
Trends
Sci.
2026;
23(7):
11687
Exploratory Analysis of the Anti-Inflammatory Potential of the Ethyl Acetate-Soluble Fraction of Selaginella doederleinii Hieron in IL-1β-Induced Rat Hepatocytes
Honesty Nurizza Pinanti1, Risa Tanaka2, Yosuke Saito2, Keita Minamisaka2,
Kaho Takayasu2, Mikio Nishizawa3 and Muhammad Sasmito Djati4,5,*
1Department of Biology, Faculty of Mathematics and Natural Sciences, State University of Surabaya,
East Java, Indonesia
2Department of Biomedical Sciences, Graduate School of Life Sciences, College of Life Sciences,
Ritsumeikan University, Biwako Kusatsu Campus, Shiga, Japan
3College of Life Sciences, Ritsumeikan University, Biwako Kusatsu Campus, Shiga, Japan
4Department of Biology, Faculty of Mathematics and Natural Sciences, Brawijaya University, East Java, Indonesia
5Dewan Jamu East Java Region, East Java, Indonesia
(*Corresponding author’s e-mail: [email protected])
Received: 25 August 2025, Revised: 24 December 2025, Accepted: 31 December 2025, Published: 1 March 2026
Abstract
Interleukin-1β (IL-1β), a key pro-inflammatory cytokine, promotes inducible nitric oxide synthase (iNOS) expression in hepatocytes, leading to excessive nitric oxide (NO) production and liver inflammation. Selaginella doederleinii Hieron is a medicinal plant traditionally used to treat various diseases, but its anti-inflammatory potential in hepatic inflammation models remains insufficiently explored. This exploratory study evaluated the anti-inflammatory potential of crude fractions of S. doederleinii, with implications for hepatoprotection, using an IL-1β-induced primary rat hepatocyte model and explored potential underlying molecular mechanisms. Fractions A (ethyl acetate), B (n-butanol), and C (aqueous) were assessed for NO inhibition, iNOS expression, and cytotoxicity. Fraction A (ethyl acetate) exhibited the strongest NO inhibitory activity (IC₅₀ = 15.69 µg/mL), showed no cytotoxicity, and qualitatively displayed a trend toward reduced iNOS expression. The representative Western blot analysis of iNOS provided visual support for the NO inhibition observed in the assay. Exploratory transcriptomic profiling suggested altered expression of genes related to the NF-κB and TNF pathways, including Rela and Nos2, thereby providing hypothesis-generating insights. LC–HRMS analysis tentatively annotated several compounds in Fraction A. In silico docking predicted that emodin-8-O-β-D-glucopyranoside showed relatively higher binding potential to IL-1β at site A, suggesting a possible interference with the IL-1β/IL-1R1 interaction. Collectively, these findings provide preliminary and exploratory in vitro and in silico evidence that the ethyl acetate-soluble fraction of S. doederleinii may exert anti-inflammatory activities in IL-1β-induced hepatocytes, warranting further validation.
Keywords: Anti-inflammatory activity, Hepatocyte, IL-1β, Inflammation, iNOS, NF-κB, Nitric oxide, Selaginella doederleinii, Hieron
Introduction
Inflammation in the liver can be induced by pathogen infections, including those caused by bacteria and viruses [1]. Lipopolysaccharides from bacteria’s outer membrane promote Kupffer cells to release IL-1β [2]. However, liver inflammation can also develop without pathogenic infection, where sterile injury or cellular stress is involved in the development of non-
alcoholic and alcoholic steatohepatitis as well as drug-induced liver injury [1,3]. Elevated levels of IL-1β and inducible nitric oxide synthase (iNOS or NOS2) expression have indeed been reported in several drug-induced liver injury models, particularly those induced by acetaminophen and cisplatin [4,5]. In hepatocytes, IL-1β activates iNOS, leading to excessive nitric oxide (NO) production [6,7]. This NO reacts with superoxide anions to produce peroxynitrite [8]. NO and peroxynitrite are members of the reactive nitrogen species (RNS), exerting destructive effects and nitrosative stress on cells [9]. Therefore, inhibition of NO production is commonly used as an indicator of anti-inflammatory activity. Furthermore, NO can be used as a marker to determine the potency of medicinal plant extracts in hindering inflammation [10].
Although conventional anti-inflammatory agents, including steroidal and non-steroidal classes, are widely prescribed, their prolonged use is often limited by adverse effects, thereby promoting the exploration for safer alternatives [11]. Medicinal plants, which contain abundant bioactive compounds, have gained increasing attention as potential therapeutic agents, as they can modulate multiple inflammatory signaling pathways with minimal side effects [12]. Among them, Selaginella doederleinii Hieron is a pteridophyte predominantly found in China, South Korea, and Southeast Asia. The plant has long been traditionally used to manage or relieve symptoms of cancers, cardiovascular diseases, rheumatoid arthritis, and sore throat [13]. A previous study revealed that S. doederleinii contains bioactive compounds, such as biflavonoids, β-citronellolvanillic acid, chrysophanol, ferulic acid, and several others, which have been reported to exert pharmacological properties, including anticancer, anti-inflammation, and antioxidant [14,15]. Furthermore, many studies have investigated the biological activities of S. doederleinii fractions, particularly its anticancer properties [13,16,17]. However, the anti-inflammatory activity of S. doederleinii crude fractions against inflamed hepatocytes remains underexplored. Therefore, this exploratory study aimed to evaluate the anti-inflammatory potential of crude fractions of S. doederleinii Hieron, with potential implications for hepatoprotection, against IL-1β-induced hepatic inflammation and to provide preliminary insights into their possible molecular mechanisms.
Materials and methods
Plant material
Aerial part samples of S. doederleinii were procured from UPT Materia Medica, Batu, Indonesia (7°52'01.2"S and 112°31'13.2"E) in October 2022. The determination number of the specimen was 074/695/102.20-A/2022, authenticated by UPT Materia Medica. The powder of the plant was also obtained from the same institution.
Extraction and crude fractionation
The extraction and fractionation were performed using methods described by Ohno et al. [18]. S. doederleinii powder was extracted with absolute methanol (1:10 ratio) under reflux at 50 ℃ for an hour twice. The total filtrate was evaporated at above 50 ℃ until the solvent had entirely evaporated. Furthermore, the methanol extract was used for the ABC fractionation with ethyl acetate and n-butanol, yielding the A, B, and C fractions, which were soluble in ethyl acetate, n-butanol, and water, respectively (Figure 1). First, the methanol extract dissolved in laboratory-grade Elix water produced by an in-house purification system was added to ethyl acetate (1:2 ratio) in a separatory funnel, vigorously shaken, and incubated to obtain two different layers. The ethyl acetate-soluble fraction was separated from the aqueous-soluble fraction. The procedure was done in triplicate. Subsequently, the n-butanol mixture (4×n-butanol: 1×elix water) was mixed with the aqueous layer (1:1 ratio) in a separatory funnel and strongly shaken to obtain n-butanol and aqueous-soluble fractions. The fractionation steps were also performed in three replicates. Each fraction was evaporated at above 70 ℃.
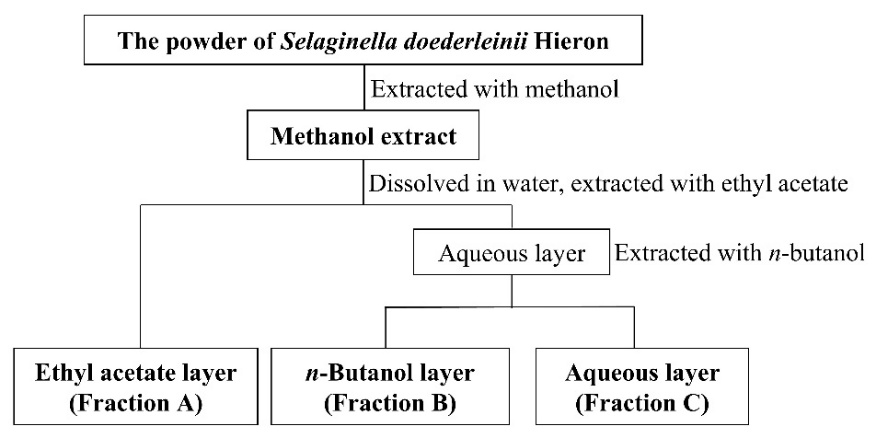
Figure 1 Flow chart of ABC fractionation of S. doederleinii extract.
Primary cultured rat hepatocytes
Wistar rats (Rattus norvegicus; male; 5 - 6 weeks of age; 250 - 300 g of body weight; specific pathogen-free) were acquired from Charles River Laboratories Japan, Inc., Yokohama, Japan. The rats were maintained at 21 - 23 ℃ for one week to acclimate, with the radiation-induced CRF-1 diet (Charles River Laboratories, Japan) and unrestricted access to water. The rats were anesthetized with sodium pentobarbital. Every attempt was made to reduce animal pain. The collagenase perfusion method was used to detach hepatocytes from the liver [19]. The hepatocytes were cultured in the Williams’ E (WE) medium (Sigma-Aldrich Corp., St. Louis, USA) at 1.2×106 cells per dish and maintained for 2 h at 37 °C [10]. The medium substitution was conducted twice before the overnight incubation. The Japanese government’s laws and regulatory guidelines have been applied to all animal handling and experimental practices. This study has obtained ethical approval from the Animal Care Committee of Ritsumeikan University, Biwako-Kusatsu Campus (No. BKC2021-031).
NO and Lactate Dehydrogenase (LDH) assays
IL-1β-induced hepatocytes were incubated with S. doederleinii crude fraction at 37 °C for 8 h. Rat IL-1β (PeproTech, Rocky Hill, NJ, USA) was added at 1 nmol/L to trigger inflammation, as hepatocytes lack TLR4 expression and do not respond to LPS stimulation. Both endotoxin and residual solvent checks were performed to ensure assay reliability. All fractions were evaporated to dryness to minimize residual solvent before conducting biological assays. The final concentrations of DMSO were set to below 1.0 %(v/v) in the medium [20]. The Griess method was carried out to determine the NO level through the amount of nitrite in the cell-free medium, whereas the LDH assay was done to check the possible toxicity of the crude fraction. All samples were measured in technical triplicate from three independent biological replicates. The IC50 value of NO suppression was calculated when the crude fraction was considered non-toxic to hepatocytes. For the NO assay, the cell medium was treated with Griess reagent (Promega, USA) in the 96-well plate [21]. Although a pharmacological positive control was not included, this assay system has been validated in previous studies [22]. Therefore, the results should be interpreted within the context of this previously validated experimental system. In addition, fresh culture medium contained < 5% of the nitrite measured in whole-cell extract, indicating that the majority of nitrite detected originated from hepatocyte activity rather than the medium. To further corroborate assay performance, vehicle controls (DMSO at the final concentration used) were included in all experiments. Sodium nitrite was also utilized as the standard for this assay. Absorbance at 540 nm was determined after a 5-min incubation period. For the LDH assay, cells were lysed in 2% Triton X-100 to prepare whole cell extracts. The cell medium and the supernatant of whole cell extracts were diluted 10 and 100 times, respectively, added with CytoTox96 Reagent (Promega, USA) in the 96-well plate for 30 min, mixed with the stop solution, and detected their absorbance at 490 nm. The standard for this assay was LDH.
Western blot
Rat hepatocytes administered with S. doederleinii crude fraction were incubated for 8 h. IL-1β was also introduced to the cell medium to stimulate hepatocyte inflammation. Cells were lysed using 1×SDS sample buffer supplemented with a protease inhibitor cocktail. Pooled lysates prepared from three independent hepatocyte isolations were used for each treatment condition. Proteins were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) before transfer to a Sequi-Blot membrane (Bio-Rad, Hercules, USA). Primary antibodies used were mouse monoclonal antibody against rat iNOS (1:1000; Thermo Fisher Scientific, Waltham, MA, USA) and rabbit polyclonal antibody against rat β-tubulin (1:1000; Cell Signaling Technology Inc., Danvers, MA, USA). Secondary antibodies were horseradish peroxidase–conjugated anti-mouse IgG (1:5000; Cell Signaling Technology Inc.) for iNOS and horseradish peroxidase–conjugated anti-rabbit IgG (1:1000; Cell Signaling Technology Inc.) for β-tubulin. Bands were visualized using Enhanced Chemiluminescence Blotting Detection Reagent (GE Healthcare Biosciences Corp., Piscataway, USA), with exposure times of 60 s for iNOS and 300 s for β-tubulin. The Western blots shown are representative, illustrative only, and no statistical analyses were performed for Western blot data.
Microarray and functional enrichment analysis
Rat hepatocytes administered with 20 µg/mL of S. doederleinii crude fraction were incubated for 4 h with or without IL-1β stimulation. The purification of whole RNA was conducted using the RNAqueous kit (Applied Biosystems, CA, USA), while the quality control examination of RNA was carried out with the 2100 Bioanalyzer instrument (Agilent, CA, USA). The fragmentation and labeling of RNA were done with the GeneChip™ WT PLUS Reagent Kit (Thermo Fisher Scientific, MA, USA). Clariom™ S Assay, rat (Thermo Fisher Scientific, MA, USA) was used to analyze the gene expression profiles of the rat hepatocytes. The GeneChip™ Scanner 3000 7G (Thermo Fisher Scientific, MA, USA) and Microarray Data Analysis Tool Ver 3.2 (Filgen Inc., Japan) were utilized for scanning the array and analyzing the data, respectively. Microarray experiments were performed without biological replicates for each condition. Because of the absence of replicates, statistical estimation of variance, moderated t-tests, and false-discovery rate (FDR) correction could not be conducted. Accordingly, differentially expressed genes (DEGs) were identified using fold-change thresholds alone. Genes exhibiting a signal ratio ≥ 2.0 were considered increased transcripts, while genes exhibiting a signal ratio ≤ 0.5 were considered decreased transcripts [22]. Due to these limitations, all transcriptomic results should be interpreted as exploratory and hypothesis-generating rather than confirmatory. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were executed with the R package clusterProfiler (v4.6.2) and ShinyGO (v0.77), respectively [23,24]. A heatmap of differentially expressed genes (DEGs) related to the inflammatory pathway and immune system was generated using the R package ggplot2 (v3.5.1) [25]. A protein–protein interaction (PPI) network was mapped from the DEGs using the STRING database (https://string-db.org/) at the highest confidence level (score ≥ 0.9). The visualization of the PPI network was carried out with Cytoscape (v3.10.2), with unconnected nodes removed [26].
Liquid chromatography-high-resolution mass spectrometry (LC-HRMS) analysis
LC-HRMS analysis of fraction A from Selaginella doederleinii was performed using a Waters ACQUITY UPLC system with a high-resolution quadrupole time-of-flight mass spectrometer equipped with an electrospray ionization (ESI) source operating in positive ion mode. Chromatographic separation was conducted on an ACQUITY UPLC® BEH C18 column (1.7 μm) at 40 °C. The mobile phase included 0.1% formic acid in water (A) and acetonitrile (B), at a flow rate of 0.30 mL min⁻¹ using a linear gradient from 95 to 10 %A over 10 min, held for 3 min, and re-equilibrated to initial conditions until 16 min. The sample was injected at a volume of 1 μL. Mass spectrometric detection was conducted in positive ESI mode over an m/z range of 50 - 1,200. The source temperature was set at 140 °C with a desolvation temperature of 550 °C. Lock-mass correction was applied to ensure mass accuracy. Data were acquired using low- and high-collision energy functions. Data processing and tentative compound identification were performed using UNIFI software (Waters). Compounds reported in this study were selected based on the quality of LC-HRMS annotation and their relevance to subsequent biological analyses.
Molecular docking simulation
The crystal structure of IL-1β (PDB: 5r88) was acquired from the RCSB PDB (https://www.rcsb.org/). Non-essential molecules were deleted from proteins. The tested ligands utilized for docking were pyrophaeophorbide A, emodin-8-O-β-D-glucopyranoside, stearidonic acid, and trichosanic acid. Their structures were sourced from PubChem (https://pubchem.ncbi.nlm.nih.gov/). (2~{S})-~{N}-(4-aminocarbonylphenyl)oxolane-2-carboxamide was used as the reference compound, obtained from the crystal structure of the IL-1β. Open Babel in PyRx 0.8-Virtual Screening Tools software (https://pyrx.sourceforge.io/) was applied for ligand energy minimization. Autodock Vina in the software was employed to perform the specific molecular docking at sites A and B of IL-1β [27-30]. The center position of Site A docking was X = 51.6363, Y = 11.2069, Z = 75.2231, and the dimensions were X = 26.4819, Y = 30.2513, Z = 27.2413. The center position of Site B docking was X = 37.2516, Y = 21.9145, Z = 62.1851, and the dimensions were X = 35.7760, Y = 31.8549, Z = 35.5607. Redocking of the reference ligand was performed to validate the docking protocol, reproducing the crystallographic pose with RMSD < 2 Å. Biovia Discovery Studio v21.1.0.20298 (Dassault Systèmes Biovia, San Diego, USA) was applied for docking visualization [31].
Statistical analysis
Statistical analyses were applied only to NO and LDH assay data. The values are presented as the means ± standard deviation (SD) (n = 3) from three independent experiments. Statistical differences between IL-1β-stimulated control and treatment groups were analyzed using Student’s t-test followed by Bonferroni correction. Significance levels were set at p < 0.05 and p < 0.01.
Results
Fractionation of S. doederleinii extract to crude fractions
The fractionation of S. doederleinii extract using the ABC fractionation method resulted in fractions A, B, and C, soluble in ethyl acetate, n-butanol, and water with yield percentages of 48.06%, 11.67%, and 40.27%, respectively (Table 1). Therefore, fraction A, containing hydrophobic constituents, was the most abundant fraction in the methanolic extract of S. doederleinii. Meanwhile, fraction B, containing amphipathic constituents, had the smallest yield percentage among all fractions.
Table 1 The result of the extraction and fractionation of S. doederleinii.
Material |
Weight (g) |
Yield (%) |
S. doederleinii powder |
68.77 |
- |
Methanol extract |
9.51 |
100 |
Fraction A |
4.57 |
48.06 |
Fraction B |
1.11 |
11.67 |
Fraction C |
3.83 |
40.27 |
The NO level and iNOS expression of IL-1β -induced rat hepatocytes affected by S. doederleinii crude fractions
According to the results of the Griess test, fractions A and B of S. doederleinii inhibited the NO level in IL-1β -induced hepatocytes, displaying a concentration-dependent effect. Fractions A and B could significantly diminish NO production at 20 and 160 µg/mL, respectively (Figure 2(A)). Meanwhile, NO production was not significantly altered by fraction C. Furthermore, to exclude the possible toxicity of S. doederleinii crude fractions, which might affect their anti-inflammatory activity, we conducted the LDH assay [32]. LDH percentage of the total cell extract was designed as 100%. According to the results, the LDH activity percentages of all crude fractions and control groups were very low, indicating that all crude fractions of S. doederleinii at the tested concentration ranges were non-toxic to hepatocytes (Figure 2(B), Supplementary File 1). Furthermore, Western blot analysis is presented as an illustrative depiction of iNOS protein patterns. Because the blot was generated from pooled lysates and lacks biological replicates per lane, it is not used for quantitative interpretation or statistical analysis. The visual pattern of the bands appeared consistent with the inhibitory trend observed in the NO assay for fractions A and B (Figure 2(C), Supplementary File 2). A qualitative reduction in iNOS band intensity was observed for fraction A with increasing concentration. These results indicated that fractions A and B of S. doederleinii could hinder NO generation, while Western blot data qualitatively supported a possible reduction in iNOS expression. Fraction A had the smallest IC50 score (15.69 µg/mL) (Table 2). Meanwhile, fraction B exhibited a higher IC50 value, making it less effective than fraction A. Fraction C had a very low suppressing effect on NO production.
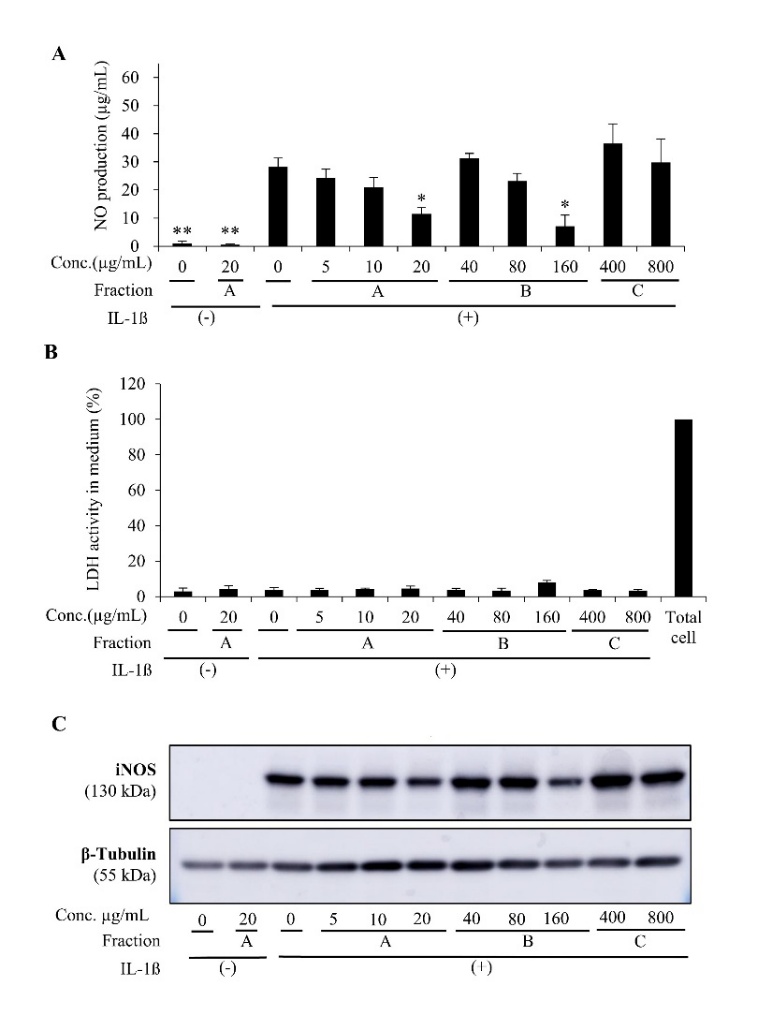
Figure 2 Inhibitory effect of S. doederleinii fractions on NO level and iNOS expression in IL-1β-induced hepatocytes. A. The total medium NO level after treatment with fractions (A) - (C), which were soluble in ethyl acetate, n-butanol, and water, respectively. The results displayed are means ± SD (n = 3) *p < 0.05 and **p < 0.01 vs IL-1β alone; B. LDH release level into the hepatocytes’ medium affected by the crude fractions. The total cell was the positive control with 100% LDH activity. The data are expressed as means ± SD (n = 3); C. Representative Western blot images of iNOS (130 kDa) and β-tubulin (55 kDa) obtained from pooled lysates. Because the Western blot was performed without biological replicates, these bands are illustrative only and not used for quantitative or statistical interpretation.
Table 2 IC50 values of S. doederleinii crude fractions in inhibiting medium NO levels.
Fraction of S. doederleinii |
IC50 (µg/mL)* |
Fraction A |
15.69 ± 3.96 |
Fraction B |
105.37 ± 29.65 |
Fraction C |
NA |
*NO concentration in the medium of IL-1β -induced hepatocytes was measured. The IC50 value was determined from three independent assays and shown as the mean ± SD.
GO and KEGG enrichment analysis
To explore potential molecular pathways associated with the anti-inflammatory effects of Fraction A on nitric oxide suppression, an exploratory microarray study followed by GO and KEGG pathway enrichment analyses was conducted. The results of microarray analysis showed an increase in 1,554 transcripts (signal ratio ≥ 2) and a decrease in 1,751 transcripts (signal ratio ≤ 0.5) after administration of the fraction (Supplementary Files 3, 4). The decreased transcripts were slightly larger than the increased ones. The ratio between decreased and increased transcripts was 1.13. Because this transcriptomic analysis was performed without biological replicates, all interpretations derived from these data should be considered exploratory and hypothesis-generating rather than confirmatory. The findings of GO analysis, as presented in Figure 3, revealed groups of genes downregulated after administration of the fraction, which were assigned to three categories: Biological Process (BP), Cellular Component (CC), and Molecular Function (MF). In the BP category, some groups of genes refer to the NF-κB molecular signaling pathway, reported to play an essential role in modulating inflammation by producing nitric oxide, including positive regulation of I-κB kinase/NF-κB signaling; I-κB kinase/NF-κB signaling; regulation of I-κB kinase/NF-κB signaling; and positive regulation of NF-κB transcription factor activity [33]. One of these groups showed a lower p-adjusted value than the others, namely positive regulation of I-κB kinase/NF-κB signaling (p-adjusted = 2.095×10⁻⁸). This suggests that treatment with Fraction A may be associated with a possible attenuation of upstream molecular events leading to NF-κB pathway activation. In the CC category, the transcription regulator complex had the lowest p-adjusted value (p-adjusted = 1.21×10⁻⁸). It encodes proteins that form a complex to regulate gene transcription, thus serving a critical function in controlling gene expression in the nucleus. It implies that Fraction A may influence nuclear transcriptional regulation, potentially involving NF-κB-associated processes. In the MF category, the gene group with the lowest p-adjusted value was transcription coregulator activity (p-adjusted = 4.62×10⁻¹⁵), which encodes proteins that can indirectly modulate gene transcription activity, either as co-activators or co-repressors. It suggests a possible reduction in transcriptional coregulator function following treatment.
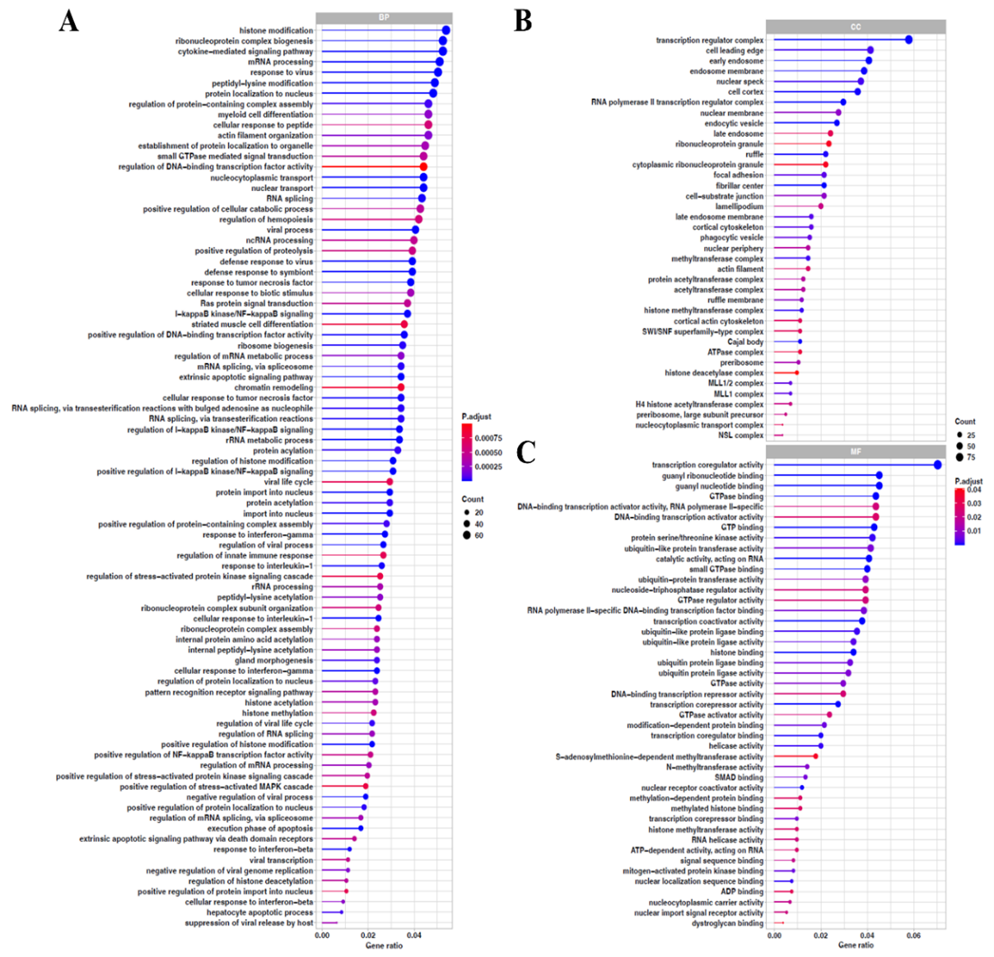
Figure 3 GO enrichment analysis of downregulated DEGs. (A) Biological process (FDR < 0.001). (B) Cellular component (FDR < 0.05). (C) Molecular function (FDR < 0.05).
KEGG analysis showed the molecular signaling pathways whose associated genes exhibited decreased expression after administration of Fraction A (Figure 4). Based on these results, several molecular pathways, such as the TNF signaling pathway, NF-κB signaling pathway, MAPK signaling pathway, Toll-like receptor signaling pathway, and NOD-like receptor signaling pathway, are frequently reported to be involved in inflammation following IL-1β induction [34-36]. Among these pathways, the TNF and NF-κB signaling pathways had lower p-adjusted values than the others, namely 1.87×10−7 and 6.94×10−6, respectively. Therefore, it indicates a potential attenuation of these pathways. Interestingly, several genes, including Tab1, Rela, Nfkb1, and LOC103694380 (a gene predicted to be homologous to Tumor Necrosis Factor), were present in all five pathways previously mentioned.
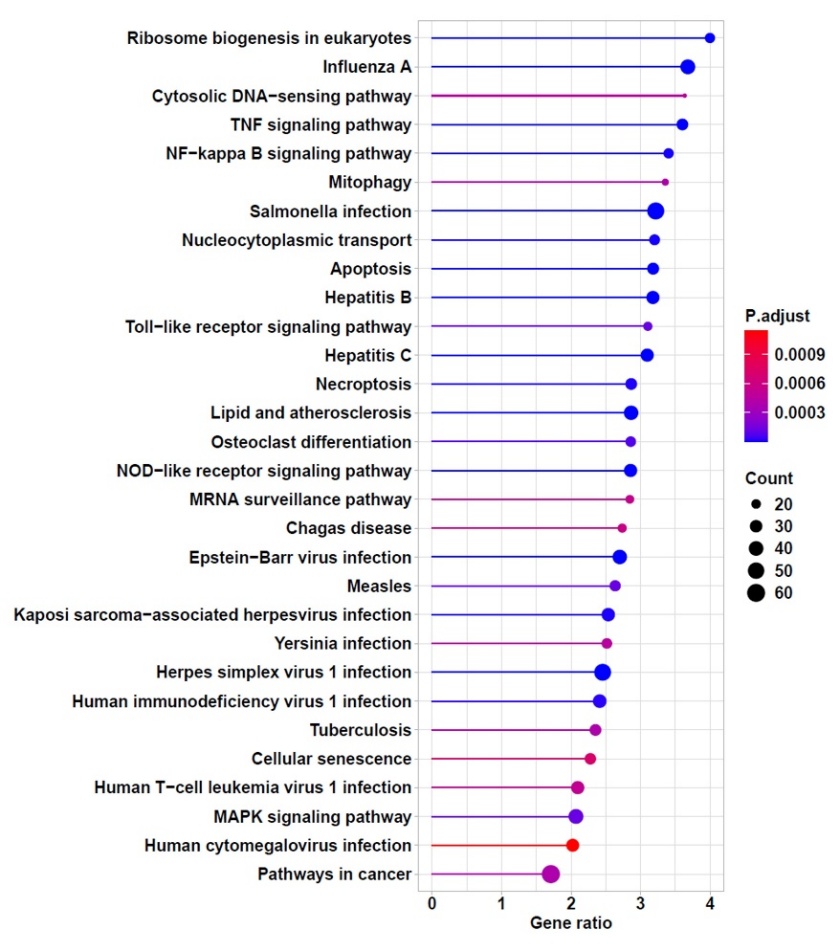
Figure 4 KEGG enrichment analysis of downregulated DEGs.
Analysis of the heatmap of selected inflammatory and immune DEGs
A heatmap of the expression profiles of 15 genes related to the molecular pathway activated by IL-1β, which subsequently modulates the NFκB and TNF pathways, leading to Nos2 expression, is presented in Figure 5. IL-1β, once induced in hepatocytes, binds to its receptor, IL-1R1. Subsequently, this complex modulates the NFκB and TNF pathways, thereby promoting Nos2 expression and ultimately increasing NO production [4]. In this study, administration of Fraction A decreased the expression of all these genes in the heatmap, except Hdac and Prkca. The largest decrease (control vs. Fraction A) was observed in the Rela and Nos2 genes. Because these observations originate from an exploratory microarray dataset, they require validation through qRT-PCR and additional functional assays in future studies.
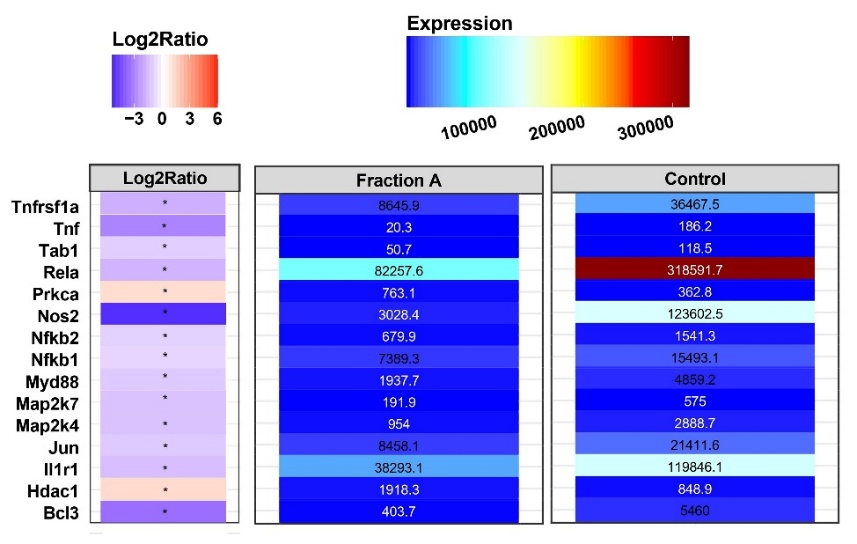
Figure 5 Heatmap of genes involved in IL-1β induction leading to Nos2 expression activation.
PPI network analysis
Based on the downregulated DEGs obtained from the microarray analysis, a protein-protein interaction (PPI) network was generated to study the potential interactions among the proteins encoded by these genes. Previous analysis revealed that Rela, the gene encoding the RelA protein, also known as the p65 subunit of NF-κB, showed the greatest decrease in expression after administration of Fraction A. PPI network analysis (highest confidence score, ≥ 0.9) revealed that RelA interacts with 14 other proteins (degree value = 14) (Figure 6), a higher number than any other proteins directly related to the NF-κB and TNF pathways in this study. Full visualization of the PPI network constructed in this study is available as Supplementary File 5. Within the exploratory framework of the microarray analysis, this pattern highlights RelA as a potential key node in the affected signaling network. Among the 14 proteins directly interacting with RelA, several are encoded by the Nfkb1, Nfkb2, and Nfkbib genes, namely the canonical NF-κB subunit p50, the non-canonical subunit p52, and IκBβ, an inhibitor of NF-κB, respectively. The microarray results in this study also showed decreased expression of the Nfkb1, Nfkb2, and Nfkbib. Additionally, Brd4 and Rock2, involved in modulating Nos2 expression, also directly interact with RelA, as shown in the heatmap [37,38]. Genes encoding both proteins exhibited decreased expression following treatment in this study (data not shown).
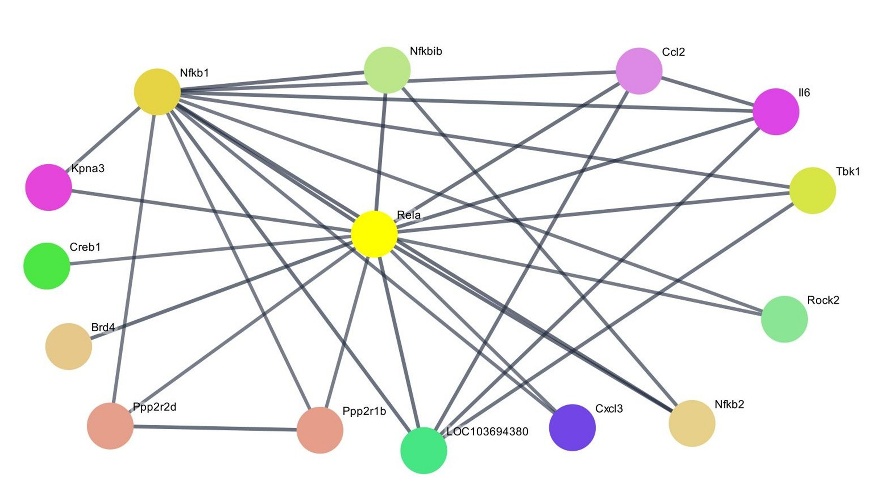
Figure 6 PPI network of RelA as a key protein (highest confidence score, ≥ 0.9).
Analysis of LC-HRMS
LC-HRMS analysis was conducted to profile the bioactive compounds present in Fraction A of Selaginella doederleinii. The total ion chromatogram (TIC) exhibited multiple peaks with different retention times, indicating the presence of chemically diverse components in the fraction (Figure 7(A)). Based on accurate mass measurements and MS² fragmentation patterns, several ion features were tentatively annotated. Among these, four compounds were selected and reported in this study based on the confidence of their annotation and their relevance to subsequent biological analysis, namely pyrophaeophorbide A, emodin-8-O-β-D-glucopyranoside, stearidonic acid, and trichosanic acid. The TIC is presented to illustrate the overall chemical profile of the fraction, while representative MS¹ and MS² spectra are presented for a single compound, pyrophaeophorbide A (Figure 7). MS¹ and MS² data for the remaining annotated compounds were used in the annotation process but are not shown for brevity. Table 3 summarizes the selected compounds along with their retention times (RT), ion types, observed m/z values, and MS² fragmentation evidence, used as the basis for compound annotation.
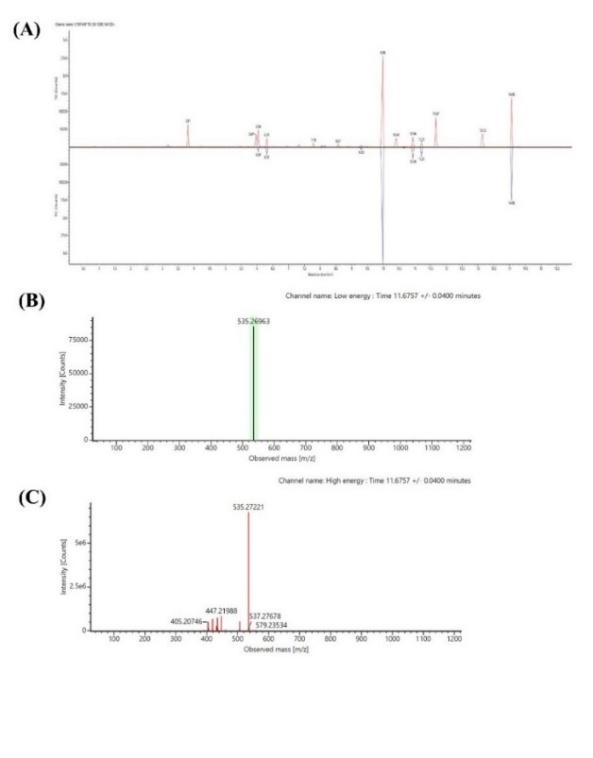
Figure 7 LC-HRMS analysis of Fraction A of Selaginella doederleinii. (A) Total ion chromatogram (TIC) showing the overall chemical profile of Fraction A. (B) Representative MS¹ spectrum of pyrophaeophorbide A acquired in positive ESI mode. (C) Representative MS² fragmentation spectrum of pyrophaeophorbide A used to support tentative compound annotation.
Table 3 Selected tentatively annotated compounds in Fraction A of Selaginella doederleinii based on LC-HRMS.
Compound |
RT (min) |
Ion |
m/z (observed) |
MS² fragmentation evidence |
Pyrophaeophorbide A |
11.68 |
[M+H]⁺ |
535.2696 |
447.22, 405.21 (fragmentation pattern consistent with porphyrin-type compounds) |
Emodin-8-O-β-D-glucopyranoside |
3.19 |
[M+H]⁺ |
433.1123 |
313.07, 283.06, 165.02 (fragmentation pattern consistent with anthraquinone O-glycosides) |
Stearidonic acid |
8.15 |
[M+H]⁺ |
277.2154 |
391.04, 384.38, 93.07 (fragmentation pattern consistent with polyunsaturated fatty acids) |
Trichosanic acid |
10.41 |
[M+H]⁺ |
279.2318 |
425.21, 124.09, 81.07 (fragmentation pattern consistent with long-chain fatty acid derivatives) |
Molecular docking towards IL-1R1
This study used IL-1β to trigger inflammation in hepatocytes. Therefore, the decreased expression of Rela and Nos2 after treatment with Fraction A may be partly explained by potential interference of tentatively annotated constituents identified in this study, namely pyrophaeophorbide A, emodin-8-O-β-D-glucopyranoside, stearidonic acid, and trichosanic acid, with the IL-1β/IL-1R1 interaction. Docking simulations were carried out at two sites on IL-1β that serve as binding sites for IL-1R, namely sites A and B. Site A consists of several amino acid residues, including Arg11, Ser13, Gln14, Gln15, Met20, Ser21, Gly22, Lys27, Leu29, His30, Leu31, Gln32, Gly33, Gln34, Asp35, Met36, Gln38, Gln126, Ala127, Glu128, Asn129, Met130, Pro131, Thr147, and Gln149, while site B consists of several amino acid residues, including Ala1, Pro2, Val3, Arg4, Leu6, Phe46, Gln48, Glu51, Asn53, Asp54, Ile56, Lys92, Lys93, Lys94, Lys103, Glu105, Ile106, Asn108, Lys109, Phe150, and Ser152 [30]. Docking simulations at site A revealed that pyrophaeophorbide A (–7.1 kcal/mol) and emodin-8-O-β-D-glucopyranoside (–6.0 kcal/mol) exhibited more negative binding affinity scores than the reference compound, (2S)-N-(4-aminocarbonylphenyl)oxolane-2-carboxamide (–5.0 kcal/mol), indicating more favorable predicted binding affinity at this site (Table 4). Among the tested ligands, emodin-8-O-β-D-glucopyranoside and trichosanic acid interacted with the same amino acid residues as the reference compound, namely Val40 and Val41 (Figure 8). The similarity of interacting residues suggests a potentially similar inhibitory interaction toward IL-1β at site A. In contrast, pyrophaeophorbide A and stearidonic acid interacted with other residues forming site A, with pyrophaeophorbide A binding to His30 and Leu31, and stearidonic acid interacting with Arg11 and Met36. Trichosanic acid also showed an additional interaction with Met20. Therefore, among the tested ligands, emodin-8-O-β-D-glucopyranoside exhibited a favorable combination of stronger binding affinity and shared interactions with key residues, indicating a higher binding potential toward IL-1β at site A. Meanwhile, docking simulations at site B demonstrated that pyrophaeophorbide A (–7.5 kcal/mol) and emodin-8-O-β-D-glucopyranoside (–7.2 kcal/mol) showed more negative binding affinity scores than the reference compound (–6.1 kcal/mol), suggesting stronger predicted binding at site B. However, none of the tested ligands interacted with the same amino acid residues as the reference compound, indicating a distinct binding mode at site B. In addition, pyrophaeophorbide A and stearidonic acid interacted with other residues constituting site B. Pyrophaeophorbide A interacted with Arg4, Glu51, Phe46, and Ile56, while stearidonic acid interacted with Ile56, Leu6, Phe46, and Phe150. Therefore, among the tested compounds, pyrophaeophorbide A showed strong binding affinity at site B but interacted with IL-1β through a binding mode distinct from that of the reference compound. Overall, the docking results demonstrate distinct ligand interaction profiles at sites A and B of IL-1β, with emodin-8-O-β-D-glucopyranoside showing the highest binding potential as an IL-1β-binding ligand at site A. Taken together, these docking outcomes offer only preliminary predictions that these compounds could interact with IL-1β. Importantly, any relevance of these in silico interactions to real biological binding events or to the transcriptional changes observed in this study requires further experimental confirmation.
Table 4 Chemical Interactions of compounds with IL-1β.
Compound |
PubChem ID |
Binding affinity at Site A (kcal/mol) |
Binding affinity at Site B(kcal/mol) |
Interacting amino acid residues at Site A |
Interacting amino acid residues at Side B |
Categories |
(2~{S})-~{N}-(4-aminocarbonylphenyl)oxolane-2-carboxamide (Reference compound) |
- |
–5.0 |
–6.1 |
Val41, Lys63 (2), Glu37 |
Gln48(2), Glu50
|
Hydrogen bond |
- |
LYS97 |
Electrostatic |
||||
Val40 |
Lys97, Val100, Ala115 |
Hydrophobic interactions |
||||
Pyrophaeophorbide A
|
3819540 |
–7.1 |
–7.5 |
- |
Arg4(2), Glu51(5), Phe46 |
Hydrogen bond |
Glu128 (3)
|
Glu51 |
Electrostatic |
||||
His30 (4) |
Ile56(4), Pro57, Phe46(2)
|
Hydrophobic interactions |
||||
Leu31 |
- |
Unfavorable |
||||
Emodin-8-O-β-D-glucopyranoside |
99649 |
–6.0 |
–7.2 |
Val41(2) |
Lys65, Glu64, Ser5, Ser43(2), Tyr68, Pro91 |
Hydrogen bond |
- |
LYS63 |
Unfavorable |
||||
Stearidonic acid |
5312508 |
–4.1 |
–4.7 |
Arg11, Thr9 |
Val47 - |
Hydrogen bond |
Val151, Met36 (2) |
Ile56, Leu6(2), Phe46(3), Phe150 |
Hydrophobic interactions |
||||
Trichosanic acid |
5281126 |
–4.0 |
–3.8 |
Met20 |
Tyr68, Leu62, Lys65 |
Hydrogen bond |
Val40(2), Val41, Lys65, Met20 (3), Leu62 |
Pro91(3) |
Hydrophobic |
||||
Lys65 |
- |
Unfavorable |
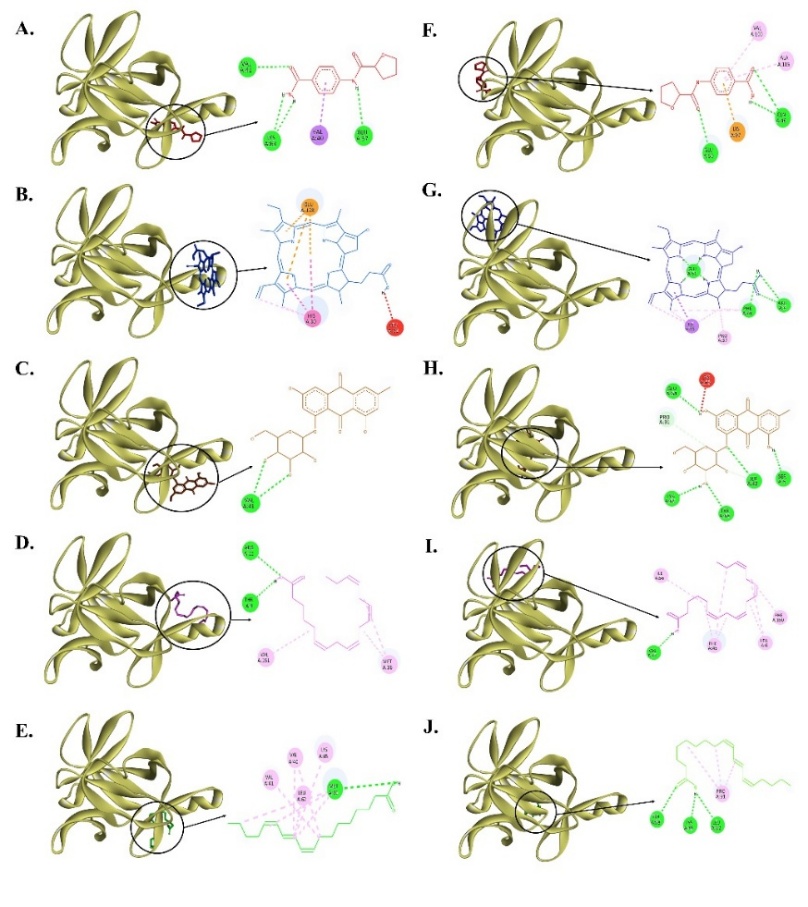
Figure 8 Interaction of compounds with amino acid residues in sites A and B of IL-1β protein. (A) (2~{S})-~{N}-(4-aminocarbonylphenyl)oxolane-2-carboxamide (reference compound) (red) at Site A, (B) Pyrophaeophorbide A (blue) at Site A, (C) Emodin-8-O-β-D-glucopyranoside (brown) at Site A, (D) Stearidonic acid (magenta) at Site A, (E) Trichosanic acid (green) at Site A, (F) (2~{S})-~{N}-(4-aminocarbonylphenyl)oxolane-2-carboxamide (reference compound) (red) at Site B, (G) Pyrophaeophorbide A (blue) at Site B, (H) Emodin-8-O-β-D-glucopyranoside (brown) at Site B, (I) Stearidonic acid (magenta) at Site B, and (J) Trichosanic acid (green) at Site B.
Discussion
Selaginella doederleinii Hieron is a wild Asian plant that has been used for centuries by Asians to treat various diseases due to its therapeutic properties [13]. This research evaluates the anti-inflammatory potential of three crude fractions of S. doederleinii in IL-1β-induced hepatocytes by assessing NO production and iNOS expression. In addition, molecular pathway analyses of the most active fraction were performed using microarray and enrichment analysis, while docking provided in silico predictions of interactions between several tentatively annotated constituents identified using LC-HRMS and IL-1β protein.
This study uses three different crude fractions of S. doederleinii, namely fraction A (ethyl acetate-soluble fraction), fraction B (n-butanol-soluble fraction), and fraction C (water-soluble fraction). The crude fractions were obtained from the fractionation of the S. doederleinii extract using the separatory funnel. The purpose was to separate the bioactive compounds in the extract based on their polarity using different solvents [39]. A crude extract generally contains a complex mixture of metabolites. Fractionation facilitates the isolation of certain bioactive compound groups with advantageous medicinal properties from the metabolite mixture to enhance their purity and biological effects. Ethyl acetate, n-butanol, and aqueous-soluble fractions majorly contain hydrophobic, amphipathic, and hydrophilic constituents, respectively. Certain fractions may exert stronger biological activity than the initial extract [40,41].
Fraction A exhibited the strongest inhibition of NO production (IC₅₀ = 15.69 µg/mL) and qualitatively showed a trend toward reduced iNOS protein expression in IL-1β-induced hepatocytes. IL-1β, interacting with its receptor, can induce iNOS expression and NO synthesis through inflammatory signaling pathways [34]. While NO has physiological roles, excessive iNOS-derived NO in hepatocytes forms reactive nitrogen species, causing macromolecular damage and liver inflammation [8,42,43]. Therefore, the NO inhibitory activity of fraction A suggests potential anti-inflammatory effects, with possible relevance to hepatocyte protection [44]. Additionally, we investigated the potential cytotoxicity of S. doederleinii crude fractions. All crude fractions, including fraction A, did not exhibit any notable cytotoxicity in the primary culture of rat hepatocytes. According to previous reports, the ethyl acetate-soluble fraction of this plant demonstrated no acute toxicity in mice and robust antioxidant activity (IC₅₀ value = 12.5 µg/mL) [45,46]. The antioxidant effect of the fraction may protect cells from NO free radicals and their deleterious effects. Bioactive compounds can donate their electrons to free radicals to maintain cellular redox balance and prevent oxidative/nitrosative damage [47]. In this study, all fractions were dissolved in DMSO at a final concentration ≤ 1 %(v/v), minimizing solvent effects. Western blotting was performed on pooled lysates and is shown as representative data. Although biological replicates were not included for the Western blot analysis, the observed iNOS expression pattern was consistent with the NO assay results. Functional conclusions are therefore based primarily on the NO assay. In addition, the present study did not include a pharmacological positive control, such as a known iNOS inhibitor or standard anti-inflammatory compound. Thus, the inhibitory potency of Fraction A cannot be directly benchmarked against established agents and should be interpreted cautiously. Future work will focus on incorporating replicates, standard comparators, and additional validation.
Microarray and GO enrichment analyses revealed that Fraction A downregulated genes involved in the NFκB signaling pathway, especially those linked with the regulation of gene transcription in the nucleus, including genes encoding transcriptional co-activators and co-repressors. Additionally, KEGG enrichment analysis also indicated that the TNF and NF-κB pathways are two signaling pathways whose molecular activity decreased after treatment with Fraction A. IL-1β activates NF-κB and MAPK pathways through MyD88/IRAK/TAK1 signaling, leading to iNOS induction and NO overproduction [48,49]. TAK1 further stimulates NF-κB, JNK, and p38 pathways, where NF-κB activation through IKKβ-mediated IκBα degradation allows binding to pro-inflammatory gene promoters such as Nos2 [6,50]. Activated NF-κB, p38, and JNK also increase TNF expression, creating crosstalk as TNF binds TNFR1 and recruits TRADD, TRAF2, RIP1, and c-IAP1/2 to reactivate NF-κB and MAPK via TAK1 [49,51]. Subsequently, MAPK, JNK, ERK1/2, and p38 activate AP-1 (c-Jun/c-Fos), further promoting iNOS transcription [50]. Meanwhile, the results of microarray analysis also reveal decreased expression of several genes important in the inflammatory pathway, including Rela and Nos2, as shown in the heatmap in Figure 5. Therefore, this result suggests that Fraction A may suppress iNOS expression and NO production, potentially through modulation of the NF-κB and TNF pathways. However, because the microarray analysis lacked biological replicates and DEG identification relied solely on fold-change, these results should be interpreted as preliminary and hypothesis-generating. Future studies should include replicate-based designs and qRT-PCR validation to confirm key DEGs.
PPI network analysis further supported the involvement of NF-κB signaling, revealing that RelA (p65), a key NF-κB subunit, interacts with p50, p52 (RelB), IκBα/β, Brd4, and Rock2, all encoded by genes downregulated by Fraction A [37,38,52,53]. RelA/p50 forms inactive dimers in the cytoplasm via IκBα, while IκBβ maintains sustained NF-κB signaling [53]. Upon IL-1β or TNF stimulation, IKKβ phosphorylates IκBα, releasing p65/p50 to enter the nucleus and activate proinflammatory genes, including Nos2 [6,50]. Non-canonical p52/RelB dimers, activated via receptors such as CD40, BAFF, and RANKL, also translocate to the nucleus to regulate Nos2 [52]. These interactions indicate that proteins downregulated by Fraction A form a tightly connected NF-κB network, consistent with GO and KEGG analyses [48,49,51]. Support for NF-κB modulation by S. doederleinii has also been reported previously, where extracts derived from the ethyl acetate-soluble fraction were shown to suppress IKKβ and IκBα activity, leading to reduced expression of NF-κB/COX-2 downstream effectors in cellular models [54]. Although this effect was demonstrated in a cancer context and involved a chemically distinct extract, as discussed further below, it underscores NF-κB as a convergent signaling pathway that can be targeted by diverse constituents of S. doederleinii. Given the central role of NF-κB in both tumorigenesis and inflammation, these findings lend further plausibility to the KEGG and PPI-based inference of NF-κB involvement observed in the present study.
LC-HRMS analysis revealed that several compounds were selected and reported in this study based on the quality of their annotation and their relevance to subsequent biological analyses, namely pyrophaeophorbide A, emodin-8-O-β-D-glucopyranoside, stearidonic acid, and trichosanic acid. A previous study revealed that the ethyl acetate fraction of S. doederleinii contains abundant biflavonoids [14]. Differences in the chemical profile of Fraction A compared to previous studies may be attributed to variations in plant origin, growth conditions, post-harvest handling, extraction solvent, fractionation procedures, and analytical conditions, which are known to influence the types of compounds detected by LC-HRMS [55,56]. Furthermore, the molecular docking study suggests that the selected compounds in Fraction A may potentially interfere with IL-1β signaling, which could be associated with altered expression of inflammatory genes in NF-κB and TNF pathways, including Rela and Nos2. Molecular docking indicated that pyrophaeophorbide A and emodin-8-O-β-D-glukopyranoside interact with site A and B of IL-1β, regions involved in receptor recognition, with predicted binding affinities comparable to the reference compound, (2~{S})-~{N}-(4-aminocarbonylphenyl)oxolane-2-carboxamide, a reported IL-1R1/IL-1β inhibitor [57]. Notably, only emodin-8-O-β-D-glucopyranoside showed the highest binding potential as an IL-1β-binding ligand at site A based on binding affinity and shared interactions with key residues. Such interactions suggest a potential interference with the formation of the IL-1β/IL-1R1 complex. Previous studies have reported that emodin-8-O-β-D-glucopyranoside exhibits anti-inflammatory and anti-oxidative effects, supporting the relevance of our docking prediction [58-60]. In contrast, all tested compounds had a different binding mode from the reference compound at site B of IL-1β. However, these findings represent computational predictions only and should not be interpreted as direct evidence of a mechanistic interaction. Further validation using biophysical or cellular assays (e.g., p65 translocation, IκBα degradation, or binding/competition studies) is required. Accordingly, our docking findings primarily serve as hypothesis-generating evidence that complements the transcriptomic and pathway analyses, suggesting a potential molecular basis, which requires experimental validation, for the observed downregulation of NF-κB signaling genes upon treatment with Fraction A.
Overall, our findings provide preliminary, exploratory insights into the anti-inflammatory potential of the ethyl acetate-soluble fraction of S. doederleinii. Further in vivo and mechanistic studies will be essential to substantiate these exploratory results.
Conclusions
The ethyl acetate–soluble Fraction A of Selaginella doederleinii demonstrated preliminary anti-inflammatory activity in an in vitro hepatocyte model, suggesting potential hepatoprotective properties. Fraction A reduced NO release and qualitatively showed a trend toward reduced iNOS expression. Exploratory transcriptomic analysis suggested altered expression of genes associated with the NF-κB and TNF signaling pathways, including Rela and Nos2. Furthermore, LC-HRMS analysis and in silico docking predicted that emodin-8-O-β-D-glucopyranoside, one of the selected tentatively annotated compounds in Fraction A, exhibited the highest binding potential to IL-1β at site A, suggesting a possible interference with the IL-1β/IL-1R1 interaction. Collectively, these findings provide exploratory and hypothesis-generating insights into the possible molecular basis underlying the anti-inflammatory potential of S. doederleinii. Given the limitations of this study, these conclusions should be considered preliminary. Future studies incorporating replicate-based transcriptomic analyses, standard pharmacological positive controls, and in vivo validation are warranted to verify and extend the present findings.
Acknowledgements
We thank the Directorate General of Higher Education, Research and Technology, Ministry of Education, Culture, Research, and Technology of the Republic of Indonesia, for facilitating this study through the Enhancing International Publication Scholarship for PMDSU. We also appreciate the Asia-Japan Research Institute, Ritsumeikan Asia-Japan Research Organization, Ritsumeikan University (Ibaraki, Osaka, Japan) for providing partial support for this study.
Declaration of Generative AI in Scientific Writing
The authors acknowledge that generative AI tools (such as QuillBot and ChatGPT by OpenAI) were utilized during the preparation of this manuscript solely for language editing and grammatical refinement. AI was not used for generating content or interpreting data. Full responsibility for the accuracy, content, and conclusions of this manuscript remains with the authors.
CRediT Author Statement
Honesty Nurizza Pinanti: Conceptualization, methodology, formal analysis, investigation, writing – original draft, visualization, and funding acquisition. Risa Tanaka: formal analysis and investigation. Yosuke Saito: formal analysis and investigation. Keita Minamisaka: formal analysis and investigation. Kaho Takayasu: formal analysis and investigation. Mikio Nishizawa: Conceptualization, methodology, validation, resources, writing– review and editing, project administration, and supervision. Muhammad Sasmito Djati: Conceptualization, methodology, validation, writing– review and editing, and supervision.
[1] AL Kiss. Inflammation in focus: The beginning and the end, pathology and oncology research. Pathology and Oncology Research 2022; 27, 1610136.
[2] C Xu, Y Xia, BW Zhang, EK Drokow, HY Li, S Xu, Z Wang, SY Wang, P Jin, T Fang, XM Xiong, P Huang, N Jin, JH Tan, Q Zhong, YX Chen, Q Zhang, Y Fang, F Ye and QL Gao. Macrophages facilitate tumor cell PD-L1 expression via an IL-1β-centered loop to attenuate immune checkpoint blockade. MedComm 2023; 4(2), e242.
[3] A Allameh, R Niayesh-Mehr, A Aliarab, G Sebastiani and K Pantopoulos. Oxidative stress in liver pathophysiology and disease. Antioxidants 2023; 12(9), 1653.
[4] L Barbier, M Ferhat, E Salamé, A Robin, A Herbelin, JM Gombert, C Silvain and A Barbarin. Interleukin-1 family cytokines: Keystones in liver inflammatory diseases. Frontiers in Immunology 2019; 10, 2014.
[5] AF AlAsmari, M Alharbi, F Alqahtani, F Alasmari, M Al-Swayyed, SI Alzarea, IA Al-Alallah, A Alghamdi, HM Hakami and N Ali. Diosmin alleviates Doxorubicin-Induced liver injury via modulation of oxidative Stress-Mediated hepatic inflammation and apoptosis via NfkB and MAPK pathway: A preclinical study. Antioxidants 2021; 10(12), 1998.
[6] H Inaba, E Yoshigai, T Okuyama, M Murakoshi, K Sugiyama, H Nishino and M Nishizawa. Antipyretic analgesic drugs have different mechanisms for regulation of the expression of inducible nitric oxide synthase in hepatocytes and macrophages. Nitric Oxide 2015; 44, 61-70.
[7] MBM Pinheiro, SV Rozini, AC Quirino-Teixeira, G Barbosa-Lima, JF Lopes, CQ Sacramento, FA Bozza, PT Bozza and ED Hottz. Dengue induces iNOS expression and nitric oxide synthesis in platelets through IL-1R. Frontiers in Immunology 2022; 13, 1029213.
[8] M Batty, MR Bennett and E Yu. The role of oxidative stress in atherosclerosis. Cells 2022; 11, 3843.
[9] S Yoon, GH Eom and G Kang. Nitrosative stress and human disease: Therapeutic potential of denitrosylation. International Journal of Molecular Sciences 2021; 22, 9794.
[10] FN Ningsih, T Okuyama, S To, Y Nishidono, T Okumura, K Tanaka, Y Ikeya and M Nishizawa. Comparative analysis of anti-inflammatory activity of the constituents of the rhizome of Cnidium officinale using rat hepatocytes. Biological and Pharmaceutical Bulletin 2020; 43(12), 1867-1875.
[11] A Nisar, S Jagtap, S Vyavahare, M Deshpande, A Harsulkar, P Ranjekar and O Prakash. Phytochemicals in the treatment of inflammation-associated diseases: The journey from preclinical trials to clinical practice. Frontiers in Pharmacology 2023; 14, 1177050.
[12] YH Gonfa, FB Tessema, A Bachheti, N Rai, MG Tadesse, AN Singab, KK Chaubey and RK Bachheti. Anti-inflammatory activity of phytochemicals from medicinal plants and their nanoparticles: A review. Current Research Biotechnology 2023; 6, 100152.
[13] FW Muema, Y Liu, Y Zhang, G Chen and M Guo. Flavonoids from Selaginella doederleinii Hieron and their antioxidant and antiproliferative activities. Antioxidants 2022; 11, 1189.
[14] H Yao, B Chen, Y Zhang, H Ou, Y Li, S Li, P Shi and X Lin. Analysis of the total biflavonoids extract from Selaginella doederleinii by HPLC-QTOF-MS and its in vitro and in vivo anticancer effects. Molecules 2017; 22, 325.
[15] M Ren, S Li, Q Gao, L Qiao, Q Cao, Z Yang, C Chen, Y Jiang, G Wang and S Fu. Advances in the anti-tumor activity of Biflavonoids in Selaginella. International Journal of Molecular Sciences 2023; 24, 7731.
[16] HN Pinanti, YI Christina, M Rifa’I, N Widodo and MS Djati. Aerial parts of Selaginella doederleinii Hieron as an anticancer agent against Luminal A breast cancer (T47D) cell line. Tropical Journal of Natural Product Research 2024; 8(6), 7452-7458.
[17] S Li, X Wang, G Wang, P Shi, S Lin, D Xu, B Chen, A Liu, L Huang, X Lin and H Yao. Ethyl acetate extract of Selaginella doederleinii Hieron induces cell autophagic death and apoptosis in colorectal cancer via PI3K-Akt-mTOR and AMPKα-signaling pathways. Frontiers in Pharmacology 2020; 11, 565090.
[18] N Ohno, E Yoshigai, T Okuyama, Y Yamamoto, T Okumura, K Sato, Y Ikeya and M Nishizawa. Chlorogenic acid from the Japanese herbal medicine Kinginka (Flos Lonicerae japonicae) suppresses the expression of inducible nitric oxide synthase in rat hepatocytes. HOAJ Biology 2012; 1(2), 1-10.
[19] T Kanemaki, H Kitade, Y Hiramatsu, Y Kamiyama and T Okumura. Stimulation of glycogen degradation by prostaglandin E2 in primary cultured rat hepatocytes. Prostaglandins 1993; 45(5), 459-474.
[20] E Yoshigai, T Machida, T Okuyama, M Mori, H Murase, R Yamanishi, T Okumura, Y Ikeya, H Nishino and M Nishizawa. Citrus nobiletin suppresses inducible nitric oxide synthase gene expression in interleukin-1β-treated hepatocytes. Biochemical and Biophysical Research Communications 2013; 439(1), 54-59.
[21] LC Green, DA Wagner, J Glogowski, PL Skipper, JS Wishnok and SR Tannenbaum. Analysis of nitrate, nitrite, and [15N] nitrate in biological fluids. Analytical Biochemistry 1982; 126(1), 131-138.
[22] MH Widyananda, DR Dwijayanti, A Fujii, K Minamisaka, Y Nishidono, M Nishizawa and N Widodo. Anti-obesity properties of Boesenbergia rotunda rhizome extract: Regulation of inflammation, lipid metabolism, and insulin signaling in ob/ob mice. Molecules 2025; 30, 501.
[23] G Yu, LG Wang, Y Han, QY He. ClusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 2012; 16(5), 284-287.
[24] SX Ge, D Jung and R Yao. ShinyGO: A graphical gene-set enrichment tool for animals and plants. Bioinformatics 2020; 36(8), 2628-2629.
[25] H Wickham. ggplot2: Elegant graphics for data analysis, second edition. Springer-Verlag, New York, 2016.
[26] P Shannon, A. Markiel, O Ozier, NS Baliga, JT Wang, D Ramage, N Amin, B Schwikowski and T Ideker. Cytoscape: A software Environment for integrated models of biomolecular interaction networks. Genome Research 2003; 13(11), 2498-2504.
[27] O Trott and AJ Olson. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. Journal of Computational Chemistry 2010; 31(2), 455-461.
[28] NM O’Boyle, M Banck, CA James, C Morley, T Vandermeersch and GR Hutchison. Open Babel: An open chemical toolbox. Journal of Cheminformatics 2011; 3, 33.
[29] S Dallakyan and AJ Olson. Small-molecule library screening by docking with PyRx. Methods in Molecular Biology 2015; 1263, 243-250.
[30] SA Halim, M Jawad, M Ilyas, Z Mir, AA Mirza and T Husnain. In silico identification of novel IL-1β inhibitors to target protein-protein interfaces. Computational Biology and Chemistry 2015; 58, 158-166.
[31] M Mic, A Pîrnău, CG Floare and M Bogdan. Study of the binding affinity between imatinib and α-1 glycoprotein using nuclear spin relaxation and isothermal titration calorimetry. International Journal of Biological Macromolecules 2020; 147, 326-332.
[32] AC Gonçalves, AR Costa, JD Flores-Félix, A Falcão, G Alves and LR Silva. Anti-inflammatory and antiproliferative properties of sweet cherry phenolic-rich extracts. Molecules 2022; 27(1), 268.
[33] T Zhang, C Ma, Z Zhang, H Zhang and H Hu. NF-κB signaling in inflammation and cancer. MedComm 2021; 2(4), 618-653.
[34] N Ahmad, MY Ansari, S Bano and TM Haqqi. Imperatorin suppresses IL-1β-induced iNOS expression via inhibiting ERK-MAPK/AP1 signaling in primary human OA chondrocytes. International Immunopharmacology 2020; 85, 106612.
[35] ML Hassan, FE Ali and AGS Shalkami. Role of TLR-4/IL-6/TNF-α, COX-II and eNOS/iNOS pathways in the impact of carvedilol against hepatic ischemia reperfusion injury. Human & Experimental Toxicology 2021; 40(8), 1362-1373.
[36] S Hu, Q Pi, M Luo, Z Cheng, X Liang, S Luo and Y Xia. Contribution of the NLRP3/IL-1β axis to impaired vasodilation in sepsis through facilitation of eNOS proteolysis and the protective role of melatonin. International Immunopharmacology 2021; 93, 107388.
[37] JA Nord, SL Wynia-Smith, AL Gehant, RA Jones Lipinski, A Naatz, I Rioja, RK Prinjha, JA Corbett and BC Smith. N-terminal BET bromodomain inhibitors disrupt a BRD4-p65 interaction and reduce inducible nitric oxide synthase transcription in pancreatic β-cells. Frontiers in Endocrinology 2022; 13, 923925.
[38] Q Lu, J Diao, Y Wang, J Feng, F Zeng, Y Yang, Y Kuang, N Zhao and Y Wang. 3D printed pore morphology mediates bone marrow stem cell behaviors via RhoA/ROCK2 signaling pathway for accelerating bone regeneration. Bioactive Materials 2023; 26, 413-424.
[39] AR Abubakar and M Haque. Preparation of medicinal plants: Basic extraction and fractionation procedures for experimental purposes. Journal of Pharmacy and Bioallied Sciences 2020; 12(1), 1-10.
[40] FA Ashu, J Na-Iya, BEN Wamba, J Kamga, P Nayim, B Ngameni, VP Beng, BT Ngadjui and V Kuete. Antistaphylococcal activity of extracts, fractions, and compounds of Acacia polyacantha Wild (Fabaceae). Evidence-Based Complementary and Alternative Medicine 2020; 2020, 2654247.
[41] PR Sapkal, AU Tatiya, SD Firke, VK Redasani, SS Gurav, M Ayyanar, PG Jamkhande, SJ Surana, RE Mutha and MG Kalaskar. Phytochemical profile, antioxidant, cytotoxic, and anti-inflammatory activities of stem bark extract and fractions of Ailanthus excelsa Roxb.: In vitro, in vivo, and in silico approaches. Heliyon 2023; 9(5), e15952.
[42] SM Andrabi, NS Sharma, A Karan, SMS Shahriar, B Cordon, B Ma and J Xie. Nitric oxide: Physiological functions, delivery, and biomedical applications. Advanced Science 2023; 10, 2303259.
[43] Y Iwakiri and MY Kim. Nitric oxide in liver diseases. Trends in Pharmacological Sciences 2015; 36(8), 524-536.
[44] F Beheshti, M Hosseini, M Taheri Sarvtin, A Kamali and A Anaeigoudari. Protective effect of aminoguanidine against lipopolysaccharide-induced hepatotoxicity and liver dysfunction in rat. Drug and Chemical Toxicology 2021; 44(2), 215-221.
[45] Y Sui, S Li, P Shi, Y Wu, Y Li, W Chen, L Huang, H Yao and X Lin. Ethyl acetate extract from Selaginella doederleinii Hieron inhibits the growth of human lung cancer cells A549 via caspase-dependent apoptosis pathway. Journal of Ethnopharmacology 2016; 190, 261-271.
[46] G Wang, S Yao, XX Zhang and H Song. Rapid screening and structural characterization of antioxidants from the extract of Selaginella doederleinii Hieron with DPPH-UPLC-Q-TOF/MS method. International Journal of Analytical Chemistry 2015; 2015, 849769.
[47] J Zhou, RC Nie, YX Yin, XX Cai, D Xie and MY Cai. Protective effect of natural antioxidants on reducing cisplatin-induced nephrotoxicity. Disease Markers 2022; 2022, 1612348.
[48] M Pereira and RT Gazzinelli. Regulation of innate immune signaling by IRAK proteins. Frontiers in Immunology 2023; 14, 1133354.
[49] J Totzke, SA Scarneo, KW Yang and TAJ Haystead. TAK1: A potent tumour necrosis factor inhibitor for the treatment of inflammatory diseases. Open Biology 2020; 10, 200099.
[50] Z Wang and L Yang. Chinese herbal medicine: Fighting SARS-CoV-2 infection on all fronts. Journal of Ethnopharmacology 2021; 270, 113869.
[51] JD Webster and D Vucic. The balance of TNF-mediated pathways regulates inflammatory cell death signaling in healthy and diseased tissues. Frontiers in Cell and Developmental Biology 2020; 8, 365.
[52] D Iacobazzi, P Convertini, S Todisco, A Santarsiero, V Iacobazzi and V Infantino. New insights into NF-κB signaling in innate immunity: Focus on immunometabolic crosstalks. Biology 2023; 12, 776.
[53] Q Guo, Y Jin, X Chen, X Ye, X Shen, M Lin, C Zeng, T Zhou and J Zhang. NF-κB in biology and targeted therapy: New insights and translational implications. Signal Transduction Target Therapy 2024; 9, 53.
[54] S Wang, D Wan, W Liu, X Kang, X Zhou, F Sefidkon, MMZ Hosseini, T Zhang, X Pan and X Yang. A biflavonoid-rich extract from Selaginella doederleinii Hieron against throat carcinoma via Akt/Bad and IKKβ/NF-κB/COX-2 pathways. Pharmaceuticals 2022; 15, 1505.
[55] SN Poetzsch, M Poetzsch, T Kraemer and AE Steuer. Toward a cannabis terroir: Untargeted metabolomic profiling of authentic samples using gas chromatography–high-resolution mass spectrometry (GC-HRMS) and liquid chromatography–high-resolution tandem mass spectrometry (LC-HRMS/MS). Drug Testing and Analysis 2025; 17, 2086-2095.
[56] D Ballesteros-Vivas, G Alvarez-Rivera, E Ibáñez, F Parada-Alfonso and A Cifuentes. Integrated strategy for the extraction and profiling of bioactive metabolites from Passiflora mollissima seeds combining pressurized-liquid extraction and gas/liquid chromatography–high-resolution mass spectrometry. Journal of Chromatography A 2019; 1595, 144-157.
[57] C Nichols, J Ng, A Keshu, G Kelly, MR Conte, MS Marber, F Fraternali and GFD Nicola. Mining the PDB for tractable cases where x-ray crystallography combined with fragment screens can be used to systematically design protein-protein inhibitors: Two test cases illustrated by IL1β-IL1R and p38α-TAB1 complexes. Journal of Medicinal Chemistry 2020; 63(14), 7559-7568.
[58] X Wu, Q Yu, Y Hou, X Zhang, SS Ocholi, L Wang, Z Yan, J Li and L Han. Emodin-8-O-β-D-glucopyranoside alleviates cholestasis by maintaining intestinal homeostasis and regulating lipids and bile acids metabolism in mice. Journal of Pharmaceutical and Biomedical Analysis 2025; 258, 116734.
[59] SM Ulfa, S Shirako, M Sato, DR Dwijayanti, T Okuyama, S Horie, J Watanabe, Y Ikeya and M Nishizawa. Anti-inflammatory effects of anthraquinones of Polygonum multiflorum roots. Bioactive Compounds in Health and Disease 2022; 5(6), 136-148.
[60] P Wang, Q He and J Zhu. Emodin-8-O-glucuronic acid, from the traditional Chinese medicine Qinghuobaiduyin, affects the secretion of inflammatory cytokines in LPS-stimulated RAW 264.7 cells via HSP70. Molecular Medicine Reports 2016; 14(3), 2368-2372.