadienoic acid |
–6.084 |
–4.953 |
–4.965 |
21 |
9,12-Octadecadienoic acid (Z,Z) |
–6.043 |
–5.22 |
–5.315 |
22 |
1H-Pyrrole-2,5-dione, 3-ethyl-4-methyl |
–4.892 |
–5.67 |
–3.983 |
23 |
2(4H)-Benzofuranone, 5,6,7,7a-tetrahydro-4,4,7a-trimethyl |
–6.212 |
–5.547 |
–4.912 |
Positive Control |
||||
24 |
Acarbose |
–7.029 |
–5.901 |
–6.226 |
ΔG values (kcal/mol) represent binding free energies estimated by AutoDock Vina. Lower (more negative) values indicate stronger predicted binding affinity. Data are from single docking runs; RMSD validation values for native ligand redocking were 0.7302553 Å, confirming protocol reliability.
Several compounds showed relatively strong binding (ΔG ≤ –6.0 kcal∙mol−1) to the yeast α-glucosidase (3A4A). The top-scoring ligand was loliolide (–6.351 kcal∙mol−1), followed by 2(4H)-benzofuranone, 5,6,7,7a-tetrahydro-4,4,7a-trimethyl (–6.212 kcal∙mol−1), phytol (–6.092 kcal∙mol−1), 10E,12Z-octadecadienoic acid (–6.084 kcal∙mol−1). 9,12-octadecadienoic acid (–6.043 kcal∙mol−1) and neophytadiene (–6.019 kcal∙mol−1) were just slightly below this threshold. These values approach the docking score of the known inhibitor acarbose on 3A4A (–7.029 kcal∙mol−1), indicating that in silico, some Padina metabolites can bind nearly as well as acarbose to the yeast enzyme.
For
the human intestinal enzyme (3L4T), binding affinities were
generally slightly weaker than for yeast, which is expected, given
3L4T’s active site is more adapted to human substrates and may be
less accommodating to some algal metabolites. The best 3L4T binders
were neophytadiene (–5.655 kcal∙mol−1),
9,12-octadecadienoic acid (–5.315 kcal/mol), phytol
(–5.134
kcal∙mol−1),
2-pentadecanone,6,10,14-trimethyl (–5.104 kcal∙mol−1),
and 10E,12Z-octadecadienoic acid (–4.965 kcal∙mol−1).
Interestingly, loliolide scored
–4.523 kcal∙mol−1
on
3L4T, which is significantly lower in affinity than its yeast enzyme
score, suggesting loliolide is much better suited to the yeast
binding pocket than to the human intestinal site.
Meanwhile,
the fatty acids and hydrophobic terpenes
(neophytadiene,
phytol) maintained moderate affinity
in 3L4T’s site.
Acarbose’s score on 3L4T was –6.226 kcal∙mol−1,
again stronger than any single compound, but some ligands like
neophytadiene were within
~0.6 kcal∙mol−1.
For
the human lysosomal enzyme (5NN6), all compounds bound weaker, on
average, than for the
other 2 receptors. The highest affinity
observed on
5NN6 was for 1H-pyrrole-2,5-dione,
3-ethyl-4-methyl (–5.670 kcal∙mol−1)
followed by 2(4H)-benzofuranone,
5,6,7,7a-tetrahydro-4,4,7a-trimethyl (–5.547 kcal∙mol−1),
neophytadiene (–5.424 kcal∙mol−1),
9,12-octadecadienoic acid (–5.220 kcal∙mol−1),
and phytol (–5.101 kcal∙mol−1).
Loliolide only scored –4.887 kcal∙mol−1
on 5NN6. Notably, acarbose also had its weakest score on 5NN6
(–5.901 kcal∙mol−1)
compared to yeast and 3L4T. This overall weaker binding to 5NN6 is
likely due to structural differences: The lysosomal enzyme has a
more confined active site tailored to its function in glycogen
hydrolysis, so it does not accommodate these bulkier phytochemicals
as easily.
In addition to the global binding energies summarized in Table 2, the specific amino acid residues involved in stabilizing the top five ligands in each receptor were examined in Tables 3 - 5 and visualised in Figures 4 - 6. For 3A4A, loliolide, phytol, neophytadiene, and polyunsaturated fatty acids such as 10E,12Z‑octadecadienoic acid formed hydrogen bonds and hydrophobic contacts with key catalytic and neighboring residues (e.g. Asp214, Glu276, His348, and Arg442), similar to those observed for acarbose. In 3L4T, neophytadiene, linoleic acid, and related fatty acids interacted with residues within the intestinal α‑glucosidase active site, including Asp327, Asp443, His600, and Arg526, whereas loliolide displayed fewer and weaker contacts, consistent with its lower affinity on this receptor. For 5NN6, the number and strength of interactions were generally reduced, with ligands forming mainly hydrophobic contacts in a narrower catalytic pocket, in line with the lower binding energies recorded for this lysosomal isoform. These interaction patterns, together with the energy scores, suggest that P. australis metabolites preferentially target yeast and intestinal α‑glucosidases over the lysosomal enzyme, which is desirable from a therapeutic selectivity perspective.
Table 3 Key amino acid residue interactions for the top five ligands in Saccharomyces cerevisiae α-Glucosidase (PDB code: 3A4A) compared to the reference inhibitor acarbose.
No |
Compounds |
Binding affinity (Kcal∙mol−1) |
Amino acid residues |
|
Hydrogen bonds |
Hydrophobic bonds |
|||
A. |
Top Five Compounds Identified in the Pre-Screening Analysis |
|||
1. |
Loliolide |
–6.351 |
Glu411 |
Glu277, Gln353, Phe303, Asp352, Gln279, Tyr158, Arg442, Arg315 |
2. |
2(4H)-Benzofuranone, 5,6,7,7a-tetrahydro-4,4,7a-trimethyl |
–6.212 |
- |
|
3. |
Phytol |
–6.092 |
Tyr158, Asn415, Glu411, |
Tyr316, Arg442, Phe159, Asp352, Phe178, Glu277, Gln279, Phe303, Gln353, Arg315 |
4. |
10E,12Z-Octadecadienoic acid |
–6.084 |
Arg442, His351, Asp215 |
Tyr72, Phe78, Asp352, Glu277, Asp307, Phe303, His280, Arg315, Tyr158, Glu411 |
5. |
9,12-Octadecadienoic acid (Z,Z)- |
–6.043 |
His351, Arg213, Glu277 |
Tyr158, Val216, Asp215, Tyr72, Asp352, Gln353, Phe303, Glu411, Arg442, Arg315, Gln279 |
B. |
Positive Control |
|||
6. |
Acarbose |
–7.029 |
Arg213, Asp215, Gln279, His280, Lys156, Tyr158, His112, Gln182, Asp69, Arg442 |
Glu277, Val216, Tyr72, Phe159, Phe178, Glu411, Ser157, Asp352, Phe303, Arg315 |
Table 4 Key amino acid residue interactions for the top five ligands against the N-terminal human maltase glucoamylase receptor (PDB code: 3L4T) compared with the reference inhibitor acarbose.
No |
Compounds |
Binding affinity (kcal∙mol−1) |
Amino acid residues |
|
Hydrogen bonds |
Hydrophobic bonds |
|||
A. |
Top Five Compounds Identified in the Pre-Screening Analysis |
|||
1. |
Neophytadiene |
–5.655 |
- |
Lys480, Phe450, Trp441, Ile328, Tyr299, His600, Phe575, Trp406 |
2. |
9,12-Octadecadienoic acid (Z,Z)- |
–5.315 |
- |
Phe450, Trp406, Tyr299 |
3. |
Phytol |
–5.134 |
Arg202 |
Lys480, Asp203, Met444, Phe450, Trp406, Phe575, Tyr299 |
4. |
2-Pentadecanone, 6,10,14-trimethyl |
–5.104 |
- |
Trp406, Phe575, Tyr299, Tyr605 |
5. |
10E,12Z-Octadecadienoic acid |
–4.965 |
Gln603 |
Trp406, Phe575, Tyr299, His600, Trp441 |
B. |
Positive Control |
|||
6. |
Acarbose |
–6.226 |
Tyr299, Gln603, Asp542, Asp203, Arg202 |
- |
Table 5 Key amino acid residue interactions for the top five ligands against the human lysosomal acid α-glucosidase receptor (PDB code: 5NN6) compared with the reference inhibitor acarbose.
No |
Compounds |
Binding affinity (kcal∙mol−1) |
Amino acid residues |
|
Hydrogen bonds |
Hydrophobic bonds |
|||
A. |
Top five compounds identified in the pre-screening analysis |
|||
1. |
1H-Pyrrole-2,5-dione, 3-ethyl-4-methyl- |
–5.670 |
Arg600, Asp616, His674 |
Asp518, Trp613, Phe649, Trp376, Asp404, Leu405, Trp481 |
2. |
Neophytadiene |
–5.424 |
- |
Trp481, Leu405, Asp616, Asp518, Phe649, Asp404, His674, Trp376, Leu678, Leu677 |
3. |
2(4H)-Benzofuranone, 5,6,7,7a-tetrahydro-4,4,7a-trimethyl |
–5.547 |
Arg600 |
Met519, Trp376, Phe525, Phe649, Trp481 |
4. |
9,12-Octadecadienoic acid (Z,Z)- |
–5.22 |
Asp616, His674 |
Trp516, Trp612, Asp515, Asp404, Trp481, Asp282, Leu650, Ser676, Trp376, Phe649 |
5. |
Phytol |
–5.101 |
His674, Asp404 |
Trp516, Trp376, Asp518, Trp481, Leu405, Asp616, Phe525, Asp282, Phe649 |
B. |
Positive Control |
|||
6. |
Acarbose |
–5.901 |
Asp616, Asp282, Asp518, Asp404, Asp443 |
Phe525, Met519, Phe649, Trp516, Trp376, Ile441, Leu405, Trp481 |
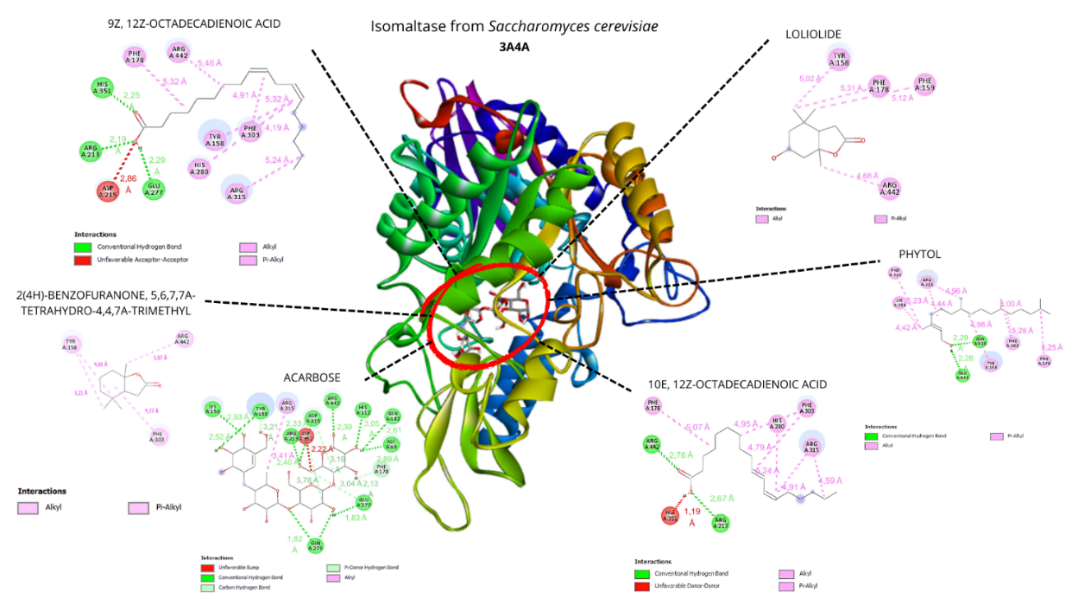
Figure 4 Visualization of the top five ligands and acarbose interacting with amino acid residues of Saccharomyces cerevisiae α-glucosidase (PDB code: 3A4A).
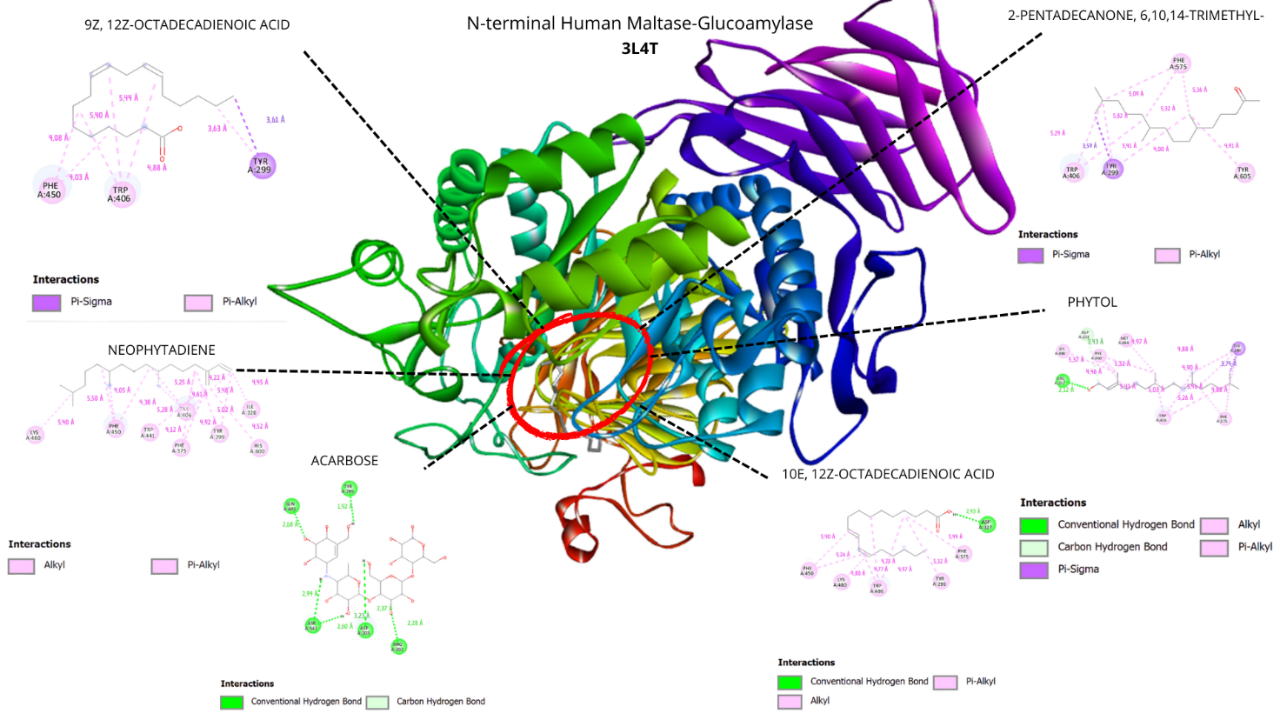
Figure 5 Visualization of the top five ligands and acarbose interacting with amino acid residues of N-Terminal Human Maltase Glucoamylase (PDB code: 3L4T).
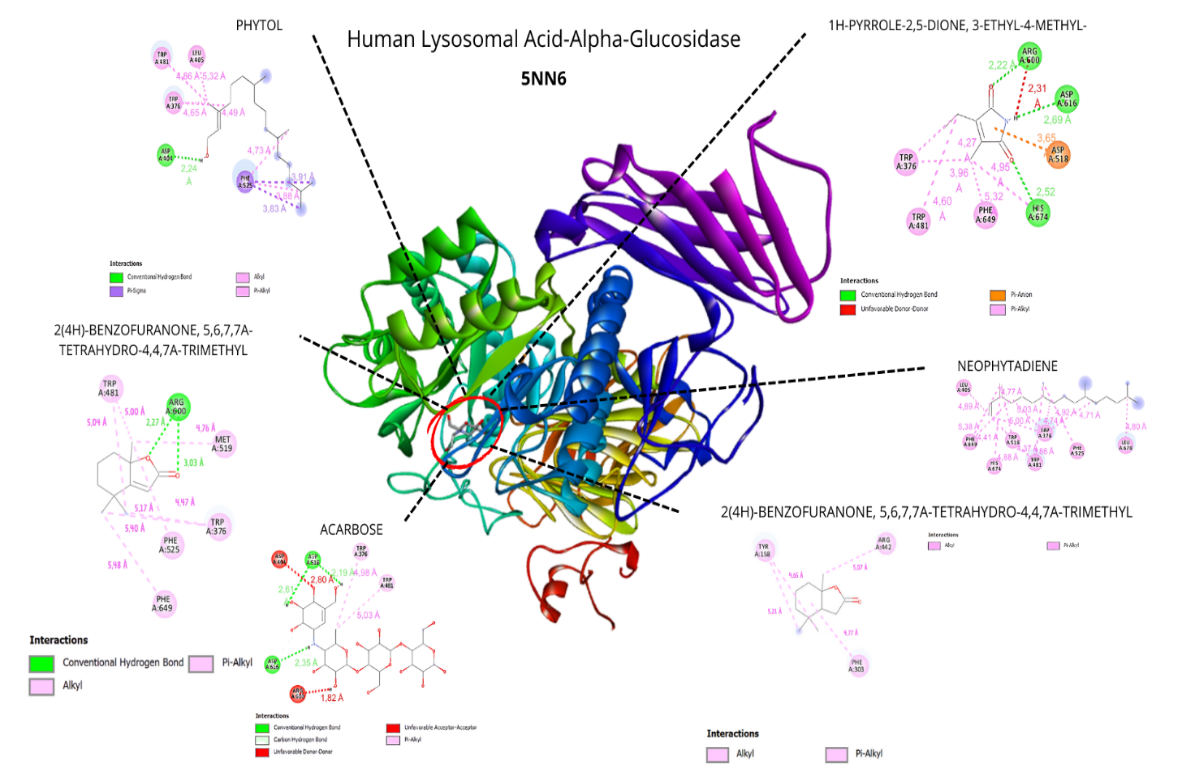
Figure 6 Visualization of the top five ligands and acarbose interacting with amino acid residues of human lysosomal acid α-glucosidase (PDB code: 5NN6).
The docking analysis across the three α-glucosidase structures, yeast isomaltase (3A4A), human intestinal maltase-glucoamylase (3L4T), and human lysosomal acid α-glucosidase (5NN6), demonstrated a coherent and biologically meaningful gradient of binding affinities that reflects the functional and structural divergence of these enzymes. Compounds from Padina australis consistently exhibited the highest interactions with 3A4A, a receptor widely used in natural-product inhibitor screening due to its well-characterized catalytic site and relevance to α-glucosidase assays by Abudurexeti et al. [29]. This affinity pattern was preserved, though moderately reduced, in 3L4T, which is frequently applied as a human intestinal α-glucosidase model in docking studies and provides a structurally reliable representation of the mammalian catalytic domain [30]. The inclusion of 5NN6 further strengthened the analysis, as this human lysosomal α-glucosidase structure has been validated in recent computational studies investigating potential antidiabetic agents, ensuring that the docking results capture ligand interactions across multiple physiologically relevant α-glucosidase conformations [31]. Together, these findings support the translational relevance of the in vitro inhibition data and highlight the conserved features of ligand recognition across yeast and human α-glucosidase systems.
In contrast, all compounds displayed noticeably weaker binding to 5NN6, the lysosomal isoform, whose catalytic architecture differs substantially because of its role in intracellular glycogen degradation rather than luminal carbohydrate processing. This marked reduction in affinity suggests a favorable degree of selectivity, as ligands preferentially interact with the intestinal α-glucosidase rather than the lysosomal enzyme, reducing the likelihood of off-target effects associated with lysosomal dysfunction. The internal consistency of these trends, supported by the positive control acarbose, which followed the same affinity hierarchy (3A4A > 3L4T > 5NN6), reinforces the validity of the multi-receptor docking approach and highlights the extract’s potential efficacy in modulating the clinically relevant human α-glucosidase while maintaining a promising safety profile with respect to unintended lysosomal interactions.
The stability of the enzyme–ligand complexes was largely driven by hydrogen bonds (2.7 - 3.2 Å) and hydrophobic interactions (3.5 - 4.0 Å), both of which contribute to decreasing Gibbs free energy (ΔG) and improving overall binding stability [22]. Fatty acids and terpenoid derivatives demonstrated good interaction potential due to their amphiphilic nature, which allows them to engage simultaneously in polar and hydrophobic contacts with key catalytic residues, including Ser, His, and Asp. These interactions occur within or near the catalytic pocket, suggesting that these ligands may directly interact with residues in or near the catalytic pocket, consistent with reported direct binding of unsaturated fatty acids to α-glucosidase. Such binding can obstruct substrate access and reduce enzymatic turnover, consistent with the mechanism reported for unsaturated fatty acids like oleic and linoleic acids that inhibited α-glucosidase through reversible, non-catalytic complex formation [23].
Toxicity prediction and safety evaluation
Toxicity assessment of the key compounds from the ethyl acetate extract of Padina australis was performed using ProTox-3.0, an AI-based platform that predicts acute oral toxicity (LD₅₀), toxicity class, and potential adverse effects from chemical structures [32]. ProTox-3.0 assessment (Table 6) showed that most compounds fell into low-toxicity classes (GHS Classes V–VI), with two compounds in Class IV and loliolide in Class II, with LD₅₀ values ranging from 34 to 10,000 mg/kg. Compounds such as 2-Pentadecanone, 6,10,14-trimethyl, 10E,12Z-octadecadienoic acid, and 9,12 - octadecadienoic acid (Z, Z) were categorized as “may be harmful” to “non-toxic,” consistent with their natural occurrence in marine organisms and known low acute toxicity based on the previous study by Anacleto-santos et al. [33]. Similarly, phytol and neophytadiene were predicted to be safe, supporting previous reports of their antioxidant and anti-inflammatory properties [34,35].
Table 6 Predicted acute oral toxicity of major compounds identified from the ethyl acetate extract of Padina australis based on ProTox-3.0 analysis.
Compound |
LD₅₀ (mg∙kg−1) |
Toxicity class |
Predicted toxicity |
1,800 |
4 |
Harmful if swallowed |
|
Loliolide |
34 |
2 |
Fatal if swallowed |
Neophytadiene |
5,050 |
6 |
Non-toxic |
Phytol |
5,000 |
5 |
May be harmful if swallowed |
10E,12Z-Octadecadienoic acid |
3,200 |
5 |
May be harmful if swallowed |
9,12-Octadecadienoic acid (Z,Z)- |
10,000 |
6 |
Non-toxic |
2(4H)-Benzofuranone, 5,6,7,7a-tetrahydro-4,4,7a-trimethyl |
1,190 |
4 |
|
2-Pentadecanone, 6,10,14-trimethyl |
5,000 |
5 |
May be harmful if swallowed |
Neophytadiene and 9,12-octadecadienoic acid (Z,Z) were classified as non-toxic (Class VI), while phytol, 10E,12Z-octadecadienoic acid, and 2-pentadecanone, 6,10,14-trimethyl showed low toxicity (Class V). Two compounds, 1H-pyrrole-2,5-dione, 3-ethyl-4-methyl and 2(4H)-benzofuranone, 5,6,7,7a-tetrahydro-4,4,7a-trimethyl, were classified as Class IV, indicating moderate acute toxicity at high doses.
Loliolide showed the highest predicted toxicity (Class II; LD₅₀ = 34 mg/kg). These results suggest that, except for loliolide, the major constituents of the extract fall within acceptable toxicity ranges for early-stage drug discovery. Further assessment using in vitro cytotoxicity, hepatotoxicity assays, and in vivo tolerability studies is necessary to confirm safety beyond the in-silico predictions.
Conclusions
This study provides preliminary indicator that the ethyl acetate extract of Padina australis exhibits α-glucosidase inhibitory activity (94.55 ± 0.16% at 500 µg∙mL−1) and contains volatile and semi-volatile metabolites, mainly long-chain fatty acids and terpenoid-like derivatives. Molecular docking analyses suggest that several metabolites, particularly loliolide, neophytadiene, phytol, and polyunsaturated fatty acids, show favourable binding towards yeast and human intestinal α-glucosidase, with weaker interactions observed for the lysosomal isoform, indicating a potential degree of selectivity. In silico toxicity predictions indicated generally low acute toxicity for most compounds (Classes V–VI), with two compounds showing moderate toxicity (Class IV) and loliolide warranting further safety evaluation (Class II). Overall, these findings support Padina australis as a potential source of α-glucosidase-modulating metabolites at a screening level. However, comprehensive studies including dose-response assays, enzyme kinetic characterization, fractionation, and experimental toxicity testing are required to validate and extend these preliminary observations.
Acknowledgements
The authors would like to thank Universitas Udayana, Bali, Indonesia and National Research and Innovation Agency, South Tangerang, Indonesia, for providing the research facilities for this study.
Declaration of Generative AI in Scientific Writing
The authors acknowledge the use of Quillbot Premium during the preparation of this manuscript to improve clarity and readability. All text generated or modified using this tool was subsequently reviewed, edited, and verified by the authors, who bear full responsibility for the content’s accuracy and integrity.
CRediT Author Statement
Ni Made Puspawati: Conceptualization; Supervision; Methodology; Validation; Project administration; Resources; Writing - Reviewing and Editing. Putu Faraditha Maharani: Methodology; Visualization; Data Curation; Software; Investigation; Writing - Original draft preparation. I Made Oka Adi Parwata: Supervision; Methodology; Validation. Setyani Budiari: Investigation; Supervision; Validation. Rajesh K Patel: Methodology; Resources.
References
[1] L Zhang, W Liao, Y Huang, Y Wen, Y Chu and C Zhao. Global seaweed farming and processing in the past 20 years. Processing and Nutrition 2022; 4, 23.
[2] MA Rimmer, S Larson, I Lapong, AH Purnomo, PR Pong-masak, L Swanepoel and NA Paul. Seaweed aquaculture in Indonesia contributes to social and economic aspects of livelihoods and community wellbeing. Sustainability 2021; 13(19), 10946.
[3] M Basyuni, M Puspita, R Rahmania, H Albasri, SS Al, F Menne, Y Ihrami, SG Salmo, A Susilowati, SH Larekeng and E Ardli. Heliyon current biodiversity status , distribution , and prospects of seaweed in Indonesia: A systematic review. Heliyon 2024; 10(10), e31073.
[4] S Lomartire and AMM Gonçalves. An overview of potential seaweed-derived bioactive compounds for pharmaceutical applications. Marine Drugs 2022; 20(2), 141.
[5] Y Bouafir, M Mounir, A Nebbak, L Belfarhi, B Aouzal, F Boufahja, H Bendif and M Bruno. Fitoterapia algal bioactive compounds : A review on their characteristics and medicinal properties. Fitoterapia 2025; 183, 106591.
[6] THP Brotosudarmo, D Pringgenies and AA Wibawa. Current progress in exploring structural changes in brown algae fucoxanthin and its potential bioactivity for human health. Trends in Sciences 2024; 21(9), 7984.
[7] N Moheimanian, H Mirkhani, A Purkhosrow, J Sohrabipour and A Reza. In Vitro and In Vivo antidiabetic , α -glucosidase inhibition and antibacterial activities of three brown algae , polycladia myrica , padina antillarum , and sargassum boveanum , and a red alga , palisada perforata from the persian gulf. The IJ Pharmaceutical Research 2023; 22(1), e133731.
[8] A Maheswari and D Sdep. In vitro correlation studies of antidiabetic, antioxidant activity and HPLC-ESI-MS/MS analysis of marine seaweeds from Gulf of Mannar. Regional Studies in Marine Science 2022; 56, 102682.
[9] YL Cheryl-Low, KL Theam and HV Lee. Alginate-derived solid acid catalyst for esterification of low-cost palm fatty acid distillate. Energy Conversion and Management 2015; 106, 932-940.
[10] J Naveen, R Baskaran and V Baskaran. Profiling of bioactives and in vitro evaluation of antioxidant and antidiabetic property of polyphenols of marine algae Padina tetrastromatica. Algal Research 2021; 55, 102250.
[11] G Gomori. Preparation of buffers for use in enzyme studies. Methods in Enzymology 1955; 1, 138-146.
[12] H Widwiastuti, RY Asworo, YS Tjahjaningsih, NC Wulandari and A Dewi. Pengaruh ukuran simplisia dan lama kontak pada ekstraksi senyawa aktif simplisia kayu jawa (Lannea Coromandelica) menggunakan metode maserasi. Jurnal Kimia Mulawarman 2022; 19(2), 86-90.
[13] L Efriani, I Hadi, A Irawan, M Ulfah and TA Putra. uji aktivitas inhibisi enzim a-glukosidase terhadap ekstrak aseton, etanol, dan methanol daun mangrove (rhizopora mucronata) sebagai antidiabetes. Medical Sains Jurnal Ilmiah Kefarmasian 2023; 8(4), 1371-1377.
[14] NNW Win, T Hanyuda, S Arai, M Uchimura, A Prathep, SGA Draisma, SM Phang, IA Abbott, AJK Millar and H Kawai. A taxonomic study of the genus padina ( dictyotales , phaeophyceae ) including the descriptions of four new species from japan , hawaii , and the andaman sea. Journal of Phycology 2011; 47, 1193-1209.
[15] CWF Fu, CW Ho, WTL Yong, F Abas, TB Tan and CP Tan. Extraction of phenolic antioxidants from four selected seaweeds obtained from Sabah. International Food Research Journal 2016; 23(6), 2363-2369.
[16] C Reichardt and T Welton. Solvents and solvent effects in organic chemistry. 4th ed. Wiley, New Jersey, 2010.
[17] H Nawaz, MA Shad, N Rehman, H Andaleeb and N Ullah. Effect of solvent polarity on extraction yield and antioxidant properties of phytochemicals from bean (Phaseolus vulgaris) seeds. Brazilian Journal of Pharmaceutical Sciences 2015; 56, e17129.
[18] A Permatasari, I Batubara, M Nursid and K Kelautan. Pengaruh konsentrasi etanol dan waktu maserasi terhadap rendemen , kadar total fenol dan aktivitas antioksidan ekstrak rumput laut padina australis. Majalah Ilmiah Biologi Biosfera: A Scientific Journal 2020; 37(2), 78-84.
[19] M Ponnanikajamideed, M Malini, C Malarkodi and S Rajeshkumar. Bioactivity and phytochemical constituents of marine brown seaweed (padina tetrastromatica) extract from various organic solvents. International Journal of Pharmacy & Therapeutics 2014; 5(2), 108-112.
[20] M Gazali, O Jolanda, A Husni, F Adibah and A Majid. In vitro α-Amylase and α-Glucosidase Inhibitory Activity of Green Seaweed Halimeda tuna Extract from the Coast of Lhok Bubon, Aceh. Plants 2023; 12(2), 393.
[21] SN Hikmah, R Setianto, BA Dewi and R Utami. Antibacterial activities of N-hexan ethyl acetate fraction and water from ethanol extract of yodium leaves (Jatrophora multifida L.) Against pseudomonas aeruginosa ATCC 27853. STRADA Jurnal Ilmiah Kesehatan 2021; 10(1), 1242-1249.
[22] S Chen, B Lin, J Gu, T Yong, X Gao, Y Xie, C Xiao, JY Zhan and Q Wu. Binding interaction of betulinic acid to α -glucosidase and its alleviation on postprandial hyperglycemia. Molecules 2022; 27(8), 1-12.
[23] Y Isono, H Watanabe, M Kumada and T Takara. Black tea decreases postprandial blood glucose levels in healthy humans and contains high-molecular-weight polyphenols that i nhibit α -g lucosidase and α -amylase in vitro: A randomized, double blind, placebo-controlled, crossover trial. Functional Foods in Health and Disease 2021; 11(5), 222-237.
[24] ND Tuan, NC Quoc, DN Ly, BT Tuyen, LD Quang, HH Phien, TQ De and TT Men. bioactive extracts from padina boryana thivy from Phu Quoc Island, Vietnam: In vitro antioxidant, anticancer, alpha-glucosidase inhibitory, anti-inflammatory, antimicrobial, and hepatoprotective activities. Tropical Journal of Natural Product Research 2024; 8(12), 9555-9559.
[25] NSA Elalal, GM El Seedy and YA Elhassaneen. Chemical composition, nutritional value, bioactive compounds content and biological activities of the brown alga (Sargassum Subrepandum) collected from the mediterranean Sea, Egypt. Alexandria Science Exchange Journal 2021; 42(4), 893-906.
[26] S Palaniyappan, A Sridhar, ZA Kari, T Guillermo and T Ramasamy. Evaluation of phytochemical screening, pigment content, in vitro antioxidant, antibacterial potential and GC-MS metabolite profiling of green seaweed caulerpa racemosa. Marine Drugs 2023; 21(5), 278.
[27] MKA Sobuj, A Islam, S Islam, M Islam, Y Mahmud and SUM Rafiquzzaman. Effect of solvents on bioactive compounds and antioxidant activity of Padina tetrastromatica and Gracilaria tenuistipitata seaweeds collected from Bangladesh. Scientific Reports 2021; 11(1), 19082.
[28] N Astalakshmi, T Gokul, KBG Sankar, M Nandhini, HH Sudhan, S Gowtham, ST Latha and MS Kumar. Over view on molecular docking: A powerful approach for structure based drug discovery. International Journal of Pharmaceutical Sciences Review and Research 2022; 77(2), 180-198.
[29] A Abudurexiti, R Zhang, Y Zhong, H Tan, J Yan and S Bake. Identification of α -glucosidase inhibitors from Mulberry using UF-UPLC-QTOF-MS / MS and molecular docking. Journal of Functional Foods 2023; 101, 105362
[30] A Khaldan, S Bouamrane, R El-mernissi, H Maghat, M Aziz, A Sbai, M Bouachrine and T Lakhlifi. 3D-QSAR modeling , molecular docking and ADMET properties of benzothiazole derivatives as a-glucosidase inhibitors. Materials Today: Proceedings 2021; 45, 7643-7652.
[31] HK Rundla, S Soni, S Teli, A Manhas, PC Jha, S Agarwal and LK Agarwal. Theophylline hydrogen sulfate as a highly efficient catalyst for the synthesis of quinoxaline derivatives: Exploring potential antidiabetic agents through molecular docking. Catalysis Letters 2025; 155(243), 243.
[32] P Banerjee, AO Eckert, AK Schrey and R Preissner. ProTox-II: A webserver for the prediction of toxicity of chemicals. Nucleic Acids Research 2018; 46, W257-W263.
[33] M Lacueva-Arnedo, A Gómez-Barrio, A Ibáñez-Escribano, TJ López-Pérez, B Casarrubias-Tabarez, F Calzada, PY López-Camacho and N Rivera-Fernández. Antibacterial, trichomonacidal, and cytotoxic activities of pleopeltis crassinervata extracts. Pharmaceutics 2024; 16(5), 624.
[34] I Oktavianawati, U Wulandari, INA Winata, AA Ridho, YI Kedang, DT Fauziah and F Yusro. The volatile compound profiles of Fire-cured and fermented Na-oogst tobacco leaves (Nicotiana tabacum L.) and its in-silico study. Indonesian Chimica Letters 2025; 4(1), 7-15.
[35] OM Oluwakeyede and BA Odeyemi. GC-MS profiling of bioactive compounds in ethanol extract of Annona Squamosa L. Leaves. Journal of Chemical Society of Nigeria 2025; 50(4), 747-755.