
Trends
Sci.
2026;
23(7):
12666
Cyclooxygenase Modulatory Activity of γ-Oryzanol from Pathumthani 1 Rice Bran: Integration of HPLC Composition, Cell Assays, and Molecular Docking
Shisanupong Anukanon1,2, Kanyanat Kaewiad3, Sakunta Manakla4, Jureemart Deeammart4, Komgrit Saeng-ngoen5, Yawanart Ngamnon6, Ngamnetr Rapan6 and Yaiprae Chatree4,*
1School of Medicine, Mae Fah Luang University, Chiang Rai 57100, Thailand
2Integrative Natural Therapeutics and Health Innovation Research Unit, School of Medicine,
Mae Fah Luang University, Chiang Rai 57100, Thailand
3Expert Center of Innovative Herbal Products, Thailand Institute of Scientific and Technological Research,
Pathum Thani 12120, Thailand
4Faculty of Science and Technology, Valaya Alongkorn Rajabhat University under the Royal Patronage,
Pathum Thani 13180, Thailand
5Faculty of Agricultural Technology, Valaya Alongkorn Rajabhat University under the Royal Patronage,
Pathum Thani 13180, Thailand
6Science Center, Valaya Alongkorn Rajabhat University under the Royal Patronage, Pathum Thani 13180, Thailand
(*Corresponding author’s e-mail: [email protected])
Received: 15 November 2025, Revised: 12 December 2025, Accepted: 22 December 2025, Published: 25 February 2026
Abstract
Gamma (γ)-Oryzanol, a mixture of ferulic acid esters of plant sterols and triterpene alcohols, exhibits notable antioxidant and anti-inflammatory properties. This study investigated the composition and cyclooxygenase (COX) modulatory activity of γ-oryzanol extract obtained from Oryza sativa L. var. indica cv. Pathumthani 1 (PTT1) rice bran. Quantitative HPLC analysis indicated a total γ-oryzanol content of 71.93 ± 8.74 mg/100 g extract, with cycloartenyl ferulate, 24-methylenecycloartanyl ferulate, campesteryl ferulate, and sitosteryl ferulates as the primary constituents. COX activity assays in LPS-stimulated RAW 264.7 macrophages demonstrated that the extract selectively inhibited COX-2 activity in a dose-dependent manner, whereas COX-1 inhibition was not statistically significant, indicating enzyme selectivity. Complementary molecular docking analysis further supported these findings, showing that all steryl ferulates exhibited stable binding conformations within the COX-2 catalytic cleft with low RMSD values. Among them, campesteryl ferulate displayed the most favorable binding energy toward both COX isoforms. Collectively, these results emphasize the possible use of γ-oryzanol from PTT1 rice bran as a natural COX-2 selective modulator with therapeutic relevance in inflammatory regulation.
Keywords: Pathumthani 1 (PTT1) rice bran, γ-Oryzanol, Steryl ferulates, Campesteryl ferulate, Anti-inflammatory, COX-2 selectivity, Molecular docking
Introduction
Gamma-oryzanol (γ-Oryzanol) is a valuable bioactive compound predominantly found in rice bran extracts with nutraceutical benefits, including antioxidant and anti-inflammatory effects [1]. γ-Oryzanol is a mixture of ferulic acid esters of phytosterols and triterpene alcohols (Steryl ferulates), primarily composed of cycloartenyl ferulate, 24-methylenecycloartanyl ferulate, campesteryl ferulate, and sitosteryl ferulate [2]. Recently, γ-oryzanol has gained extensive attention due to its diverse biological activities, not only antioxidant and anti-inflammatory but also for treating menopausal symptoms and enhancing pituitary function, as well as lipid-lowering effects [3,4]. The important role of γ-oryzanol in anti-inflammatory properties was underlying the modulation of cyclooxygenase (COX) enzymes, particularly COX-1 and COX-2, which are responsible for the conversion of arachidonic acid into prostaglandins during inflammatory responses [5,6]. These lipid mediators bind to specific receptors to modulate inflammatory responses and support physiological processes such as cell proliferation and tissue repair [7].
COX-1
is constitutively expressed in most tissues and is responsible for
producing prostaglandins that perform housekeeping functions. These
vital functions include maintaining the integrity of the
gastrointestinal lining, regulating blood flow in the kidneys, and
promoting platelet aggregation for blood clotting [8,9]. Inducible
and upregulated COX-2 occurs during inflammation, particularly in
macrophages stimulated by lipopolysaccharide (LPS) and other
inflammatory mediators. This differential regulation provides an
opportunity for targeted therapies that inhibit COX-2 while
preserving the protective functions of COX-1, thus minimizing side
effects associated with traditional nonsteroidal anti-inflammatory
drugs (NSAIDs) [10,11]. Inflammatory mediators, particularly an
inducible COX-2, play an important role in prostaglandin production,
mediating pain and fever
in response to inflammatory stimuli
such as
pro-inflammatory cytokines and inflammatory
substances. It has been reported that γ-oryzanol in rice bran oil
inhibited LPS-mediated COX-2 expression by suppressing
Erk1/2-mediated Egr-1 expression [6]. Individual steryl ferulates
attenuate inflammation through inhibition of COX activity and
downregulation of inflammatory mediators in macrophages [12].
Pathum Thani 1 (PTT1) rice (Oryza sativa L. var. indica cv. Pathumthani 1) is a high-yielding and widely cultivated Thai rice variety known for its fragrant aroma and low-amylose content [13]. Its bran provides substantial nutritional value, containing balanced amounts of carbohydrates, proteins, and lipids in the pericarp [14,15]. Notably, PTT1 rice bran has been reported to contain relatively high levels of γ-oryzanol compared with other Thai rice varieties, along with considerable total polyphenol content that contributes to its antioxidant potential [16]. These characteristics underscore the relevance of PTT1 rice bran as a rich source of bioactive compounds, yet data specifically linking the composition of its γ-oryzanol to COX-modulating activity remain limited. Moreover, few studies have combined compositional analysis with computational modeling to clarify these potential interactions. Integrating HPLC with molecular docking enables both precise identification of γ-oryzanol constituents and prediction of their binding behavior toward COX isoenzymes, providing a more coherent framework for understanding its biological activity. Therefore, this study aimed to characterize γ-oryzanol from PTT1 rice bran using high-performance liquid chromatography (HPLC) and to evaluate its inhibitory effects on COX-1 and COX-2 activities in LPS-stimulated RAW264.7 macrophage cells. In addition, molecular docking analysis was performed to elucidate the binding interactions of γ-oryzanol with COX isoenzymes, highlighting its potential as a selective cyclooxygenase inhibitor and a promising rice bran-derived anti-inflammatory nutraceutical.
Materials and methods
Chemicals and reagents
Analytical-grade ethanol and acetone (Thermo Fisher Scientific, Milford, USA) were used as extraction solvents. For HPLC analysis, HPLC-grade methanol, acetonitrile, dichloromethane, and acetic acid (Thermo Fisher Scientific, Milford, USA) were employed as mobile-phase solvents. The γ-oryzanol standard was obtained from TCI America (Portland, OR, USA).
Cell cultures
RAW264.7 macrophages were obtained from the American Type Culture Collection (ATCC, Manassas, Virginia, USA). Cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM; Gibco, Thermo Fisher Scientific, England) supplemented with 10% fetal bovine serum (FBS; Gibco, Thermo Fisher Scientific, USA) and 1% antibiotics (100 g/mL streptomycin and 100 U/mL penicillin, Gibco, Thermo Fisher Scientific, USA). Cells were cultured at 37 °C in a humidified 5% CO2 incubator (Sheldon Manufacturing, USA).
Cell viability assay
RAW264.7 cells (1×103 cells/well of 96 well plate) were incubated in a humidified incubator (37 °C and 5% CO2) for 24 h. After incubation, cells were treated with oryzanol extract (0.15, 0.31, 0.62, 1.25, 2.50, 5.00, 10.00 and 20.00 mg/mL) for 24 h. Then, 10 µL of 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT) was added to each well. After 3 h at 37 °C in the dark, absorbance was read at 570 nm using a multi-well microplate reader (Infinite M200 PRO, TECAN, Australia). Sodium dodecyl sulfate (SDS) was used as a positive control due to its well-characterized membrane-disrupting cytotoxicity in in vitro viability assays [17]. A viability rate at > 80% was considered for the tested concentration.
Cyclooxygenase activity assay
Cyclooxygenase activity assay was performed using COX-1 and COX-2 activity assay kit (Cat# ab204699, Abcam, USA) and we followed manufacturer’s protocol. Raw 264.7 cells (1×106 cells in 60 mm cell culture dish) were incubated in a humidified incubator (37 °C and 5% CO2) for 24 h. After incubation, cells were treated with 10 µg/mL LPS additional of γ-oryzanol extract (0, 0.62, 1.25, and 2.50 mg/mL) for 24 h. Then, cells were harvested and lysed in ice-cold lysis buffer. Cell lysates were added to appropriate wells, followed by the addition of assay buffer, arachidonic acid, and COX-1 or 2 inhibitors (SC-560 or Celecoxib), sequentially. After 1 h at 37 °C in the dark, fluorescence was read at excitation 535 and emission 587 nm using a multi-well microplate reader (Infinite M200 PRO, TECAN, Australia).
Plant material
Rice bran sample was obtained from PTT 1 rice cultivated at Small and Micro Community Enterprise field (Lamlukka Subdistrict, Amphoe Lam Luk Ka, Pathum Thani province: 13.9912° N, 100.7950° E). Rice was cultivated using certified organic farming methods, which certified to Organic Thailand standards. Rice grain samples were harvested in September 2024 and subjected to the standard process to separate bran using a rice mill. Rice bran sample was stored immediately at −20 °C after milling until analysis.
γ-oryzanol extraction
The extraction of γ-oryzanol was carried out using a modified method according to the oryzanol-crystallization procedure [13]. The rice bran sample was macerated with 95% ethanol (1:6 w/v) on an orbital shaker at 200 rpm for 1 h. To remove solvent, supernatants were filtrated using Whatman No. 1 filter paper and evaporated to remove ethanol. The extract was dewaxed and degummed by dissolution in acetone (1:6, v/v) at 60 °C for 1 h, yielding a clear solution. For crystallization, 10 g of the extract were dissolved in an ethanol–acetone mixture (7:3, v/v) and maintained at −20 °C for 5 days. The resulting solid phase was recovered by filtration at −20 °C and used as the starting material for this study.
Determination of active compounds using high-performance liquid chromatography
Quantification of total γ-oryzanol concentrations in oryzanol extract was determined using high-performance liquid chromatography (HPLC). A reversed-phased HPLC (C18 column, 4.6×150 mm, i.d. 5 μm particle size; GL Sciences, Tokyo, Japan) with UV/Visible detection (Waters e2695 separations module, Milford, USA) was employed. The mobile phase consisted of methanol, acetonitrile, dichloromethane, and acetic acid (50, 44, 3 and 3%, respectively) under isocratic condition. The wavelength of the detector was set at 330 nm and the flow rate was 0.8 mL/min. Samples were prepared through isopropanol extraction with rigorous vortexing and incubated at 30 °C for 3 min. To collect supernatant, centrifugation at 8,000 rpm for 5 min was performed, followed by filtration using a 0.45 μm membrane. The concentrations of γ-oryzanol content were reported by calculating their standard curve.
Molecular docking validation and simulation procedures
Protein preparation and active site definition
Cyclooxygenase-1 (COX-1, PDB ID: 1EQG) and cyclooxygenase-2 (COX-2, PDB ID: 3LN1) crystal structures were obtained from the RCSB Protein Data Bank [18,19]. After checking the structures for completeness, Discovery Studio Visualizer 2021 (BIOVIA, San Diego, Dassault Systèmes) was used to eliminate heteroatoms, co-crystallized ligands, and unnecessary water molecules. AutoDockTools 1.5.7 was used to assign Gasteiger charges and add polar hydrogens [20]. Using the native co-crystallized ligands, the active site center (grid box centroid) was defined. Conserved water molecules that mediated hydrogen bonding close to the catalytic residues Tyr355 and Arg120 were kept in hydrated systems [21].
Redocking validation
The docking protocol was validated by re-docking the native ligands of COX-1 and COX-2 into their crystallographic binding sites utilizing AutoDockTools 1.5.7 [22]. Grid maps were created using box sizes of 40×40×40, 60×60×60, 80×80×80, and 100×100×100 Å, centered at the centroid of the crystallographic ligand. Each system was evaluated with and without conserved crystallographic waters, defined as those within 5 Å of the ligand heavy atoms, to examine the impact of hydration on pose accuracy. Docking employed the Lamarckian Genetic Algorithm, conducting 100 GA runs for each ligand. Each configuration’s poses were clustered with an RMSD tolerance of 2.0 Å. Rather than choosing a single “best-ranked” pose, the most populated cluster (Cluster 1) was utilized as a representative of the docking solution, with cluster-level metrics reported. The validation quality was assessed by calculating the cluster-mean root-mean-square deviation (RMSD) between the predicted ligand poses and the crystallographic data. Cluster occupancy percentages and mean binding energies were recorded for each grid configuration.
Ligand preparation
The γ-oryzanol mixture comprised four principal steryl ferulate components (P1–P4). Furthermore, ibuprofen (IBP) and celecoxib (CEL) were utilized as reference inhibitors for COX-1 and COX-2, respectively. Ligand structures were sourced from PubChem in SDF format and subsequently converted to PDBQT format utilizing Open Babel 3.1.1. Ligands underwent energy minimization using the MMFF94 force field, with all torsional bonds designated as rotatable.
Molecular docking of γ-oryzanol components
Molecular docking of P1–P4, IBP, and CEL was conducted on COX-1 and COX-2 targets utilizing the validated grid parameters. Each ligand was docked under two conditions: (i) in the absence of water and (ii) with conserved water molecules present. The docking parameters were maintained consistent with the validation phase (LGA, 100 runs). The binding energy (ΔG, kcal/mol), cluster occupancy (%), and average RMSD of Cluster 1 were evaluated for each compound. The selectivity index (SI) values were determined by calculating the ratio of the estimated Ki for COX-1 to that for COX-2. The binding modes were analyzed using Discovery Studio Visualizer, emphasizing hydrogen bonding, π-π interactions, and hydrophobic contacts with critical residues (Arg120, Tyr355, Tyr385, Ser530).
Integration of HPLC composition with molecular docking analysis
To assess the individual contributions of steryl ferulates (P1–P4) to the overall COX inhibitory potential of the γ-oryzanol extract, quantitative HPLC data were combined with molecular docking results. The mass fractions (fi) of each compound were calculated based on their concentrations determined by HPLC as follows: fi = Ci/∑Ci, where Ci represents the concentration (mg/100 g) of compound i within the extract. The validated binding energies (ΔGi) derived from AutoDock (grid size 60 Å, including water) were utilized for weighting purposes. This composition–affinity weighting model was formulated as a semi-empirical hybrid approach that integrates fundamental principles from thermodynamic binding theory, Boltzmann population weighting, and quantitative mixture modeling [23-25]. The model assumes that the magnitude of the binding free energy (|ΔG|) is proportional to inhibitory potency and that each compound contributes to the overall activity in proportion to its abundance (fᵢ) and binding strength [23]. The contribution of each compound to the overall activity was estimated using Eq. (1).
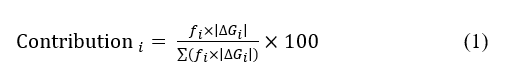
Eq. (1) represents a semi-empirical, thermodynamically inspired approximation that combines the relative abundance (fi) of each steryl ferulate with its docking-derived binding free energy (ΔGi). This model does not constitute a rigorous thermodynamic calculation, and the predicted contributions should therefore be interpreted as relative trends rather than absolute quantitative effects. Similar semi-empirical weighting approaches have been used in mixture-based computational pharmacology where experimental binding constants are unavailable.
This model was adapted and proposed in the present work to approximate the relative contribution of each compound based on the assumption that the magnitude of binding free energy correlates with inhibitory potency.
Statistical analysis
Data from cyclooxygenase activity, redocking, and production docking were compiled in GraphPad Prism 10.6.1. Data normality was assessed using the Shapiro–Wilk test to determine the appropriate correlation method. Non-parametric Spearman’s correlation (ρ) was applied to evaluate relationships between grid size, RMSD, and binding energy, as well as the influence of water presence. Significance was considered at p-value < 0.05. All numerical data were expressed as mean ± SD from three independent docking repetitions (n = 3 per grid configuration).
Results and discussion
Total γ-oryzanol content and composition of steryl ferulates
Triplicate HPLC analyses of the γ-oryzanol extract identified four consistent peaks corresponding to individual steryl ferulates (Figure 1). The mean retention times (Rt, min ± SD, n = 3) were as follows: P1 (cycloartenyl ferulate) 30.05 ± 1.05, P2 (24-methylenecycloartanyl ferulate) 33.38 ± 1.19, P3 (campesteryl ferulate) 35.82 ± 1.33, and P4 (sitosteryl ferulate) 41.54 ± 1.58. Minor variations in Rt among replicates (< 1.5 min) demonstrated high chromatographic reproducibility.
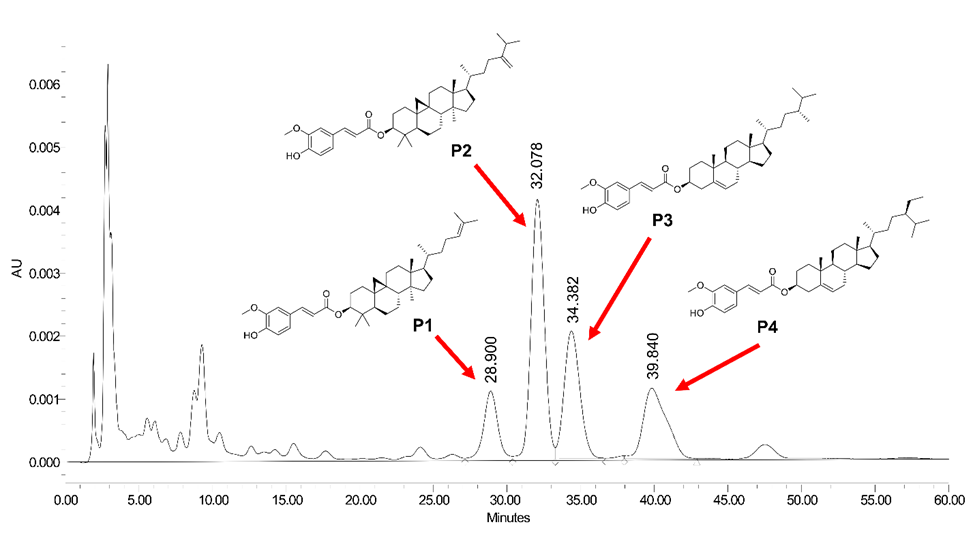
Figure 1 Representative HPLC chromatogram of γ-oryzanol extract showing four major steryl ferulate components: P1 cycloartenyl ferulate), P2 (24-methylenecycloartanyl ferulate), P3 (campesteryl ferulate), and P4 (sitosteryl ferulate).
Table 1 Total γ-oryzanol content and composition of steryl ferulates (P1-P4) in extract.
|
Steryl ferulate contents (mg/100 g) |
Total γ-oryzanol content (mg/100 g) |
|||
Cycloartenyl ferulate P1 |
24-methylenecycloartanyl ferulate P2 |
Campesteryl ferulate P3 |
Sitosteryl ferulate P4 |
||
γ-oryzanol extract |
12.45 ± 0.10 |
30.82 ± 0.07 |
16.24 ± 0.12 |
12.41 ± 0.19 |
71.93 ± 8.74 |
The value is the mean ± S.D. from three independent experiments (n = 3).
The total γ-oryzanol content and individual steryl ferulate composition of the γ-oryzanol extracted from PTT1 rice bran, as determined by HPLC analysis, are presented in Table 1. The total γ-oryzanol concentration was 71.93 ± 8.74 mg/100 g extract. Among the four major steryl ferulate constituents, 24-methylenecycloartanyl ferulate (P2) was the predominant component (30.82 ± 0.07 mg/100 g), followed by campesteryl ferulate (P3, 16.24 ± 0.12 mg/100 g), cycloartenyl ferulate (P1, 12.45 ± 0.10 mg/100 g), and sitosteryl ferulate (P4, 12.41 ± 0.19 mg/100 g). Steryl ferulate composition might be responsible for major phytostituent in anti-inflammation modulation of cyclooxygenase activity.
Cell viability
The control group exhibited 100% cell viability, whereas the positive control (sodium dodecyl sulfate, 0.1 mg/mL) reduced viability to 16%. Treatment with the test extract at concentrations of 0.15, 0.31, 0.62, 1.25, 2.50, 5.00, 10.00, and 20.00 mg/mL resulted in cell viabilities of 108.19 ± 0.67%, 107.05 ± 0.85%, 106.30 ± 1.52%, 105.16 ± 1.63%, 96.54 ± 2.78%, 79.87 ± 0.58%, 76.04 ± 1.06%, and 68.25 ± 0.58%, respectively. The calculated IC₅₀ was 25.52 ± 1.21 mg/mL. Based on these results, concentrations maintaining viability above 80% were selected for subsequent COX activity assays in RAW264.7 macrophages.
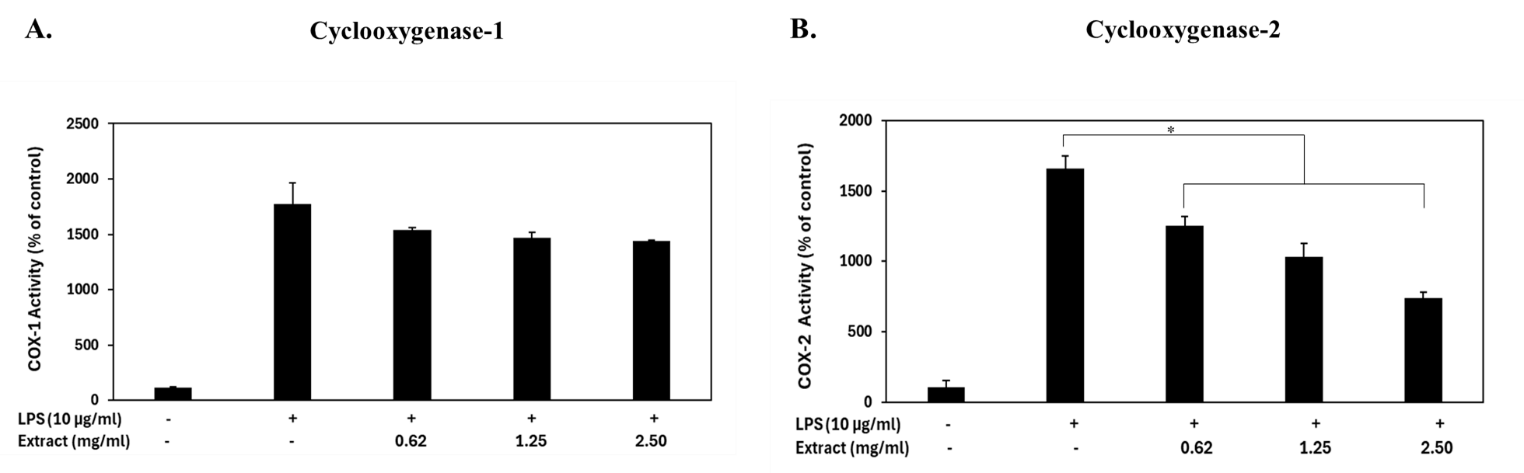
Figure 2 Cyclooxygenase activity in RAW 264.7 macrophage cells treated with γ-oryzanol extract from rice bran and lipopolysaccharide (LPS) for 24 h. Enzyme activities are expressed as a percentage of the untreated control. (A) COX-1 and (B) COX-2 activities were determined based on prostaglandin G₂ (PGG₂) formation, which reacts with a COX-specific probe to generate a fluorescent signal proportional to enzyme activity. Data represent the mean ± standard deviation (SD) of three independent experiments. *p-value < 0.05 indicates a significant difference compared with the LPS-only treatment group.
Cyclooxygenase inhibitory activity of γ-oryzanol on LPS-induced inflammation
γ-oryzanol extracts contributed to inhibiting LPS-stimulated cyclooxygenase activity in Raw 264.7 macrophage cells (Figures 2(A) - 2(B)). γ-oryzanol extracts decreased the activity of particular COX-2 enzymes in LPS-stimulated Raw 264.7 cells, while COX-1 activity remained largely unaffected. In the COX-1 assay, LPS stimulation markedly increased COX-1 activity to 1770.41 ± 194.80% relative to the unstimulated control. Treatment with γ-oryzanol extract at concentrations of 0.62, 1.25, and 2.50 mg/mL resulted in COX-1 activity levels of 1539.53 ± 21.64%, 1467.38 ± 50.50%, and 1438.52 ± 7.21% of control, respectively. These values correspond to 13.04 ± 1.73%, 17.12 ± 4.03%, and 18.75 ± 0.58% inhibition, indicating no significant difference compared with the LPS-stimulated group.
In
contrast, γ-oryzanol exhibited a marked effect on COX-2
activity. LPS stimulation elevated COX-2 activity to 1662.19 ±
64.93% of the control. Treatment with γ-oryzanol extract at 0.62,
1.25, and 2.50 mg/mL significantly reduced COX-2 activity to
1,250.93 ± 64.93%, 1,034.48 ± 93.79%, and
738.67 ±
43.28% of control, corresponding to
24.74 ± 5.52%, 37.76 ±
7.98%, and 55.56 ± 3.68% inhibition, respectively (p-value
< 0.05 compared with LPS group). These findings indicate that
γ-oryzanol selectively suppresses COX-2 activity in LPS-induced
inflammatory conditions in RAW 264.7 cells. Additionally, γ-oryzanol
exhibits a dose-dependent relationship that could potentially
enhance its therapeutic potential and significantly modulate
inflammatory responses. A molecular approach is required to fully
comprehend its mechanisms and potential therapeutic applications.
Correlation between grid parameters and docking performance
Spearman’s rank correlation analysis was conducted to assess the impact of grid box size and the presence of conserved water molecules on the reproducibility of docking for COX-1 and COX-2. The data presented in Table 2 indicates a positive monotonic correlation between grid size and both reference RMSD and mean binding energy, suggesting that an increase in search box dimensions diminishes redocking precision and reduces binding affinity.
The association between grid size and RMSD was mild for COX-1 (p-value = 0.54 - 0.56, p-value > 0.05), while it was stronger and statistically significant for COX-2 (p-value = 0.86, p-value = 0.0008 for the water-containing system). It was further confirmed that larger boxes provided fewer favorable (less negative) binding energies, since grid size showed a strong positive association with the mean binding energy in both enzymes (p-value = 0.91 - 0.95, p-value < 0.05).
The presence of water molecules did not produce statistically significant differences in RMSD (p-value = 0.29 - 0.88, p-value > 0.05) or binding energy (p-value = −0.88, p-value > 0.05). Nonetheless, a weak negative trend in binding energy indicated that the inclusion of conserved water could marginally stabilize ligand–protein interactions.
A grid box size of 60×60×60 Å was chosen for the following docking of γ-oryzanol components (P1 - P4) based on these correlations, as it represents the ideal trade-off between search-space coverage and docking accuracy. Both with-water and without-water configurations were maintained to assess the impact of hydration on ligand binding.
Table 2 Correlation between grid size, water inclusion, and docking performance parameters (RMSD and binding energy) for COX-1 and COX-2 docking simulations.
Correlation pair |
Target enzyme |
ρ (Spearman’s rho) |
95% CI |
p-value (two-tailed) |
Grid size vs RMSD |
COX-1 (no water) |
0.540 |
−0.07 - 0.86 |
0.079 |
COX-1 (with water) |
0.561 |
−0.04 - 0.86 |
0.066 |
|
COX-2 (no water) |
0.669 |
0.14 - 0.90 |
0.022 |
|
COX-2 (with water) |
0.864 |
0.56 - 0.96 |
0.0008 |
|
Grid size vs BE |
COX-1 (no water) |
0.907 |
0.68 - 0.97 |
0.0002 |
COX-1 (with water) |
0.928 |
0.75 - 0.98 |
< 0.0001 |
|
COX-2 (no water) |
0.821 |
0.45 - 0.95 |
0.0022 |
|
COX-2 (with water) |
0.950 |
0.82 - 0.99 |
< 0.0001 |
|
Water (0 = remove,1 = with) vs RMSD |
COX-1 |
0.293 |
- |
0.700 |
COX-2 |
0.878 |
- |
0.100 |
|
Water (0 = remove, 1 = with) vs BE |
COX-1 |
−0.878 |
- |
0.100 |
COX-2 |
−0.878 |
- |
0.100 |
Molecular docking performance
All γ-oryzanol steryl ferulates (P1 - P4) exhibited a consistent binding orientation within the COX-2 catalytic cleft. The bulky hydrophobic steryl moieties were consistently oriented toward the hydrophobic wall formed by Phe504, Leu338, Val335, and Leu517, whereas the polar ferulate heads extended toward the solvent-accessible entrance of the active site. None of the γ-oryzanol constituents interacted with the key catalytic residues of COX-2 (Arg120, Tyr355, and Ser530)
The comparative docking results of the γ-oryzanol steryl ferulates (P1 - P4) and reference COX inhibitors are summarized in Table 3 and Figure 3. P3 (campesteryl ferulate) demonstrated the highest affinity for COX-1 and COX-2, with the lowest binding energies recorded at −11.28 and −12.86 kcal/mol, respectively. The optimal ΔG indicates the most advantageous single docking pose that aligns with the theoretical minimum energy configuration between the ligand and enzyme. The mean ΔG of Cluster 1 (−4.61 and −5.20 kcal/mol) indicates the average stability within the predominant ensemble of poses. These findings indicate that P3 forms a deep-binding conformation and maintains a stable and reproducible interaction within the catalytic pocket. The negative ΔΔG (−1.58 kcal/mol) and selectivity index (SI = 0.88) suggest a slight preference for COX-2, aligning with its moderate cluster occupancy (43% compared to 22%).
The binding conformation of P3 in the COX-2 active site exhibited distinct variations between the hydrated and non-hydrated docking systems (Figure 4). In the hydrated COX-2 model, the steryl backbone of P3 was closely integrated within the hydrophobic cleft, establishing van der Waals interactions with Phe504, Leu338, Val509, Ala513, Met99, Val335, and Leu517 at interatomic distances of 3.5 - 4.0 Å, significantly shorter than the distances recorded in the non-hydrated system (5.4 - 5.6 Å). The ferulate aromatic ring formed π-σ and π-alkyl interactions with Val74 and Arg106, while the hydroxyl and methoxyl groups engaged in hydrogen bonding with Pro71, Glu510, and a bridging water molecule (HOH625), exhibiting bond lengths of 2.6 - 2.9 Å. The non-hydrated model displayed a reduced number of hydrogen bonds with extended distances (3.2 - 3.8 Å), leading to a less compact ligand orientation and a slight displacement of the ferulate moiety from Glu510.
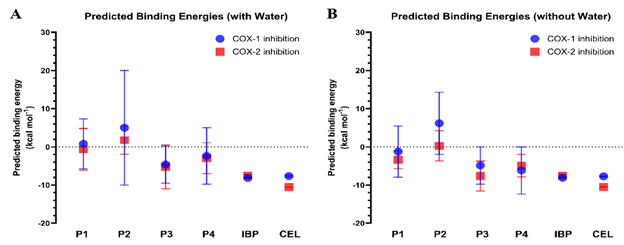
Figure 3 Predicted binding energies (ΔG) of γ-oryzanol steryl ferulates (P1 - P4) toward COX-1 and COX-2 under hydrated and non-hydrated conditions. (A) Predicted binding energies of P1 - P4 and reference inhibitors (ibuprofen, IBP; celecoxib, CEL) docked into COX-1 and COX-2 with conserved water molecules retained in the active sites. (B) Predicted binding energies obtained from docking simulations without water molecules. Data are expressed as mean ± SD from the most populated cluster of poses.
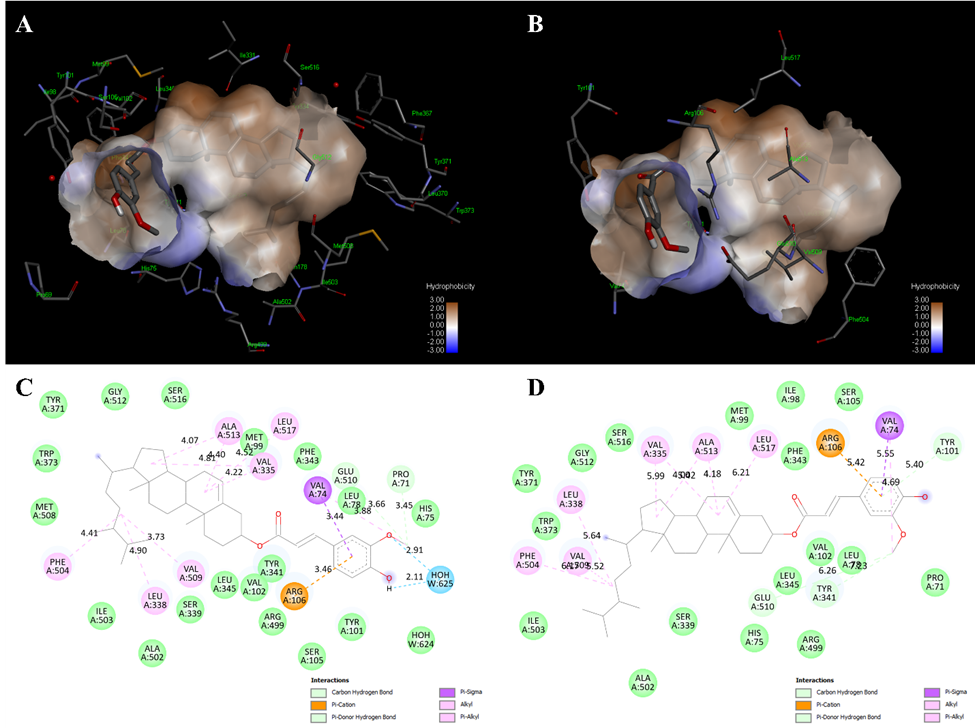
Figure 4 Comparative binding modes of campesteryl ferulate (P3) in COX-2 under hydrated and non-hydrated conditions. (A) - (B) Hydrophobic receptor surface representation of COX-2 complexed with P3 in the presence (A) and absence (B) of conserved water molecules. (C) - (D) Two-dimensional interaction diagrams of P3 bound to COX-2 with (C) and without (D) water molecules. Hydrogen bonds (green and blue dashed lines), π–π/π–σ interactions (purple), and hydrophobic contacts (pink) are indicated.
P1
(cycloartenyl ferulate) also exhibited a favorable affinity toward
COX-2 (ΔΔG = −1.61 kcal/mol, SI = 0.80). Although its best pose
was weaker than that of P3, the mean ΔG values (0.79 and −0.63
kcal/mol) suggest higher conformational variability and less stable
clustering compared to P3, possibly due to partial accommodation of
the cycloartenyl ring system within the COX-2 pocket. In contrast,
P2
(24-methylenecycloartanyl ferulate) showed the weakest
binding energies (−5.21 and −5.13 kcal/mol) and minimal energy
difference between isoforms
(ΔΔG = +0.08 kcal/mol, SI =
1.02), indicating nearly non-selective binding. P4 (sitosteryl
ferulate), with its bulkier sterol moiety, favored COX-1 binding
(ΔΔG = +3.12 kcal/mol, SI = 1.42). Despite showing
relatively low best ΔG for COX-1 (−10.61 kcal/mol), its small
COX-2 cluster occupancy (7%) and modest mean ΔG (−2.98 kcal/mol)
suggest unstable interactions within the narrower COX-2 secondary
pocket, possibly due to steric hindrance.
For comparison, the reference NSAIDs reflected their known pharmacological selectivity. Celecoxib showed a strong and consistent COX-2 selectivity (ΔΔG = −2.84 kcal/mol, SI = 0.73) with complete clustering (100%), validating the reliability of the docking protocol. Ibuprofen, in contrast, produced nearly equivalent affinities for both isoforms (ΔΔG = +0.59 kcal/mol, SI = 1.08), confirming its non-selective binding profile.
Table 3 Comparative docking results of γ-Oryzanol-Derived steryl ferulates and reference COX inhibitors (Ligands prepared from PubChem, validated grid 60×60×60 Å, with conserved water molecules).
Ligand |
Best ΔG (kcal/mol)a |
ΔΔGb |
Cluster1 Occupancy (%)c |
Mean ΔG of Cluster1d |
Selectivity Index (SI) |
|||
COX-1 |
COX-2 |
COX-1 |
COX-2 |
COX-1 |
COX-2 |
|||
P1 |
−6.36 |
−7.97 |
−1.61 |
45 |
36 |
0.79 |
−0.63 |
0.80 |
P2 |
−5.21 |
−5.13 |
+0.08 |
29 |
22 |
5.01 |
1.74 |
1.02 |
P3 |
−11.28 |
−12.86 |
−1.58 |
43 |
22 |
−4.61 |
−5.20 |
0.88 |
P4 |
−10.61 |
−7.49 |
+3.12 |
31 |
7 |
−2.35 |
−2.98 |
1.42 |
Ibuprofen |
−8.34 |
−7.75 |
+0.59 |
100 |
63 |
−8.14 |
−7.50 |
1.08 |
Celecoxib |
−7.86 |
−10.70 |
−2.84 |
36 |
100 |
−7.65 |
−10.56 |
0.73 |
aValues represent the best (lowest) binding free energy (ΔG, kcal/mol) and corresponding cluster characteristics derived from 100 genetic algorithm runs.
bΔΔG = ΔG(COX-2) − ΔG(COX-1); negative values indicate COX-2 preference.
cCluster1 occupancy denotes the percentage of docking poses belonging to the top-ranked conformational cluster.
dMean ΔG of Cluster1 represents the average energy of all poses within that cluster.
eSelectivity index (SI) was calculated as | Best ΔG(COX-1)| / | Best ΔG(COX-2)|, where SI < 1 indicates COX-2 selectivity and SI > 1 indicates COX-1 preference.
Composition-activity integration
HPLC quantification identified four main steryl ferulates in the γ-oryzanol extract: P1 (cycloartenyl ferulate, 17.3%), P2 (24-methylenecycloartanyl ferulate, 42.9%), P3 (campesteryl ferulate, 22.6%), and P4 (sitosteryl ferulate, 17.3%). To assess the energetic contribution of each compound, the composition fractions (fᵢ) were combined with their mean ΔG values from Cluster 1, which reflects the average binding energy within the most populated docking cluster (grid 60 Å, with water) (Table 4).
Based on the semi-empirical weighting model, P3 was predicted to have the largest relative contribution to COX-2 binding (~37%), followed by P2 (~28%), P1 (~18%), and P4 (~17%). These values should be interpreted qualitatively as indicative trends rather than exact biochemical contributions. P3 (34%) and P4 (32%) were the primary contributors for COX-1, aligning with their higher mean binding energies. These findings suggest at the computational level that steryl ferulates with more favorable ΔG values (such as P3 and P4) may contribute more strongly to the predicted inhibitory profile; however, this does not establish their quantitative biological contribution in the cellular context. The application of mean ΔG, as opposed to best-pose energy, highlights the comparative stability of ligands and offers a more accurate assessment of their relative contributions within the mixed extract.
An important limitation of our composition activity integration is that the predicted energetic contributions depend on a semi-empirical model that uses docking-derived ΔG values. As a result, compounds such as P3 exhibit disproportionately high predicted contributions because of their favorable ΔG values, despite being less abundant than P2. This discrepancy reflects the inherent limitation of relying on docking energies to infer quantitative contributions within a complex phytochemical mixture. Thus, the results should be interpreted qualitatively, and future validation using purified constituents and experimental binding assays is warranted.
Table 4 Composition-weighted contributions of individual steryl ferulates to COX-1 and COX-2 inhibitory potential (linear model, grid 60 Å with water).
Compound |
Fraction (fi)a |
ΔG (COX-1)b |
ΔG (COX-2)b |
Contribution (%) COX-1c |
Contribution (%) COX-2c |
P1 |
0.173 |
−6.36 |
−7.97 |
19 |
18 |
P2 |
0.429 |
−5.21 |
−5.13 |
15 |
28 |
P3 |
0.226 |
−11.28 |
−12.86 |
34 |
37 |
P4 |
0.173 |
−10.61 |
−7.49 |
32 |
17 |
Total (Σ)d |
1.000 |
|
- |
100 |
100 |
a Fraction (fi) was derived from the relative HPLC-determined concentration of each steryl ferulate as fi = Ci / ΣCi, where Ci is the compound concentration (mg/100 g extract).
b ΔG represents the validated binding energy (kcal/mol) obtained from AutoDock simulations.
c Contribution (%) was calculated using the equation Contributioni = (fi×|ΔGi|) / Σ(fi×|ΔGi|)×100, assuming a linear relationship between binding affinity and inhibitory potency.
d The total contribution (Σ = 100%) indicates the relative proportion of each steryl ferulate to the overall predicted COX inhibitory potential of the γ-oryzanol extract.
Discussion
γ-oryzanol
extracted from PTT1 rice bran selectively inhibited
COX-2 activity in LPS-stimulated RAW 264.7 macrophages, while COX-1
activity was not significantly affected. Regarding COX-1
biological function, it is constitutively expressed in enterocytes,
where it regulates prostaglandin
synthesis essential for maintaining intestinal integrity and
gastrointestinal homeostasis [10]. Consequently, the suppression of
COX-1, particularly by non-selective NSAIDs, has been associated
with gastrointestinal toxicity resulting from impaired mucosal
defense [26]. In the present study,
γ-oryzanol extract
exerted nonsignificant inhibitory effects on COX-1, which may be
advantageous by reducing the risk of gastrointestinal side effects
while preserving enterocyte integrity. Under LPS-induced
inflammatory conditions, γ-oryzanol extract demonstrated a
significant, dose-dependent inhibition of COX-2 activity. This
observation aligns with the biological role of COX-2 as an inducible
enzyme upregulated in response to pro-inflammatory stimuli such as
LPS, cytokines, and growth factors [27]. The elevation of COX-2
activity upon LPS stimulation reflects its central role in the
biosynthesis of pro-inflammatory prostaglandins, which mediate
macrophage activation and inflammatory signaling. Therefore, the
selective suppression of COX-2 by
γ-oryzanol suggests its
potential as a natural anti-inflammatory agent capable of
attenuating inflammatory responses without compromising
COX-1–mediated physiological functions. Our results are consistent
with previous evidence indicating that
γ-oryzanol
downregulates COX-2 expression in LPS-stimulated RAW 264.7
macrophages, primarily through inhibition of the ROS-ERK1/2-Egr-1
signaling cascade independently of NF-κB activation [6]. Previous
studies have demonstrated the anti-inflammatory potential of rice
bran–derived compounds. Rice bran extract has been shown to
inhibit COX-1 and COX-2 activities in a dose-dependent manner, with
preferential COX-2 inhibition in embryonic kidney 293 cells [28]. In
an in
vivo
study, γ-oryzanol suppressed COX-2 upregulation in dextran sulfate
sodium-induced colitis mice, supporting its COX-modulatory effects
under physiological conditions [29,30]. A recent in
silico
study further reported that γ-oryzanol exhibits favorable binding
affinities toward COX-1 (−7.8 kcal/mol) and COX-2 (−6.2
kcal/mol) at multiple active sites [5], consistent with our docking
observations and in
vitro
inhibitory activity results.
Molecular docking simulations were conducted to enhance the biochemical findings and clarify the structural and energetic foundations of COX selectivity among the γ-oryzanol steryl ferulates. Overall binding orientations inside the COX-2 catalytic cleft were identical for the four main components detected by HPLC, P1 P4. The bulky steryl moieties consistently occupied the hydrophobic wall formed by Phe504, Leu338, Val335, and Leu517, while the polar ferulate heads extended toward the solvent-exposed entrance, establishing secondary hydrogen-bonding interactions. None of these compounds interacted with the canonical catalytic residues Arg120, Tyr355, or Ser530, which serve as typical anchoring sites for acidic NSAIDs [31]. The lack of ionic interaction indicates the non-acidic ester characteristics of γ-oryzanol derivatives, implying that they function as neutral hydrophobic modulators instead of traditional carboxylate-anchored inhibitors.
Among the four steryl ferulates, P3 (campesteryl ferulate) demonstrated the most favorable interaction energies toward both COX isoforms. The negative ΔΔG and selectivity index indicate a modest preference for COX-2, aligning with the extract’s observed selectivity in cell-based assays. Importantly, comparison between hydrated and non-hydrated COX-2 models revealed that the presence of conserved water molecules significantly enhanced binding compactness. In the hydrated model, P3 formed shorter hydrogen bonds with Pro71, Glu510, and a bridging water molecule, while the hydrophobic contacts with Phe504, Leu338, Val509, Ala513, Met99, Val335, and Leu517 were tighter. Upon removal of water, the number and strength of H-bonds decreased, and the interatomic distances increased, indicating a less stable and more expanded configuration.
These findings highlight the critical role of hydration-mediated stabilization in the COX-2–γ-oryzanol interaction. Water molecules likely facilitate indirect hydrogen-bond bridges that reinforce ligand anchoring in the absence of the typical Arg120–Tyr355 ionic gate, thus improving binding affinity and conformational stability [32,33]. The predominance of hydrophobic and water-mediated interactions may underlie the COX-2 selectivity and reduced COX-1 interference observed for the γ-oryzanol extract, suggesting a pharmacophore distinct from classical NSAIDs but consistent with non-acidic natural anti-inflammatory compounds.
Integration of HPLC composition data with docking-derived mean binding energies (mean ΔG of Cluster 1) further revealed that γ-oryzanol’s overall COX-inhibitory potential is not solely determined by compound abundance but rather by the combined effects of molecular potency and extract composition. P3 contributed the greatest energetic share to COX-2 inhibition, followed by P2, P1, and P4. For COX-1, P3 and P4 were the major contributors, indicating a partial but mixed selectivity profile. Despite P2 being the most abundant steryl ferulate, its relatively weak binding energy suggests limited functional impact on overall COX inhibition.
The extract demonstrated a dose-dependent inhibition of COX-2 activity at the cellular level, reaching a maximum of 55% inhibition at a concentration of 2.50 mg/mL, while preserving COX-1 function. Biochemical and computational data collectively support a model wherein γ-oryzanol selectively reduces COX-2-mediated prostaglandin synthesis through direct enzyme binding (specifically via P3 and P1) and the upstream suppression of oxidative and MAPK-dependent signaling pathways [6]. The dual mechanism indicates that γ-oryzanol may serve as a safer anti-inflammatory agent, presenting a lower gastrointestinal risk relative to conventional non-selective NSAIDs.
Although docking analysis provides a mechanistic perspective on steryl ferulate-COX interactions, the biological COX-2 inhibition observed in RAW 264.7 cells reflects the integrated pharmacological effect of the extract in a complex cellular environment. Parameters such as cellular uptake, metabolic stability, solubility, protein binding, and possible synergistic or antagonistic interactions among steryl ferulates are not captured by docking simulations. Therefore, the docking results should be interpreted as supportive mechanistic evidence rather than direct predictors of cellular potency. The cell-based findings remain the primary biological evidence of COX modulation, and the computational results serve to complement, rather than replace, experimental observations. A further limitation is that the concentrations required to achieve COX inhibition in vitro are relatively high and may not be physiologically attainable. Addressing this issue will require additional research on the metabolism, bioavailability, and in vivo pharmacokinetics of γ-oryzanol to establish the physiological and pharmacological relevance of the current results.
Conclusions
γ-Oryzanol
extracted from PTT1 rice bran contained a high proportion of steryl
ferulates, particularly 24-methylenecycloartanyl ferulate, while
computational analysis indicated that campesteryl ferulate exhibited
the strongest predicted binding affinity toward COX isoforms.
Consistent with these findings, the extract demonstrated
dose-dependent
anti-inflammatory activity in RAW 264.7
macrophages through selective COX-2 inhibition. This selectivity
might suggest potential therapeutic benefits with reduced
gastrointestinal risks compared to non-selective COX inhibitors.
Further
in
vivo
studies are needed to validate COX-2
selectivity
and to evaluate pharmacokinetics, bioavailability, and safety.
Acknowledgments
This
project was supported by
Valaya
Alongkorn Rajabhat University under the Royal Patronage, Pathum
Thani Province, Thailand (Contract number
VRU-FF67/004). We
also gratefully acknowledge the School of Medicine, Mae Fah Luang
University, for providing academic resources and research guidance
throughout the study.
Declaration of generative AI and AI-assisted technologies in the writing process
During the preparation of this work, the authors used QuillBot to correct grammar during manuscript preparation. After using these tools, the authors reviewed and edited the content as needed and took full responsibility for the publication.
CRediT author statement
Shisanupong Anukanon: Conceptualization, Methodology, Computational analysis, and Writing original draft. Kanyanat Kaewiad: Investigation. Sakunta Manakla: Investigation. Jureemart Deeammart: Investigation. Komgrit Saeng-ngoen: Investigation. Yawanart Ngamnon: Investigation. Ngamnetr Rapan: Investigation. Yaiprae Chatree: Conceptualization, Methodology, Investigation, Writing original draft, Review and editing, and Funding acquisition.
References
[1] MG Sahini and E Mutegoa. Extraction, phytochemistry, nutritional, and therapeutical potentials of rice bran oil: A review. Phytomedicine Plus 2023; 3(2), 100453.
[2] H Juricic, M Cuccioloni, L Bonfili, M Angeletti, D Uberti, AM Eleuteri and V Cecarini. Biochemical, biological, and clinical properties of γ-Oryzanol. Antioxidants 2025; 14(9), 1099.
[3] T Hudson and J Katzinger. 196 - Menopause. In: J E Pizzorno and MT Murray (Eds.). Textbook of natural medicine. St. Louis MO, Churchill Livingstone, 2020.
[4] X Tian, X Wang, M Fang, L Yu, F Ma, X Wang, L Zhang and P Li. Nutrients in rice bran oil and their nutritional functions: A review. Critical Reviews in Food Science and Nutrition 2025; 65(14) 2840-2857.
[5] P Christine Natalia, S Anna, W Ernanin Dyah and F Fatchiyah. Virtual prediction of brown rice gamma oryzanol as anti-inflammation via COX-1 and COX-2 inhibitions. Berkala Penelitian Hayati 2023; 29(2), 57-66.
[6] SY Shin, HW Kim, HH Jang, YJ Hwang, JS Choe, JB Kim, et al. γ-Oryzanol suppresses COX-2 expression by inhibiting reactive oxygen species-mediated Erk1/2 and Egr-1 signaling in LPS-stimulated RAW264.7 macrophages. Biochemical and Biophysical Research Communications 2017; 491(2), 486-492.
[7] K Jin, C Qian, J Lin and B Liu. Cyclooxygenase-2–Prostaglandin E2 pathway: A key player in tumor-associated immune cells. Frontiers in Oncology 2023; 13, 1099811.
[8] B Hinz and A Pahl. Cyclooxygenase-1. In: SJ Enna and DB Bylund (Eds.). xPharm: The comprehensive pharmacology reference. New York, Elsevier, 2007, p. 1-5.
[9] M Tomić, A Micov, U Pecikoza, and R Stepanović-Petrović. Chapter 1 - Clinical uses of nonsteroidal anti-inflammatory drugs (NSAIDs) and potential benefits of NSAIDs modified-release preparations. In: B Čalija (Ed.). Microsized and nanosized carriers for nonsteroidal anti-inflammatory drugs. Academic Press, Boston, 2017.
[10] Y Faki and A Er. Different chemical structures and physiological/pathological roles of cyclooxygenases. Rambam Maimonides Medical Journal 2021; 12(1), e0003.
[11] I Morita. Distinct functions of COX-1 and COX-2. Prostaglandins & Other Lipid Mediators 2002; 68-69, 165-175.
[12] Y Liu, L Shi, W Qiu and Y Shi. Ferulic acid exhibits anti-inflammatory effects by inducing autophagy and blocking NLRP3 inflammasome activation. Molecular & Cellular Toxicology 2022; 18(4), 509-519.
[13] S Chimthai, S Cheabu, W Aesomnuk, S Ruengphayak, S Arikit, A Vanavichit and C Malumpong. Breeding for heat tolerant aromatic rice varieties and identification of novel QTL regions associated with heat tolerance during reproductive phase by QTL-Seq. Rice Science 2025; 32(1), 67-80.
[14] C Chinvongamorn and S Sansenya. The γ-oryzanol content of Thai rice cultivars and the effects of gamma irradiation on the γ-oryzanol content of germinated Thai market rice. Oriental Journal of Chemistry 2020; 36, 812-818.
[15] C Siripattanakulkajorn, P Sombutsuwan, A Nakornsadet, S Chumsantea, S Lilitchan, K Krisnangkura and K Aryusuk. Policosanol and other bioactive compounds in different Thai rice varieties. Journal of Food Composition and Analysis 2024; 126, 105891.
[16] M Yamsaray, T Sreewongchai, C Phumichai and P Chalermchaiwat. Yield and nutritional properties of improved red pericarp Thai rice varieties. ScienceAsia 2023; 49, 155-160.
[17] J Welch, J Wallace, AB Lansley and C Roper. Evaluation of the toxicity of sodium dodecyl sulphate (SDS) in the MucilAir™ human airway model in vitro. Regulatory Toxicology and Pharmacology 2021; 125, 105022.
[18] BS Selinsky, K Gupta, CT Sharkey and PJ Loll. Structural analysis of NSAID binding by prostaglandin H2 synthase: Time-dependent and time-independent inhibitors elicit identical enzyme conformations. Biochemistry 2001; 40(17), 5172-5180.
[19] JL Wang, D Limburg, MJ Graneto, J Springer, JR Hamper, S Liao, JL Pawlitz, RG Kurumbail, T Maziasz, JJ Talley, JR Kiefer and J Carter. The novel benzopyran class of selective cyclooxygenase-2 inhibitors. Part 2: The second clinical candidate having a shorter and favorable human half-life. Bioorganic & Medicinal Chemistry Letters 2010; 20(23), 7159-7163.
[20] GM Morris, R Huey, W Lindstrom, MF Sanner, RK Belew, DS Goodsell and AJ Olson. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. Journal of Computational Chemistry 2009; 30(16), 2785-2791.
[21] SD Oniga, L Pacureanu, CI Stoica, MD Palage, A Crăciun, LR Rusu, EL Crisan and C Araniciu. COX inhibition profile and molecular docking studies of some 2-(trimethoxyphenyl)-thiazoles. Molecules 2017; 22(9), 1507.
[22] LG Ferreira, RN Dos Santos, G Oliva and AD Andricopulo. Molecular docking and structure-based drug design strategies. Molecules 2015; 20(7), 13384-13421.
[23] MK Gilson and HX Zhou. Calculation of protein–ligand binding affinities. Annual Review of Biophysics and Biomolecular Structure 2007; 36, 21-42.
[24] DDL Minh. Implicit ligand theory: Rigorous binding free energies and thermodynamic expectations from molecular docking. Journal of Chemical Physics 2012; 137(10), 104106.
[25] TC Chou. Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Research 2010; 70(2), 440-446.
[26] KA Ko and DK Lee. Nonsteroidal anti-inflammatory drug-induced peptic ulcer disease. The Korean Journal of Helicobacter and Upper Gastrointestinal Research 2025; 25(1), 34-41.
[27] J Park, HJ Chang, KJ Hwang, SH Yum, CE Park, JH Kim and M Kim. Association of COX-2 selectivity in pain medication use with endometriosis incidence: Retrospective cohort study. Yonsei Medical Journal 2025; 66(6), 374-382.
[28] B Roschek Jr, RC Fink, D Li, M McMichael, CM Tower, RD Smith and RS Alberte. Pro-inflammatory enzymes, cyclooxygenase 1, cyclooxygenase 2, and 5-lipooxygenase, inhibited by stabilized rice bran extracts. Journal of Medicinal Food 2009; 12(3), 615-623.
[29] MS Islam, T Murata, M Fujisawa, R Nagasaka, H Ushio, AM Bari, M Hori and H Ozaki. Anti-inflammatory effects of phytosteryl ferulates in colitis induced by dextran sulphate sodium in mice. British Journal of Pharmacology 2008; 154(4), 812-824.
[30] X Xia, H Lin, F Luo, X Wu, L Zhu, S Chen and H Luo, F Ye, X Peng, Y Zhang, G Yang and Q Lin. Oryzanol ameliorates DSS-stimulated gut barrier damage via targeting the gut microbiota accompanied by the TLR4/NF-κB/NLRP3 cascade response in vivo. Journal of Agricultural and Food Chemistry 2022; 70(50), 15747-15762.
[31] A Zarghi and S Arfaei. Selective COX-2 inhibitors: A review of their structure–activity relationships. Iranian Journal of Pharmaceutical Research 2011; 10(4), 655-683.
[32] CA Rouzer and LJ Marnett. Structural and chemical biology of the interaction of cyclooxygenase with substrates and non-steroidal anti-inflammatory drugs. Chemical Reviews 2020; 120(15), 7592-7641.
[33] S Xu, DJ Hermanson, S Banerjee, K Ghebreselasie, GM Clayton, RM Garavito and LJ Marnett. Oxicams bind in a novel mode to the cyclooxygenase active site via a two-water-mediated H-bonding network. The Journal of Biological Chemistry 2014; 289(10), 6799-6808.