
Trends
Sci.
2026;
23(7):
12614
Characterization and Biotechnological Potential of Halophilic Fungi Producing Salt-Tolerant Amylase and Protease from Marine Environments
Ninadia Jitprasitporn and Lakkhana Kanhayuwa Wingfield*
Division of Biological Science, Faculty of Science, Prince of Songkla University, Songkhla 90110, Thailand
(*Corresponding author’s e-mail: [email protected])
Received: 13 November 2025, Revised: 18 December 2025, Accepted: 28 December 2025, Published: 1 March 2026
Abstract
This study aimed to characterize halophilic and halotolerant fungi from marine environments and evaluate their potential as sources of salt-tolerant hydrolases for biotechnological applications in the food and detergent industries. Seven fungal isolates were identified through morphological and molecular characterization of the ITS region and phylogenetic analysis using the Maximum Likelihood method. Salt tolerance was evaluated on PDA supplemented with 0% - 20% NaCl, while amylase and protease activities were assessed by plate assays and quantified spectrophotometrically during 7 - 21 days of cultivation. Crude enzyme extracts were further examined for orange juice clarification and stain-removal efficiency on cotton fabrics, both individually and in combination with sodium dodecyl sulfate (SDS). All isolates belonged to the genera Aspergillus and Penicillium and exhibited facultative halophilism, with optimal growth at 5% - 10% NaCl. A. subalbidus NM-16 and NM-17.1 showed the highest amylase activities (3.16 ± 0.43 and 2.63 ± 0.24 EI), whereas protease production occurred exclusively in P. citrinum isolates. Among them, P. citrinum I22M1 exhibited maximal enzyme activities at day 21 (amylase = 0.93 ± 0.01 U/mL; protease = 1.58 ± 0.01 U/mL). Crude enzyme preparations achieved 32.53 ± 2.01% juice clarification and up to 95.67 ± 8.85% and 88.99 ± 1.21% removal of chocolate and blood stains, respectively. The synergistic combination of enzyme and SDS completely restored fabric whiteness. These findings demonstrate that marine-derived Aspergillus and Penicillium produce stable, halotolerant amylase and protease enzymes capable of functioning under saline conditions, highlighting their potential as eco-friendly biocatalysts for sustainable industrial processes.
Keywords: Halophilic fungi, Amylase, Protease, Salt tolerance, Detergent enzyme, Biocatalyst
Introduction
The marine environment is a highly saline ecosystem inhabited by diverse microbial communities, including bacteria, fungi, algae, plankton, and viruses. These microorganisms play essential ecological roles through their involvement in critical biogeochemical processes [1]. Among them, fungi have gained increasing scientific attention, particularly those thriving in extreme conditions. These organisms serve as valuable models for studying functional evolution and offer promising biotechnological applications [2]. Marine fungi are exposed to harsh environmental pressures such as fluctuating temperatures, high salinity, nutrient limitations, and intense microbial competition. These challenges drive the development of unique genetic and metabolic traits that distinguish marine fungi from their terrestrial counterparts [3]. High salinity, in particular, imposes osmotic and ionic stress, which marine fungi counteract via a conserved high-osmolarity-glycerol (HOG) signaling pathway to maintain cellular homeostasis [4].
The diversity of marine fungi is vast, with estimates suggesting over 10,000 species [5]. Most belong to the phyla Ascomycota and Basidiomycota, which together account for more than 90% of known marine fungal taxa [6]. These fungi are generally categorized as either obligate or facultative. Obligate marine fungi exclusively grow and reproduce in marine or estuarine environments, while facultative marine fungi originate from terrestrial or freshwater habitats but can adapt to marine conditions [7]. Fungi inhabiting saline environments are further classified as halophilic or halotolerant. Halophilic fungi require high concentrations of salt, typically sodium chloride (NaCl), for optimal growth and often fail to grow in salt-free conditions. Some can tolerate NaCl concentrations exceeding 1.5 M, with certain species growing in saturated salt solutions (up to 5.2 M NaCl) [8,9]. Wallemia ichthyophaga, for instance, is an extremely halophilic basidiomycete capable of growing in high-salt environments [10]. In contrast, halotolerant fungi do not require salt for growth but can survive and proliferate under saline conditions. These fungi exhibit broad osmotolerance by accumulating compatible solutes, such as glycerol, and regulating ion transporters to maintain cellular function under osmotic stress [11,12]. Genera such as Aspergillus and Penicillium are commonly halotolerant and frequently isolated from saline or hypersaline habitats [13].
Marine fungi are prolific producers of structurally diverse secondary metabolites, including polyketides, alkaloids, terpenes, peptides, and hybrid compounds, that often display potent antibacterial, antiviral, and anticancer activities [14]. Additionally, they are increasingly recognized for producing industrially important enzymes, particularly amylases and proteases, which exhibit favorable traits such as thermostability, salinity and pH tolerance, and activity in unconventional reaction environments [15,16]. These features make marine fungal enzymes valuable to various industries, including food, pharmaceuticals, textiles, leather, biofuels, detergents, and waste management [15]. Several marine-derived fungal genera, including Aspergillus, Penicillium, Rhizopus, Mucor, and Humicola, have been identified as producers of enzymes with exceptional activity and stability [16]. To further explore this potential, we screened fungal strains isolated from diverse marine environments for amylase and protease activity. This study aimed to enhance the understanding of marine halophilic fungi by evaluating their enzymatic capabilities. We examined their halophilic growth characteristics, conducted morphological and molecular identification, and characterized crude extracellular enzymes for protease and amylase activities, with a focus on assessing their potential applications in the detergent and food industries.
Materials
and methods
Fungal strains and maintenance
Fungal strains were obtained from the Mycology Laboratory, Division of Biological Science, Prince of Songkla University, and maintained on potato dextrose agar (PDA) slants. Strains NM16 and NM17.1 were isolated from marine salterns, while strains F11M6, F12M4.1, F12M5, F12M2, and I22M1 were isolated from marine sea cucumbers (Holothuria scabra). All strains were subsequently cultured on PDA supplemented with 5% (w/v) NaCl and incubated at 28 °C for 5 - 7 days for further use.
Identification of fungal isolates
For morphological Identification, fungal isolates were identified to the genus and species levels based on macroscopic and microscopic characteristics, following the taxonomic keys and descriptions provided by Samson et al. [17]; Dugan [18]; Ellis et al. [19]. Each isolate was cultured by transferring a mycelial plug onto a PDA plate and incubated at 28 °C for 7 days. After incubation, colony morphology was recorded. Microscopic examination of vegetative and reproductive structures was performed using a compound microscope (Olympus CX23, Japan). To observe detailed structures, the slide culture technique was employed. Molecular analysis was carried out using a genetic marker, the internal transcribed spacer (ITS) region. Genomic DNA was extracted from fresh mycelia using the DNeasy® Plant Mini Kit (QIAGEN, UK) following the manufacturer’s protocol. PCR amplification was performed using a primer set and condition described by Wingfield et al. [20]. An ITS primer set had the following sequences: ITS5 (5’-GGAAGTAAAAGTCGTAACAAGG-3’) and ITS4N (5’-TCCTCCGCTTATTGATATGC-3’). Each 50 µL reaction mixture contained 5 µL of 10x Taq buffer, 5 µL of dNTP mix (2mM each), 2 µL of each primer at 1.0 µM, 0.5 µL of Taq DNA polymerase (1.25U) (New England Biolabs, USA) and 1 µg of genomic DNA. The PCR reaction was performed using a DNA Engine DYAD ALD 1244 thermocycler (MJ Research Inc., USA) with the following cycles: Initial denaturation at 94 °C for 5 min, 35 cycles of denaturation at 94 °C for 1 min, annealing at 55 °C for 1 min, and extension at 72 °C for 2 min, followed by a final extension at 72 °C for 10 min. Amplified products were verified by agarose gel electrophoresis and subsequently purified using the MinElute® Gel Extraction Kit (QIAGEN). DNA sequencing was conducted on a 3730xl DNA Analyzer (Life Technologies, Carlsbad, CA, USA) by Macrogen (Seoul, Korea).
The ITS sequences were queried using BLASTn against the NCBI GenBank database to identify the closest matches. Phylogenetic analysis was performed in MEGA version 11 [21], using MUSCLE for multiple sequence alignment with manual refinement. A maximum-likelihood tree was constructed, and branch support was evaluated via 1,000 bootstrap replicates. Newly generated sequences were submitted to GenBank (Table S1).
Determination of salt tolerance and optimal NaCl concentration for fungal growth
Fungal isolates were cultured on PDA supplemented with varying concentrations of NaCl at 0%, 5%, 10%, 15%, and 20% (w/v) to assess their salt tolerance and determine the optimal salinity for growth. A 5 µL aliquot of spore suspension (1×105 spores/mL) was inoculated at the center of each PDA plate. Plates were incubated at 28 °C for 21 days. Fungal growth was evaluated by measuring the diameter of the colony, and morphological characteristics of each isolate were observed daily. The experiment was conducted in triplicate, and results were expressed as mean ± standard deviation (n = 3).
Enzyme assays
The ability of fungal isolates to produce extracellular enzymes was evaluated using plate-based assays. Amylase production was assessed on nutrient agar supplemented with 2 g/L of soluble starch. Each isolate was inoculated onto the medium and incubated at 28 °C for 2 to 5 days. After incubation, the plates were flooded with an iodine solution. The presence of a clear halo around the colony indicated amylase activity [22]. To enhance enzyme visualization, the cultures were subsequently incubated at 50 °C for 16 h, followed by flooding with 5 mL of 1% Congo red solution. Plates were rinsed with distilled water, and the formation of a hydrolysis zone further confirmed amylase activity [23]. Protease production was determined using casein agar medium prepared with 1% (w/v) skim milk, 0.01% (v/v) Triton X-100 (Sigma) and 1.5% (w/v) agar. Proteinase K (Qiagen) and sterile distilled water were used as positive and negative controls, respectively. After inoculation, the plates were incubated at 28 °C for 2 to 5 days, and proteolytic activity was evidenced by a clear zone surrounding the fungal colonies [22]. For both enzyme types, activity was quantified using the enzymatic index (EI), calculated as the ratio of the diameter of the clear zone (R) to the diameter of the fungal colony (r). Higher EI values correspond to greater extracellular enzymatic activity.
Preparation of extracellular crude enzymes
Each fungal isolate was cultured on PDA supplemented with 5% (w/v) NaCl and incubated at 28 °C for 7 days. Following incubation, 4 agar plugs were transferred into 50 mL of liquid culture medium. For amylase activity, plugs were inoculated into a starch culture broth containing (per L): 10 g soluble starch, 3 g peptone, 8 g CaCO₃, 6.6 g (NH₄)₂SO₄, 3.5 g KH₂PO₄, 0.15 g FeSO₄·7H₂O, 0.1 g MgSO₄·7H₂O, and 5% (w/v) NaCl in a potato dextrose broth (PDB) base. For protease activity, plugs were transferred into skim milk broth composed of 0.5% (w/v) skim milk, 0.01% (v/v) Triton X-100, and 5% (w/v) NaCl in PDB. All cultures were incubated at 28 °C under agitation at 100 rpm for 14 days. After incubation, the culture broths were filtered through Whatman filter paper to remove mycelial debris. The filtrates were then centrifuged at 5,000 rpm for 10 min, and the supernatants were further sterilized by syringe filtration using a 0.45 µm membrane filter. All crude enzyme preparations were stored at −20 °C until further use.
Enzyme activity assays
Amylase activity was measured using the dinitrosalicylic acid (DNS) method with minor modifications [24]. The reaction mixture contained 0.1 mL of 1% (w/v) soluble starch and 0.05 mL of crude enzyme, incubated at 37 °C for 30 min. A reaction blank was prepared by replacing the enzyme with 0.05 mL of distilled water. The reaction was terminated by adding 1.0 mL of DNS reagent, followed by heating at 90 °C for 10 min and cooling to room temperature. Absorbance was measured at 540 nm. A standard curve was prepared using glucose as the reference. One unit of amylase activity (U/mL) was defined as the amount of enzyme that releases 1 μmol of reducing sugar per minute under the assay conditions. Proteolytic activity was measured using azocasein as a substrate, following modified protocols [25,26]. A 0.05 mL aliquot of crude enzyme was incubated with 0.45 mL of 3 mg/mL azocasein in 20 mM phosphate buffer (pH 6.0) at 37 °C for 1 h. The reaction was stopped by adding 0.5 mL of 5% (w/v) trichloroacetic acid (TCA), and the mixture was centrifuged at 9,700×g for 10 min. A 0.1 mL aliquot of the supernatant was mixed with 0.1 mL of 0.5 N NaOH in a 96-well microplate, and absorbance was measured at 440 nm using a microplate reader. A reaction blank was prepared using TCA-inactivated enzyme. One unit of protease activity (U/mL) was defined as the amount of enzyme that hydrolyzes azocasein to release peptides per minute under assay conditions. The influence of cultivation time on enzyme production was assessed by harvesting crude enzymes from fungal cultures grown for 7, 14, and 21 days. Enzyme activities were then measured using the respective standard assays described above.
Estimation of soluble protein
Total protein concentration of the crude enzymes was determined using the Bradford method [27], with bovine serum albumin (BSA) as the standard. Briefly, 20 µL of crude enzyme sample was mixed with 1.0 mL of Bradford reagent (Coomassie Brilliant Blue G-250) and incubated at room temperature for 5 min. Absorbance was measured at 595 nm using a UV-Vis spectrophotometer. Protein concentrations were calculated from a standard curve prepared using serial dilutions of BSA (0 - 100 µg/mL). All measurements were performed in triplicate, and results were expressed as micrograms of protein per milliliter of enzyme solution (µg/mL).
Assessment of industrial applications of crude enzymes
Juice clarification assay
Fresh orange juice (Citrus sinensis) was treated with crude fungal amylase at a 1:1 (v/v) juice-to-enzyme ratio. Controls included (i) distilled water in place of enzyme, and (ii) 1 mL of commercial α-amylase (100 U/mL). Sodium benzoate (0.001 g) was added to all treatments to inhibit microbial growth. Samples were incubated at 25 °C for 24 h under static conditions. After incubation, samples were centrifuged at 5,000×g for 10 min to remove suspended solids. The supernatant was collected, and visual clarity was assessed by measuring the OD at 600 nm. Unclarified raw orange juice was used as a control, and its turbidity was defined as 100% [28]. The degree of clarification was calculated as a percentage reduction in OD relative to the control.
Stain removal assay
Clarification of blood and chocolate samples
Fresh sheep blood and melted chocolate were treated with crude fungal enzyme preparations at a 1:1 (v/v) sample-to-enzyme ratio. Controls included (i) distilled water in place of enzyme, and (ii) 1 mL of either commercial α-amylase or protease (100 U/mL). All samples were incubated at 37 °C for 24 h under static conditions. Following incubation, the mixtures were centrifuged at 5,000 ×g for 10 min to remove suspended solids. The resulting supernatant was collected, and visual clarity was assessed by measuring the optical density at 660 nm. The degree of clarification was calculated as the percentage reduction in OD relative to the control.
Stain removal from cotton fabrics
Cotton fabric squares (4×4 cm2) were stained with either fresh sheep blood or melted chocolate and left to air-dry for 30 min at room temperature. The stained fabrics were then subjected to enzyme treatment under the following experimental conditions: (i) distilled water (10 mL) + stained cloth, (ii) crude enzyme solution (10 mL) + stained cloth, (iii) distilled water (10 mL) + sodium dodecyl sulfate (SDS; 0.1 mL of 7 mg/mL) + stained cloth and, (iv) distilled water (10 mL) + SDS (0.1 mL of 7 mg/mL) + crude enzyme solution (10 mL) + stained cloth. All flasks were incubated at 37 °C for 30 min under static conditions. After incubation, the cloth pieces were removed, rinsed thoroughly with distilled water, and allowed to air-dry. The effectiveness of stain removal was evaluated visually. Untreated stained cloth samples served as negative controls. The methodology was adapted from a previously reported protocol [29].
Statistical analysis
All experiments were performed in triplicate across 3 independent trials. Data is presented as a mean standard deviation (SD). Statistical significance was assessed using either Student’s t-test or 1-way analysis of variance (ANOVA), followed by Tukey’s post hoc multiple comparison test to identify differences among group means. A p-value of less than 0.05 (p < 0.05) was considered statistically significant. All statistical analyses were conducted using GraphPad Prism version 8.0 (GraphPad Software, La Jolla, CA, USA).
Results and discussion
Identification of halophilic fungi
Seven halophilic fungal isolates were identified based on morphological and molecular characterization using the ITS region. Morphological observations (Figure 1(A)) revealed colony morphology and microscopic features consistent with members of the genera Aspergillus and Penicillium. Molecular identification via BLASTn analysis of ITS sequences showed ≥ 98.95% similarity to known fungal species in the NCBI GenBank database (Table S1). The phylogenetic tree constructed using the Maximum Likelihood method with the Tamura-Nei model (1,000 bootstrap replicates; Figure 1(B)) clustered isolates NM-16 and NM-17.1 with Aspergillus subalbidus (99.83% and 99.82% identity, respectively), isolate F11M6 with A. terreus (99.65%), and 4 isolates (F12M4.1, F12M5, F12M2, I22M1) with Penicillium citrinum (98.95% - 99.65%).
These findings confirm that the seven isolates belong to Aspergillus and Penicillium, which are widely recognized as dominant fungal taxa in saline and hypersaline ecosystems. Consistent with previous metagenomic and culture-dependent studies [30,31], both genera frequently occur as halotolerant filamentous fungi in marine and coastal environments. The high ITS sequence similarity to A. subalbidus, A. terreus, and P. citrinum supports their taxonomic assignment and aligns with earlier reports of Aspergillus and Penicillium species isolated from saline soils, sediments, and solar salterns [32]. The close clustering of A. subalbidus NM-16 and NM-17.1 with reference sequences further validates their phylogenetic placement. Collectively, these results highlight the adaptive capability of Aspergillus and Penicillium as facultative halophiles able to withstand fluctuating salinity levels [33]. The established morphological and molecular identities provide a robust taxonomic foundation for subsequent biochemical analyses and application-oriented studies of their halotolerant enzymes, addressing a key research gap in the underexplored domain of marine fungi as potential producers of industrially relevant amylases and proteases [34].
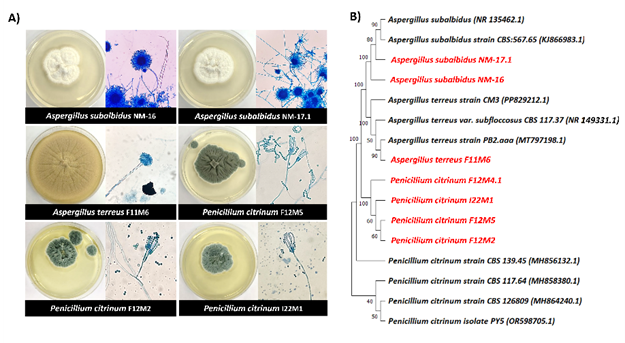
Figure 1 Identification of halophilic fungi. (A) Macroscopic and microscopic morphology of selected halophilic fungal isolates (magnification×40). All isolates were cultured on PDA without NaCl supplementation and incubated at 28 °C. (B) Phylogenetic tree based on ITS gene sequences of halophilic fungi and related species. The tree was constructed using the Maximum Likelihood method with the Tamura-Nei model. Bootstrap values (1,000 replicates) are shown at the nodes.
Halotolerance and halophilic characteristics of fungi
To evaluate the salt tolerance and halophilic behavior of the fungal isolates, colony growth was assessed on PDA plates supplemented with increasing concentrations of NaCl (0%, 5%, 10%, 15% and 20% w/v). All isolates exhibited robust growth across NaCl concentrations ranging from 0% to 20%, confirming their halotolerant nature (Figure 2). Growth was sustained even at the highest salt concentration tested (20% NaCl), although with reduced colony diameters, indicating tolerance rather than preference for extreme salinity. Differences in growth rates across NaCl concentrations were evident, with most isolates showing significantly enhanced growth at 5% and 10% NaCl compared to 0%. For example, Aspergillus subalbidus NM-16 and A. subalbidus NM-17.1 displayed maximal colony diameters at 5% - 10% NaCl, significantly higher than at 0%, 15%, or 20% NaCl. Similarly, Penicillium spp. exhibited optimal growth at 5% NaCl but showed reduced colony expansion at higher salinities.
The ability of all isolates to grow in the absence of NaCl, coupled with enhanced growth at moderate salinities, suggests that these fungi are facultative halophiles rather than obligate halophiles. Extended incubation up to 21 days (Figure 3) confirmed these patterns, with colony diameters peaking at 5% - 10% NaCl and plateauing or declining beyond 15%. Statistical analysis further supported these observations, revealing significant differences (p < 0.05) in colony diameters across salinity gradients. This growth pattern agrees with recent reports showing that halotolerant Aspergillus and Penicillium species often exhibit improved mycelial expansion under moderate salinity [30]. Such enhancement may be linked to osmotic adjustment mechanisms, including the intracellular accumulation of compatible solutes (e.g., glycerol, trehalose) and the upregulation of stress-response genes that maintain cellular homeostasis under saline conditions [33]. At higher NaCl concentrations (≥ 15%), reduced growth likely reflects the diversion of metabolic energy toward osmoregulatory processes rather than active biomass production. These findings demonstrate that marine-derived fungi possess remarkable ecological flexibility and adaptive responses to osmotic stress. Their ability to maintain metabolic activity and enzyme production across a wide salinity range underscores their potential for industrial applications in saline environments, such as in detergent and food biotechnology where salt-tolerant enzymes are valuable.
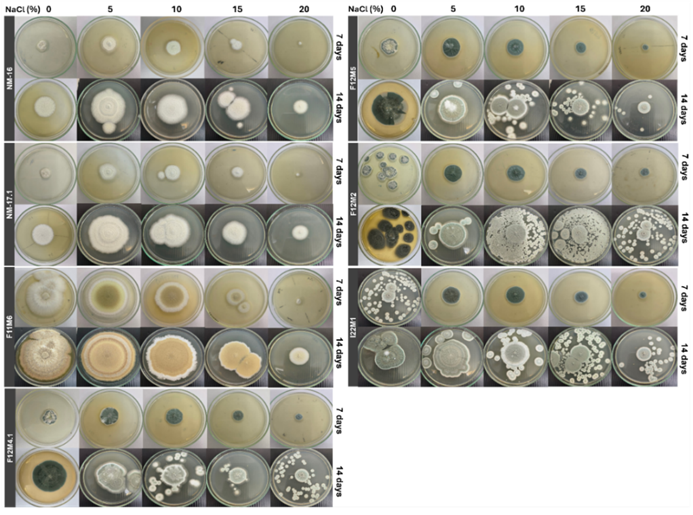

Figure 2 Halotolerance and halophilic growth of selected fungal isolates. Colony diameters were measured after 14 days of incubation at 28 °C on PDA supplemented with varying NaCl concentrations (0% - 20% w/v). Growth performance under saline conditions was used to assess halotolerance and halophilic characteristics.
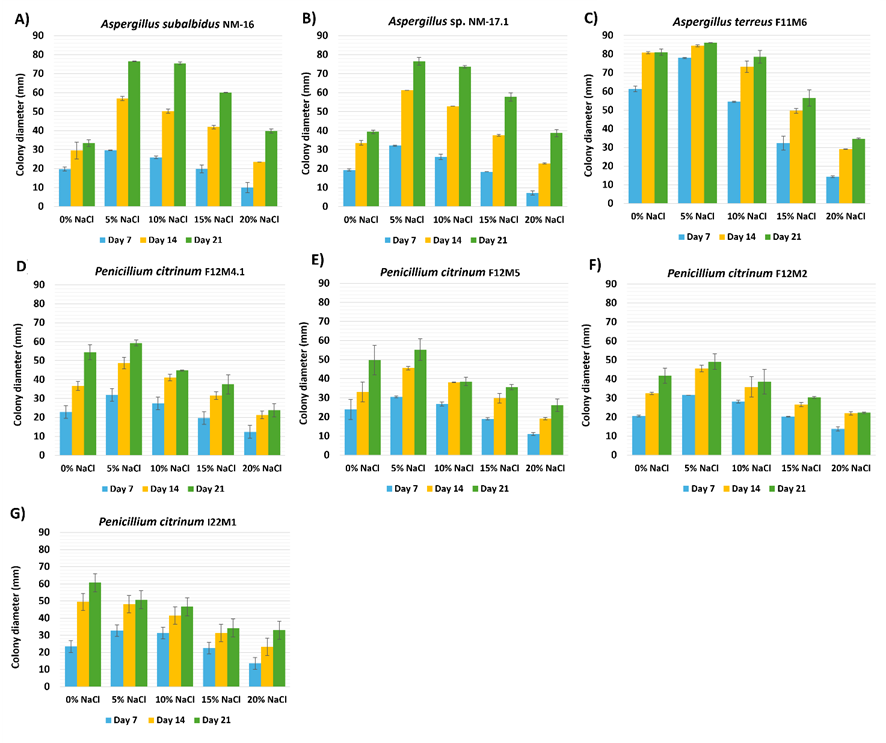
Figure 3 Effect of NaCl concentration on fungal growth (Day 21). Mean colony diameters (mm) of halophilic fungal isolates grown under different NaCl concentrations (0% - 20%) after 21 days of incubation. Error bars represent standard deviations from simulated replicates (n = 3).
Screening for enzymatic activity
All fungal isolates demonstrated extracellular hydrolase production, as confirmed by substrate-specific agar assays for amylase and protease activities (Figure 4). Clear hydrolytic zones indicated enzymatic activity, quantified by the enzymatic index (EI; Table 1). Amylase activity was detected in all isolates, with Aspergillus subalbidus NM-16 showing the highest EI (3.16 ± 0.43), followed by NM-17.1 (2.63 ± 0.24). Both isolates exhibited significantly greater activity than the others (p < 0.05), while A. terreus F11M6 displayed the lowest amylase activity (1.14 ± 0.02). Penicillium citrinum isolates showed moderate amylase production (EI 1.75 - 2.16). In contrast, protease activity occurred exclusively in P. citrinum isolates, forming clear halos on skim-milk agar with EI values of 1.55 - 1.91, with no significant variation among strains (p > 0.05). No protease activity was observed in Aspergillus isolates. Based on the classification threshold (EI ≥ 2), A. subalbidus NM-16, A. subalbidus NM-17.1, P. citrinum F12M2, and P. citrinum I22M1 were identified as promising enzyme producers. Notably, only the Penicillium isolates exhibited polyenzymatic potential, producing both amylase and protease enzymes.
These findings indicate that the tested marine-derived fungi possess broad catabolic versatility. The high α-amylase activity of A. subalbidus NM-16 and NM-17.1 supports the recognized glycosidase potential of marine Aspergillus species [33]. Meanwhile, protease production exclusively by P. citrinum aligns with reports that Penicillium species often express proteolytic enzymes in saline and alkaline conditions [30]. The dual enzyme production by P. citrinum F12M2 and I22M1 underscores their value for multifunctional industrial applications such as detergent formulation and food processing. Tukey’s HSD analysis confirmed significant interspecific variation (p < 0.05) in enzyme indices, highlighting A. subalbidus NM-16 and P. citrinum I22M1 as potential candidates for large-scale enzyme production. Overall, the secretion of multiple salt-tolerant hydrolases by these halophilic fungi reinforces their relevance as sources of extremozymes suitable for harsh industrial environments [34].
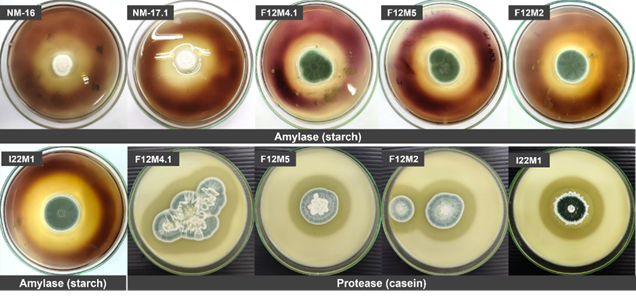
Figure 4 Enzymatic activity assay of halophilic fungi. Representative images show extracellular amylase and protease production by selected isolates on agar plates. Clear zones surrounding colonies indicate hydrolytic activity.
Table 1 Summary of enzymatic activity of halophilic fungi.
Fungi |
Isolate code |
Enzyme activity (EA)/Enzymatic index (EI) |
|||
Amylase |
Protease |
||||
EA |
EI |
EA |
EI |
||
Aspergillus subalbidus |
NM-16 |
++ |
3.16 ± 0.43a |
- |
|
Aspergillus subalbidus |
NM-17.1 |
++ |
2.63 ± 0.24a |
- |
- |
Aspergillus terreus |
F11M6 |
+ |
1.14 ± 0.02b |
- |
- |
Penicillium citrinum |
F12M4.1 |
+ |
1.75 ± 0.01b |
+ |
1.90 ± 0.22a |
Penicillium citrinum |
F12M5 |
+ |
1.99 ± 0.17b |
+ |
1.55 ± 0.00a |
Penicillium citrinum |
F12M2 |
+ |
2.13 ± 0.21b |
+ |
1.60 ± 0.40a |
Penicillium citrinum |
I22M1 |
+ |
2.16 ± 0.11b |
+ |
|
Amylase and protease production by fungal isolates was assessed based on enzyme activity (EA) in mm and enzymatic index (EI). “+” indicates low activity (0 - 3 mm), “++” moderate (3.1 - 6 mm), and “+++” high (> 6 mm). “–” indicates no enzymatic activity. Data are presented as mean ± SD (n = 3). EI with superscript letters indicate statistical groupings based on Tukey’s HSD test (p < 0.05). Isolates sharing the same letter are not significantly different.
Enzyme activity and protein production of halophilic fungi
The amylase and protease activities, along with corresponding protein content, of halophilic fungal strains cultivated under saline conditions for 7, 14, and 21 days are summarized in Table 2. Amylase production varied among strains and across cultivation periods. At 7 days, Aspergillus subalbidus NM-16 exhibited the highest amylase activity (1.39 ± 0.02 U/mL), followed by A. subalbidus NM-17.1 (0.89 ± 0.07 U/mL) and Penicillium citrinum I22M1 (0.83 ± 0.01 U/mL), whereas A. terreus F11M6 showed negligible activity (0.01 ± 0.00 U/mL). By day 14, enzyme activities generally declined, except in P. citrinum I22M1 (0.51 ± 0.02 U/mL) and F12M5 (0.38 ± 0.01 U/mL), which maintained relatively higher levels. After 21 days, P. citrinum I22M1 demonstrated the highest amylase activity (0.93 ± 0.01 U/mL), while other strains showed moderate activity (0.67 - 0.85 U/mL). Protein content across all amylase-producing strains remained stable (0.66 ± 0.04 to 0.91 ± 0.05 mg/mL), indicating consistent secretory output.
Protease activity was observed exclusively in P. citrinum isolates. At 7 days, P. citrinum I22M1 recorded the highest protease activity (1.14 ± 0.05 U/mL), followed by F12M5 (1.04 ± 0.06 U/mL). Over time, enzyme profiles revealed dynamic changes. While F12M4.1 maintained consistent activity (0.68 - 0.95 U/mL), both F12M5 and F12M2 showed sharp declines by day 21 (0.36 ± 0.00 and 0.15 ± 0.01 U/mL, respectively). In contrast, P. citrinum I22M1 exhibited a substantial increase at day 21 (1.58 ± 0.01 U/mL), indicating delayed or sustained enzyme induction under prolonged incubation. Protein content among protease producers remained relatively constant (0.71 - 0.91 mg/mL).
Temporal enzyme production patterns demonstrated clear strain- and time-dependent variability. The early amylase peak in A. subalbidus NM-16 suggests association with primary metabolism, while the delayed induction in P. citrinum I22M1, reaching maximal amylase and protease activities after 21 days, may correspond to secondary metabolic activation triggered by osmotic stress. Such delayed secretion kinetics align with reports on other extremophilic fungi, where salt stress induces complex transcriptional and post-translational regulatory mechanisms [34]. The relatively stable total protein levels across all strains and time points further imply that changes in enzyme yield stemmed from secretory regulation rather than biomass accumulation. These findings highlight the importance of optimizing fermentation parameters, particularly incubation period and salinity, to maximize enzyme productivity. The observed stability and sustained enzyme release of P. citrinum I22M1 make it a promising candidate for industrial applications requiring robust and long-lasting enzymatic performance [33].
Table 2 Amylase and protease activities (U/mL) and corresponding protein content (mg/mL) of different halophilic fungal strains cultivated for 7, 14, and 21 days.
Fungi |
Enzyme activity (U/mL) |
Protein content (mg/mL) |
||||
7 Days |
14 Days |
21 Days |
7 Days |
14 Days |
21 Days |
|
Amylase |
|
|
|
|
|
|
Aspergillus subalbidus NM-16 |
1.39 ± 0.02 |
0.31 ± 0.01 |
0.67 ± 0.03 |
0.86 ± 0.03 |
0.89 ± 0.06 |
0.73 ± 0.04 |
Aspergillus subalbidus NM-17.1 |
0.89 ± 0.07 |
0.27 ± 0.00 |
0.71 ± 0.05 |
0.85 ± 0.03 |
0.91 ± 0.05 |
0.74 ± 0.05 |
Aspergillus terreus F11M6 |
0.01 ± 0.00 |
0.33 ± 0.00 |
0.82 ± 0.01 |
0.88 ± 0.02 |
0.90 ± 0.06 |
0.73 ± 0.03 |
Penicillium citrinum F12M4.1 |
0.19 ± 0.04 |
0.32 ± 0.01 |
0.85 ± 0.06 |
0.85 ± 0.05 |
0.66 ± 0.04 |
0.73 ± 0.02 |
Penicillium citrinum F12M5 |
0.77 ± 0.03 |
0.38 ± 0.01 |
0.79 ± 0.03 |
0.82 ± 0.01 |
0.87 ± 0.03 |
0.69 ± 0.01 |
Penicillium citrinum F12M2 |
0.54 ± 0.04 |
0.30 ± 0.02 |
0.67 ± 0.02 |
0.81 ± 0.02 |
0.90 ± 0.04 |
0.71 ± 0.02 |
Penicillium citrinum I22M1 |
0.83 ± 0.01 |
0.51 ± 0.02 |
0.93 ± 0.01 |
0.81 ± 0.02 |
0.87 ± 0.01 |
0.68 ± 0.04 |
Protease |
|
|
|
|
|
|
Penicillium citrinum F12M4.1 |
0.74 ± 0.00 |
0.68 ± 0.02 |
0.95 ± 0.05 |
0.89 ± 0.02 |
0.71 ± 0.08 |
0.71 ± 0.03 |
Penicillium citrinum F12M5 |
1.04 ± 0.06 |
0.43 ± 0.00 |
0.36 ± 0.00 |
0.86 ± 0.01 |
0.72 ± 0.07 |
0.72 ± 0.01 |
Penicillium citrinum F12M2 |
0.40 ± 0.02 |
0.66 ± 0.01 |
0.15 ± 0.01 |
0.86 ± 0.03 |
0.76 ± 0.05 |
0.74 ± 0.05 |
Penicillium citrinum I22M1 |
1.14 ± 0.05 |
0.19 ± 0.00 |
1.58 ± 0.01 |
0.91 ± 0.03 |
0.74 ± 0.09 |
0.72 ± 0.08 |
Assessment of potential industrial applications of crude enzymes
Juice clarification by crude fungal amylase
The clarification efficiency of crude enzymes derived from halophilic fungi was evaluated by measuring the percentage of residual turbidity and turbidity reduction in orange juice after enzymatic treatment (Table 3; Figure 5). The untreated control exhibited 100% turbidity, while all enzyme-treated samples showed significant reductions in turbidity, demonstrating the clarification potential of these fungal enzymes. Among the tested strains, Aspergillus subalbidus NM-16 exhibited the highest clarification efficiency, reducing turbidity by 32.53 ± 2.01%, followed closely by A. subalbidus NM-17.1 (30.27 ± 3.09%). These values were significantly greater (p < 0.05) than those of A. terreus F11M6 and Penicillium citrinum F12M4.1, which achieved only 7.82 ± 2.28% and 13.83 ± 1.68% turbidity reduction, respectively. Moderate clarification efficiencies were recorded for P. citrinum F12M5, F12M2, and I22M1, which reduced turbidity by approximately 28% with residual turbidity ranging from 71.16% to 72.04%.
These findings highlight the notable potential of crude halophilic fungal enzymes for biotechnological applications, particularly in juice clarification. The high clarification efficiencies achieved by A. subalbidus NM-16 and NM-17.1 correspond with their elevated amylase activities, suggesting that enzymatic degradation of starch and other polysaccharide residues contributes to improved juice transparency. Similar outcomes have been reported in recent studies showing that fungal amylases effectively hydrolyze turbidity-causing polysaccharides, thereby enhancing juice clarity [35,36]. Moreover, the ability of these halophilic enzymes to function efficiently under moderate salinity conditions indicates their suitability for industrial applications in the fruit and beverage sectors, where osmotic fluctuations often hinder conventional enzyme performance.
Table 3 Clarification efficiency of crude enzymes from halophilic fungi.
Fungi |
% Turbidity reduction |
|||
Amylase |
Protease |
|||
Orange juice |
Chocolate |
Chocolate |
Blood |
|
A. subalbidus NM-16 |
53.01 ± 3.48c |
- |
- |
|
A. subalbidus NM-17.1 |
55.26 ± 4.18c |
- |
- |
|
A. terreus F11M6 |
30.53 ± 1.42a |
- |
- |
|
P. citrinum F12M4.1 |
13.83 ± 1.68a |
47.65 ± 4.80b |
90.82 ± 7.69c |
74.34 ± 2.79b |
P. citrinum F12M5 |
28.64 ± 1.23b |
51.11 ± 6.92c |
88.00 ± 9.70b |
69.60 ± 5.00a |
P. citrinum F12M2 |
28.84 ± 4.80b |
50.03 ± 3.30c |
81.91 ± 4.80a |
69.30 ± 6.99a |
P. citrinum I22M1 |
27.96 ± 8.71b |
48.05 ± 4.49b |
95.67 ± 8.85c |
88.99 ± 1.21c |
Values represent mean ± standard deviation (n = 3). Different superscript letters (a-c) within each column indicate statistically significant differences among fungal strains (p < 0.05), as determined by 1-way ANOVA followed by Tukey’s post hoc test. Turbidity reduction was calculated based on the percentage decrease in OD₆₆₀ relative to the respective untreated controls. Dashes (-) indicate treatments that were not tested.
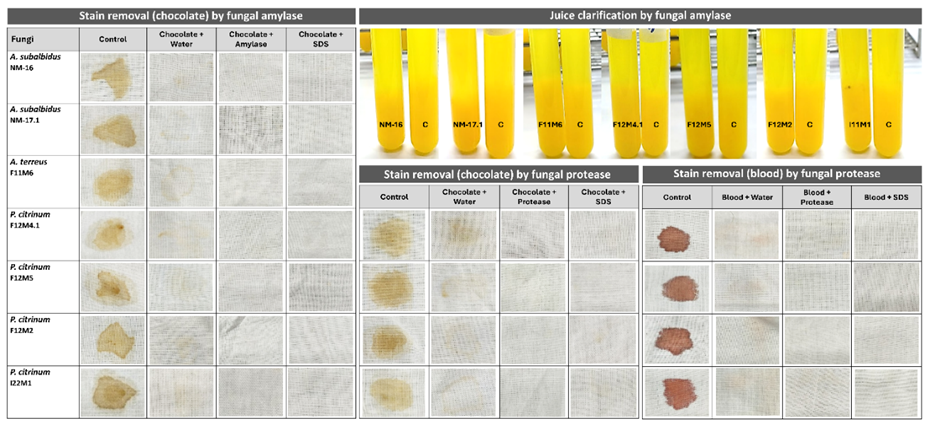
Figure 5 Juice clarification and stain removal efficiency test by crude fungal enzymes.
Stain removal efficiency test
The ability of crude fungal enzymes to clarify chocolate and blood stains was evaluated by measuring the percentage reduction. The results are summarized in Table 3. For chocolate clarification using amylase, A. subalbidus NM-17.1 (55.26 ± 4.18%) and NM-16 (53.01 ± 3.48%) exhibited the highest turbidity reduction, significantly exceeding that of A. terreus F11M6 (30.53 ± 1.42%) (p < 0.05). Among P. citrinum strains, F12M5 (51.11 ± 6.92%), F12M2 (50.03 ± 3.30%), and F12M4.1 (47.65 ± 4.80%) demonstrated comparable clarification efficiency, while I22M1 showed moderate activity (48.05 ± 4.49%). Protease treatment on chocolate stains revealed P. citrinum I22M1 as the most effective strain, achieving 95.67 ± 8.85% turbidity reduction, followed by F12M4.1 (90.82 ± 7.69%), F12M5 (88.00 ± 9.70%), and F12M2 (81.91 ± 4.80%). For blood stain clarification, I22M1 again exhibited the highest efficiency (88.99 ± 1.21%), significantly outperforming F12M5 (69.60 ± 5.00%) and F12M2 (69.30 ± 6.99%). F12M4.1 also showed high proteolytic activity (74.34 ± 2.79%). These findings indicate that P. citrinum I22M1 is particularly effective for both chocolate and blood stain removal, likely due to its strong protease activity, while A. subalbidus NM-17.1 and P. citrinum F12M5 demonstrated notable amylolytic performance.
Stain removal from cotton fabrics
The stain removal potential of crude fungal amylase and protease was evaluated on cotton fabrics stained with melted chocolate and fresh sheep blood (Figure 5). Treatment with distilled water alone was insufficient to eliminate stains, leaving visible residues on the fabric surface. In contrast, both amylase and protease treatments resulted in significant decolorization of chocolate and blood stains, while sodium dodecyl sulfate (SDS) alone also facilitated partial removal due to its surfactant properties. Remarkably, the combined application of enzyme (amylase or protease) with SDS completely removed the stains, restoring the fabric to its original whiteness.
The superior cleaning efficiency observed in enzyme-SDS treatments can be attributed to a synergistic interaction between enzymatic hydrolysis and surfactant-mediated solubilization. Amylase hydrolyzes starch-based residues in chocolate stains, whereas protease catalyzes the degradation of proteinaceous components in blood stains. SDS enhances enzyme accessibility by dispersing hydrophobic or lipidic materials, thereby promoting complete stain dissolution. This cooperative mechanism aligns with previous reports emphasizing enzyme-surfactant synergy in detergent formulations [37,38]. Among the tested isolates, Penicillium citrinum I22M1 exhibited the highest stain-removal efficiency, achieving 95.67% for chocolate and 88.99% for blood stains, consistent with its strong proteolytic activity. These results demonstrate that crude fungal enzymes, particularly from marine-derived P. citrinum, possess excellent stain-removal capacity under mild and environmentally friendly conditions. The findings underscore the potential application of halotolerant fungal enzymes in eco-sustainable detergent formulations and textile cleaning processes.
Conclusions
This study highlights the biotechnological potential of halophilic and halotolerant fungi isolated from marine environments. The identified Aspergillus and Penicillium strains demonstrated strong salt tolerance and the ability to produce stable amylase and protease enzymes under saline conditions. These crude enzymes exhibited effective juice clarification and complete stain-removal capabilities, confirming their functional robustness and eco-friendly potential. Collectively, the findings establish marine-derived halophilic fungi as promising sources of salt-tolerant hydrolases suitable for sustainable applications in the detergent and food industries. Given their halotolerant nature, these fungal isolates also show strong potential for application in highly saline or fluctuating salinity environments, such as salt-rich industrial effluents, marine-based bioprocesses, and saline wastewater treatment systems, where conventional enzymes often lose stability and activity.
Acknowledgements
This research was supported by the National Science, Research and Innovation Fund (NSRF) and Prince of Songkla University, Thailand (Grant No. AGR6701237b).
CRediT author statement
Ninadia Jitprasitporn: Methodology; Data curation; Visualization; Investigation; Lakkhana Kanhayuwa Wingfield: Conceptualization; Supervision; Validation; Writing - Original draft preparation; Writing - Reviewing and Editing.
References
[1] Y He, B Sen and G Wang. Diversity and dynamics of microbial communities in marine ecosystems. Water 2024; 16(1), 31.
[2] SM Tiquia-Arashiro and M Grube. Fungi in extreme environments ecological role and biotechnological significance. Springer Nature, Cham, Switzerland, 2019.
[3] MFM Gonçalves, AC Esteves and A Alves. Marine fungi: Opportunities and challenges. Encyclopedia 2022; 2, 559-577.
[4] H Yaakoub, NS Sanchez, L Ongay-Larios, V Courdavault, A Calenda, JP Bouchara, R Coria and N Papon. The high osmolarity glycerol (HOG) pathway in fungi. Critical Reviews in Microbiology 2022; 48(6), 657-695.
[5] GEB Jones and KL Pang. Introduction marine fungi. In: GEB Jones and KL Pang (Eds.). Marine fungi. De Gruyter, Berlin, Germany, 2012.
[6] GEB Jones, KL Pang, MA Abdel-Wahab, B Scholz, KD Hyde, T Boekhout, R Ebel, ME Rateb, L Henderson, J Sakayaroj, S Suetrong, MC Dayarathne, V Kumar, S Raghukumar, KR Sridhar, AHA Bahkali, FH Gleason and C Norphanphoun. An online resource for marine fungi. Fungal Diversity 2019; 96, 347-433.
[7] Q Li and G Wang. Diversity of fungal isolates from 3 Hawaiian marine sponges. Microbiological Research 2009; 164, 233-241.
[8] N Gunde-Cimerman, P Zalar, S de Hoog and A Plemenitaš. Hypersaline waters in salterns - natural ecological niches for halophilic black yeasts. FEMS Microbiology Ecology 2000; 32(3), 235-240.
[9] A Plemenitaš, T Vaupotič, M Lenassi, T Kogej and N Gunde-Cimerman. Adaptation of the extremely halotolerant black yeast Hortaea werneckii to salinity: A molecular perspective. Frontiers in Microbiology 2014; 5, 199.
[10] J Zajc, T Kogej, EA Galinski, J Ramos and N Gunde-Cimerman. Osmoadaptation strategy of the most halophilic fungus, Wallemia ichthyophaga, growing optimally at salinities above 15% NaCl. Applied and Environmental Microbiology 2014; 80(1), 247-256.
[11] S Hohmann. Osmotic stress signaling and osmoadaptation in yeasts. Microbiology and Molecular Biology Reviews 2002; 66(2), 300-372.
[12] C Gostinčar, J Zajc, A Plemenitaš and N Gunde-Cimerman. Fungal adaptation to extremely high salt concentrations. Fungal Biology Reviews 2011; 25(4), 133-146.
[13] L Butinar, S Santos, I Spencer-Martins, A Oren and N Gunde-Cimerman. Yeast diversity in hypersaline habitats. FEMS Microbiology Letters 2005; 244(2), 229-234.
[14] S Hasan, MI Ansari, A Ahmad and M Mishra. Major bioactive metabolites from marine fungi: A review. Bioinformation 2015; 11(4), 176-181.
[15] N Kango, UK Jana and R Choukade. Fungal enzymes: Sources and biotechnological applications. In: T Satyanarayana, SK Deshmukh and MV Deshpande (Eds.). Advancing frontiers in mycology & mycotechnology: Basic and applied aspects of fungi. Springer, Singapore, 2019, p. 515-538.
[16] H El-Gendi, AK Saleh, R Badierah, EM Redwan, YA El-Maradny and EM El-Fakharany. A comprehensive insight into fungal enzymes: Structure, classification, and their role in mankind’s challenges. Journal of Fungi 2021; 8(1), 23.
[17] RA Samson, ES Hoekstra and JC Frisvad. Introduction to food and airborne fungi. 6th ed. Centraalbureau voor Schimmelcultures, Utrecht, Netherlands, 2002.
[18] FM Dugan. The Identification of fungi-An illustrated introduction with keys, glossary, and guide to literature. The American Phytopathological Society Press, St. Paul, United States, 2015.
[19] D Ellis, S Davis, H Alexiou, R Handke and R Bartley. Description of medical fungi. 3rd ed. South Australia Nexus Print Solutions, Underdale South Australia, Australia, 2016.
[20] LK Wingfield, J Atcharawiriyakul and N Jitprasitporn. Diversity and characterization of culturable fungi associated with the marine sea cucumber Holothuria scabra. PLoS One 2024; 19(1), e0296499.
[21] K Tamura, G Stecher and S Kumar. MEGA11: Molecular evolutionary genetics analysis version 11. Molecular Biology and Evolution 2021; 38, 3022-3027.
[22] L Hankin and SL Anagnostakis. Solid media containing carboxymethylcellulose to detect C x cellulase activity of microorganisms. Journal of General Microbiology 1975; 98(1), 109-115.
[23] RM Teather and PJ Wood. Use of Congo red-polysaccharide interactions in enumeration and characterization of cellulolytic bacteria from the bovine rumen. Applied and Environmental Microbiology 1982; 43(4), 777-780.
[24] GL Miller. Use of dinitrosalicylic acid reagent for determination of reducing sugar. Analytical Chemistry 1959; 31(3), 426-428.
[25] DF Coelho, TP Saturnino, FF Fernandes, PG Mazzola, E Silveira and EB Tambourgi. Azocasein substrate for determination of proteolytic activity: Reexamining a traditional method using bromelain samples. BioMed Research International 2016; 2016(1), 8409183.
[26] C Han, L Kautto and H Nevalainen. Secretion of proteases by an opportunistic fungal pathogen Scedosporium aurantiacum. PLOS One 2017; 12(1), e0169403.
[27] MM Bradford. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analytical Biochemistry 1976; 72(1-2), 248-254.
[28] JA Elegbede and A Lateef. Valorization of corn-cob by fungal isolates for production of xylanase in submerged and solid-state fermentation media and potential biotechnological applications. Waste Biomass Valorization 2018; 9(8), 1273-1287.
[29] UC Banerjee, RK Sani, WAzmi and R Soni. Thermostable alkaline protease from Bacillus brevis and its characterization as a laundry detergent additive. Process Biochemistry 1999; 35(1-2), 213-219.
[30] W Śliżewska, K Struszczyk-Świta, A Otlewska, F Pinzari, L Canfora, K Dybka-Stȩpień, R Napoli, M Migliore, A Manfredini and O Marchut-Mikołajczyk. Halophilic and halotolerant fungi across diverse climates: A comparative study of Polish and Italian soil ecosystems. Frontiers in Microbiology 2025; 16, 1637496.
[31] L Yovchevska, Y Gocheva, G Stoyancheva, J Miteva-Staleva, V Dishliyska, R Abrashev, T Stamenova, M Angelova and E Krumova. Halophilic fungi-Features and potential applications. Microorganisms 2025; 13, 175.
[32] LK Wingfield, N Jitprasitporn and N Che-alee. Isolation and characterization of halophilic and halotolerant fungi from man-made solar salterns in Pattani Province, Thailand. PLoS One 2023; 18(2), e0281623.
[33] I Pócsi, J Dijksterhuis, J Houbraken and RP de Vries. Biotechnological potential of salt tolerant and xerophilic species of Aspergillus. Applied Microbiology and Biotechnology 2024; 108(1), 521.
[34] IB Hmad and A Gargouri. Halophilic filamentous fungi and their enzymes: Potential biotechnological applications. Journal of Biotechnology 2024; 381, 11-18.
[35] G Ozyilmaz and E Gunay. Clarification of apple, grape and pear juices by co-immobilized amylase, pectinase and cellulase. Food Chemistry 2023; 398, 133900.
[36] T Roheen, R Ramzan, W Khalid M Nadeem, FA Atif, M Munir and TM Qureshi. Synthesis and characterization of CMC/PAM-amy hydrogel and its efficacy in apple juice clarification. Processes 2024; 12, 2264.
[37] K Farooq, Z Anwar, S Hasan, F Afzal, M Zafar, U Ali, O Alghamdi, A AL-Farga and SM Al-maaqar. Optimization and detergent compatibility of protease produced from Aspergillus oryzae by utilizing agro wastes. ACS Omega 2024; 9(15), 17446-17457.
[38] X Wang, X Qin, L Tong, J Zheng, T Dong, X Wang, Y Wang, H Huang, B Yao, H Zhang and H Luo. Improving the catalytic activity of a detergent-compatible serine protease by rational design. Microbial Biotechnology 2023; 16(5), 947-960.